ISSN: 1839-9940
J Genomics 2020; 8:62-70. doi:10.7150/jgen.41196 This volume Cite
Research Paper
crispRdesignR: A Versatile Guide RNA Design Package in R for CRISPR/Cas9 Applications
Department of Biological Sciences, University of Massachusetts Lowell, 1 University Ave., Lowell, 01852 USA
Abstract
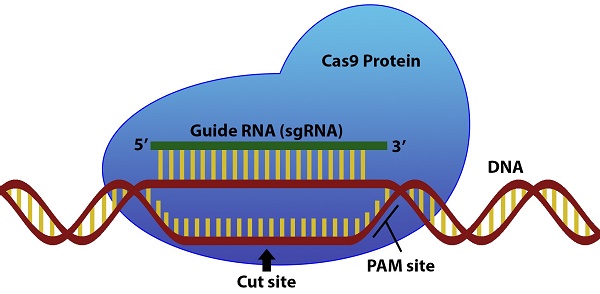
The success of CRISPR/Cas9 gene editing applications relies on the efficiency of the single guide RNA (sgRNA) used in conjunction with the Cas9 protein. Current sgRNA design software vary in the details they provide on sgRNA sequence efficiency and usually limit organism choice to a list of developer-selected species. The crispRdesignR package aims to address these limitations by providing comprehensive sequence features of the generated sgRNAs in a single program, which allows users to predict sgRNA efficiency and design sgRNA sequences for systems that currently do not have optimized efficiency scoring methods. crispRdesignR reports extensive information on all designed sgRNA sequences with robust off-target calling and annotation and can be run in a user-friendly graphical interface. The crispRdesignR package is implemented in R and has fully editable code for specialized purposes including sgRNA design in user-provided genomes. The package is platform independent and extendable, with its source code and documentation freely available at
Keywords: gRNA design, genome editing, guide efficiency, CRISPR, GUI, off-target

 Global reach, higher impact
Global reach, higher impact