ISSN: 1839-9940
J Genomics 2021; 9:1-5. doi:10.7150/jgen.46294 This volume Cite
Research Paper
Isolation, draft genome sequencing and identification of Enterobacter roggenkampii CCI9
1. Research Institute for Sustainable Chemistry, National Institute of Advanced Industrial Science and Technology (AIST), 3-11-32 Kagamiyama, Higashi-Hiroshima, Hiroshima 739-0046, Japan.
2. Department of Civil and Environmental Engineering, National Institute of Technology, Kure College, 2-2-11 Aga-minami, Kure, Hiroshima, 737-8506, Japan.
3. Graduate School of Integrated Sciences for Life, Hiroshima University, 1-3-1 Kagamiyama, Higashi-Hiroshima, Hiroshima 739-8530, Japan.
Abstract
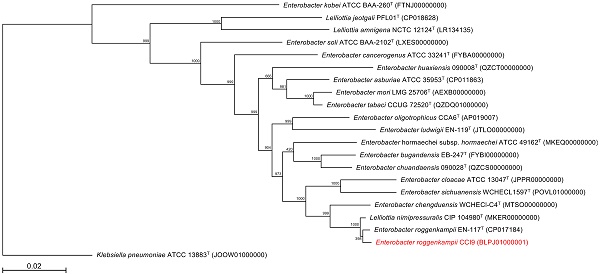
Strain CCI9, which was isolated from leaf soil collected in Japan, was capable of growth on poor-nutrient medium, at temperatures of 10°C to 45°C, at pHs of 4.5 to 10, and in the presence of 7.0% NaCl. We determined a draft genome sequence of strain CCI9, which consists of a total of 28 contigs containing 4,644,734 bp with a GC content of 56.1%. This assembly yielded 4,154 predicted coding sequences. Multilocus sequence analysis (MLSA) based on atpD, gyrB, infB, and rpoB gene sequences were performed to further identify strain CCI9. The MLSA revealed that strain CCI9 clustered tightly with Enterobacter roggenkampii EN-117T. Moreover, the average nucleotide identity value (98.6%) between genome sequences of strain CCI9 and E. roggenkampii EN-117T exceeds the cutoff value for prokaryotic subspecies delineation. Therefore, strain CCI9 was identified as E. roggenkampii CCI9. To clarify differences between E. roggenkampii EN-117T and CCI9, the coding proteins were compared against the eggNOG database.
Keywords: Enterobacter, Oligotroph, 16S rRNA, draft genome sequence, multilocus sequence analysis, average nucleotide identity

 Global reach, higher impact
Global reach, higher impact