ISSN: 1839-9940
J Genomics 2022; 10:33-38. doi:10.7150/jgen.68220 This volume Cite
Research Paper
Identification of the c.829_832delAATA Deletion Variants in the BRCA1 Gene Associated with Hereditary Breast/Ovarian Cancer ˗ Case Report
1. Department of Biotechnology, Microbiology and Human Nutrition, University of Life Sciences in Lublin, 8 Skromna St., 20-704 Lublin, Poland
2. Laboratory of Genetic Investigations, B. Dobrzanskiego 7 St., 20-262 Lublin
3. Center of Oncology of the Lublin Region St. Jana z Dukli, 7 K. Jaczewskiego St., 20-090 Lublin
4. Pathomorphology Department of Clinical Hospital I, I Independent Clinical Hospital, Stanislawa Staszica 16 St., 20-081 Lublin
Abstract
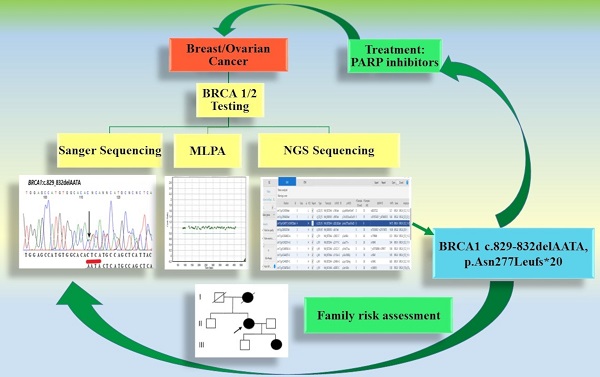
Determination of the BRCA1/BRCA2 mutation status in patients with breast and/or ovarian cancer is commonly performed using various molecular techniques. The use of targeted PCR-based tests only may not be sufficient, as not all possible variants are investigated. In the present study, we used next-generation sequencing (NGS) techniques to identify novel pathogenic variants in BRCA1 and BRCA2.
In this study, material (blood and FFPE) collected from a 67-year-old patient with ovarian cancer was used. The presence of hereditary mutations characteristic for the Polish population was examined using Sanger sequencing. BRCA1 and BRCA2 gene exons were amplified using the Devyser BRCA kit and sequenced on the Miniseq. No germline mutations characteristic for the Polish population were detected. However, 12 single nucleotide variants and 2 indels were identified. We found a new deleterious mutation of gene BRCA1 (c.829_832delAATA). To our knowledge, this mutation has not been reported yet in the Polish population and elsewhere. The use of the NGS technique increases the possibility of detecting mutational changes in patients with ovarian and/or breast cancer. Quick determination of pathogenic variants is important to facilitate specific therapy, in addition to the identification of familial predisposition to cancer.
Keywords: BRCA1, BRCA2, ovarian/breast cancer, NGS, PARP inhibitors

 Global reach, higher impact
Global reach, higher impact