ISSN: 1839-9940
J Genomics 2020; 8:80-83. doi:10.7150/jgen.50468 This volume Cite
Research Paper
Nucleotide sequencing of the HoxA gene cluster using Gorilla fosmid clones
Division of Population Genetics, National Institute of Genetics, Japan.
*Present address: Graduate School of Science and Engineering, Ibaraki University, Japan.
#Present address: Marine Ecosystem Research Center, Korea Institute of Ocean Science and Technology, Korea.
Received 2020-7-8; Accepted 2020-8-21; Published 2020-8-29
Abstract
We sequenced the western gorilla (Gorilla gorilla) HoxA cluster region using seven fosmid clones, and found that the total tiling path sequence was 214,185 bp from the 5' non-genic region of HoxA1 to the 3' non-genic region of Evx1. We compared the nucleotide sequence with the gorilla genome sequence in the NCBI database, and the overall proportion of nucleotide difference was estimated to be 0.0005-0.0007. These estimates are lower than overall genomic polymorphism in gorillas.
Keywords: Fosmid library, Gorilla, HoxA
Introduction
The euchromatic sequence of the human genome was sequenced using the hierarchical shotgun sequencing strategy, also known as clone-by-clone sequencing, by the International Human Genome Sequencing Consortium [1]. With the advent of next-generation sequencer (NGS), the hierarchical shotgun sequencing strategy is now less commonly used. However, hierarchical shotgun sequencing involving bacterial artificial chromosomes (BACs) and fosmids is still used for some purposes, such as to determine the long complete haploid in a chromosome region.
Kim et al. [2] constructed a western gorilla (Gorilla gorilla) fosmid library and established a simple polymerase chain reaction (PCR) screening system for it. They also selected seven fosmid clones, which constitutes the minimum tiling path for the entire HoxA gene cluster in the gorilla genome [2]. In this study we sequenced these fosmid clones and used them in our analysis.
Materials and Methods
Seven fosmid clones originated from one female gorilla individual “Taiko” (GGFP-562J15, GGFP-367A20, GGFP-347D05, GGFP-175G07, GGFP-452O13, GGFP-012E07, and GGFP-210K06) were screened and selected in a previous study [2]. These seven clones were used for this study. We determined nucleotide sequences primarily using the hierarchical shotgun sequencing method, as previously described [3], by sequencing each clone with more than tenfold-coverage. For base-calling, assembly, and to obtain a quality score for both raw and assembled data we used the Phred-Phrap software package [4]. Editing was performed with Consed [5]. Finishing was carried out by primer walking and PCR-coupled primer walking.
The fosmid sequences were compared with the nucleotide sequences (assembly NC_044609.1 from the Kamilah_GGO_v0 genome) available in the National Center for Biotechnology Information (NCBI) database. Pairwise sequence alignment was done manually using MEGA7 software [6]. Transitions, transversions, synonymous and nonsynonymous substitutions were counted using this software. Synonymous and nonsynonymous substitutions were estimated using Nei and Gojobori's method [7]. When counting substitutions, the cluster sequence was divided into coding sequence (CDS), intron, and inter-genic regions. The 53 amino acid residues (e.g. RTNFTTKQLTELEKEFHFNKYLTRARRVEIAASLQLNETQVKIWFQNRRMKQK from HOXA1) were used as the homeodomain regions.
Results and Discussion
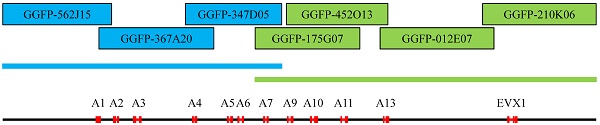
The nucleotide sequences of the seven fosmid clones are as follows: GGFP-562J15 (39,323 bp), GGFP-367A20 (41,479 bp), GGFP-347D05 (34,811 bp), GGFP-175G07 (37,837 bp), GGFP-452O13 (36,576 bp), GGFP-012E07 (41,057 bp), and GGFP-210K06 (40,950 bp) (Figure 1). We found four identical overlapping regions: 4,635 bp of nucleotides between GGFP-562J15 (AB125652) and GGFP-367A20 (AB125653); 10,316 bp of nucleotides between GGFP-367A20 (AB125653) and GGFP-347D05 (AB125654); 2,819 bp of nucleotides between GGFP-452O13 (AB125656) and GGFP-012E07 (AB125657); and 3,986 bp of nucleotides between GGFP-012E07 (AB125657) and GGFP-210K06 (AB125658).
Mapping of the seven fosmid clones that constitute the minimum tiling path for the entire HoxA gene cluster of the gorilla genome. The seven fosmid clones are indicated by white boxes. The two concatenated haploid sequences are indicated by thick bars. The CDS regions of the HoxA genes and the EVX1 gene are indicated by black boxes.

The overlap region between GGFP-175G07 (AB125655) and GGFP-452O13 (AB125656) had one indel in 26,513 bp. We used PCR to sequence this region using the genomic DNA of the gorilla (Taiko), and we confirmed the probable loss of one base in GGFP-452O13 (AB125656). The nucleotides in the overlap region between GGFP-347D05 (AB125654) and GGFP-175G07 (AB125655) differed by 0.09% (9/9,560 bp). We therefore linked and concatenated GGFP-562J15, GGFP-367A20, and GGFP-347D05 as “concatenated haploid sequence 1” (100,662 bp) and GGFP-175G07, GGFP-452O13, GGFP-012E07, and GGFP-210K06 as “concatenated haploid sequence 2” (123,103 bp) (Figure 1). The nucleotide difference of 0.09% between GGFP-347D05 (AB125654) and GGFP-175G07 (AB125655) can be considered the difference between maternal and paternal chromosomes.
We compared the nucleotide difference (p-distance) between our two concatenated haploid gorilla fosmid sequences and the whole-genome shotgun sequence of the gorilla (assembly NC_044609.1 from the Kamilah_GGO_v0 genome) (Table 1). The overall nucleotide difference in the HoxA cluster region between concatenated haploid sequence 1 and the shotgun sequence (100,577 bp) was 0.0005, and between concatenated haploid sequence 2 and the shotgun sequence (123,061 bp) was 0.0008. These estimates were lower than the genomic polymorphisms reported from two other western gorillas [Kamilah: 0.189% (0.00189), EB(JC): 0.178% (0.00178)] [8]. These results are reasonable, because nucleotide differences in the HoxA gene cluster region are expected to be lower than in other genomic regions.
Nucleotide differences in the HoxA gene cluster sequences between two gorillas
| Region | Bp compared | ts | tv | ts+tv | p-distance | s | n* |
|---|---|---|---|---|---|---|---|
| Concatenated haploid sequence 1 (AB125652-AB125654) vs. NC_044609.1 | |||||||
| 5p non-genic | 33,704 | 6 | 3 | 9 | 0.0003 | ||
| A1 CDS | 1,008 | 0 | 0 | 0 | 0.0000 | 0 | 0 |
| A1 intron | 465 | 0 | 0 | 0 | 0.0000 | ||
| intergenic A1-A2 | 4,822 | 0 | 0 | 0 | 0.0000 | ||
| A2 CDS | 1,128 | 1 | 0 | 1 | 0.0009 | 1 | 0 |
| A2 intron | 640 | 0 | 0 | 0 | 0.0000 | ||
| intergenic A2-A3 | 5,393 | 0 | 1 | 1 | 0.0002 | ||
| A3 CDS | 1,332 | 0 | 0 | 0 | 0.0000 | 0 | 0 |
| A3 intron | 1,396 | 2 | 0 | 2 | 0.0014 | ||
| intergenic A3-A4 | 18,521 | 4 | 5 | 9 | 0.0005 | ||
| A4 CDS | 963 | 0 | 1 | 1 | 0.0010 | 0 | 1 (D/H) |
| A4 intron | 546 | 1 | 0 | 1 | 0.0018 | ||
| intergenic A4-A5 | 11,081 | 8 | 5 | 13 | 0.0012 | ||
| A5 CDS | 813 | 1 | 0 | 1 | 0.0012 | 0 | 1 (G/S) |
| A5 intron | 960 | 1 | 1 | 2 | 0.0021 | ||
| intergenic A5-A6 | 2,030 | 1 | 1 | 2 | 0.0010 | ||
| A6 CDS | 702 | 0 | 0 | 0 | 0.0000 | 0 | 0 |
| A6 intron | 1,383 | 1 | 0 | 1 | 0.0007 | ||
| intergenic A6-A7 | 7,201 | 3 | 2 | 5 | 0.0007 | ||
| A7 CDS | 693 | 0 | 0 | 0 | 0.0000 | 0 | 0 |
| A7 intron | 917 | 1 | 0 | 1 | 0.0011 | ||
| intergenic A7-A9 | 4,879 | 5 | 1 | 6 | 0.0012 | ||
| sum | 100,577 | 35 | 20 | 55 | 0.0005 | ||
| Concatenated haploid sequence 2 (AB125655-AB125658) vs. NC_044609.1 | |||||||
| intergenic A6-A7 | 3,076 | 0 | 0 | 0 | 0.0000 | ||
| A7 CDS | 693 | 0 | 0 | 0 | 0.0000 | 0 | 0 |
| A7 intron | 932 | 0 | 0 | 0 | 0.0000 | ||
| intergenic A7-A9 | 7,049 | 2 | 4 | 6 | 0.0009 | ||
| A9 CDS | 819 | 0 | 1 | 1 | 0.0012 | 0 | 1 (S/C) |
| A9 intron | 1,036 | 0 | 0 | 0 | 0.0000 | ||
| intergenic A9-A10 | 6,456 | 3 | 4 | 7 | 0.0011 | ||
| A10 CDS | 1,233 | 1 | 0 | 1 | 0.0008 | 0 | 0 |
| A10 intron | 1,175 | 1 | 1 | 2 | 0.0017 | ||
| intergenic A10-A11 | 8,483 | 6 | 4 | 10 | 0.0012 | ||
| A11 CDS | 945 | 0 | 0 | 0 | 0.0000 | 0 | 0 |
| A11 intron | 1,399 | 1 | 0 | 1 | 0.0007 | ||
| intergenic A11-A13 | 13,035 | 7 | 4 | 11 | 0.0008 | ||
| A13 CDS | 1,167 | 0 | 0 | 0 | 0.0000 | 0 | 0 |
| A13 intron | 713 | 0 | 0 | 0 | 0.0000 | ||
| intergenic A13-EVX1 | 42,931 | 37 | 14 | 51 | 0.0012 | ||
| EVX1 CDS | 1,227 | 0 | 0 | 0 | 0.0000 | 0 | 0 |
| EVX1 introns | 2,177 | 0 | 0 | 0 | 0.0000 | ||
| 3p non-genic | 28,515 | 8 | 2 | 10 | 0.0004 | ||
| sum | 123,061 | 66 | 34 | 100 | 0.0008 | ||
ts: transition, tv: transversion, s: synonymous substitution, n: nonsynonymous substitution;
*Amino acid differences are shown in parentheses.
In the intron regions, the average p-distance was 0.0007, ranging from 0 [in HoxA1, HoxA2, HoxA7 (in concatenated haploid sequence 2)], HoxA9, HoxA13, and Evx1) to 0.0021 (in HoxA5).
In the CDS regions, the average p-distance was 0.0004, ranging from 0 [in HoxA1, HoxA3, HoxA6, HoxA7 (in concatenated haploid sequence 1), HoxA7 (in concatenated haploid sequence 2), HoxA11, HoxA13, and Evx1] to 0.0012 (in HoxA5 and HoxA9). One synonymous difference was observed in HoxA2. One nonsynonymous difference was observed for each of HoxA4, HoxA5, and HoxA9, although these changes were not located in homeobox regions.
An 18 bp gap was observed in exon 1 of HoxA10. The fosmid sequences were 18 bp longer than the whole-genome shotgun sequence of the gorilla (assembly NC_044609.1 from the Kamilah_GGO_v0 genome). When we examined the same region in humans (Homo sapiens), chimpanzees (Pan troglodytes), and orangutans (Pongo abelii), we found that they did not contain the 18 bp insertion. We used PCR to sequence this region in the DNA of the gorilla (Taiko) genome and confirmed that Taiko did have the 18 bp insertion. Because this insertion was observed in the non-homeobox region, we hypothesize that this insertion does not affect the fundamental function of HoxA10.
In the intergenic regions, the average p-distance was 0.0007, ranging from 0 [in the intergenic A1-A2 and A6-A7 (in concatenated haploid sequence 2)] to 0.0012 [in the intergenic A4-A5, A7-A9 (in concatenated haploid sequence 1), A10-A11, and A13-EVX1]. The differences observed in these intergenic regions were comparable with those observed in the intron regions.
The estimates for the CDS, introns, and intergenic regions were lower than the genomic polymorphisms reported from two other western gorillas (Kamilah: 0.00189, EB(JC): 0.00178) [8]. It has been previously reported that the four Hox gene clusters have the lowest density of interspersed repeats in the human genome [9], probably because of the large-scale cis-regulatory elements that cannot tolerate being interrupted by insertions [10-12]. The lower estimates we observed in the HoxA gene cluster region are therefore reasonable.
In conclusion, we successfully sequenced the HoxA cluster region of the gorilla. This region consists of two stretched haploid sequences that will be available for further analysis, including cis-regulatory element, linkage disequilibrium, and recombination.
Abbreviations
NGS: next-generation sequencer; BAC: bacterial artificial chromosome; PCR: polymerase chain reaction; NCBI: National Center for Biotechnology Information; CDS: coding sequence.
Acknowledgements
We would like to thank Akiko Aoshima, Atsuko Ide, and Yoshimi Noaki for their technical assistance. We would also like to thank Yuji Kohara of the Genome Biology Laboratory, National Institute of Genetics for the help with sequencing.
Nucleotide Sequence Accession Numbers
The nucleotide sequences from the seven fosmid clones for Gorilla gorilla have been deposited in the DDBJ/EMBL/GenBank International Nucleotide Sequence Database under accession numbers AB125652-AB125658.
Competing Interests
The authors have declared that no competing interest exists.
References
1. International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature. 2004;431(7011):931-945
2. Kim CG, Fujiyama A, Saitou N. Construction of a gorilla fosmid library and its PCR screening system. Genomics. 2003;82(5):571-574
3. Watanabe H, Fujiyama A, Hattori M. et al. DNA sequence and comparative analysis of chimpanzee chromosome 22. Nature. 2004;429(6990):382-388
4. Ewing B, Hillier L, Wendl MC. et al. Base-calling of automated sequencer traces using Phred. I. Accuracy assessment. Genome Research. 1998;8(3):175-185
5. Gordon D, Abajian C, Green P. Consed: a graphical tool for sequence finishing. Genome Research. 1998;8(3):195-202
6. Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution. 2016;33(7):1870-1874
7. Nei M, Gojobori T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Molecular Biology and Evolution. 1986;3(5):418-426
8. Scally A, Dutheil JY, Hillier LW. et al. Insights into hominid evolution from the gorilla genome sequence. Nature. 2012;483(7388):169-175
9. Lander ES, Linton LM, Birren B. et al. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860-921
10. Santini S, Boore JL, Meyer A. Evolutionary conservation of regulatory elements in vertebrate Hox gene clusters. Genome Research. 2003;13(6a):1111-1122
11. Matsunami M, Sumiyama K, Saitou N. Evolution of conserved non-coding sequences within the vertebrate Hox clusters through the two-round whole genome duplications revealed by phylogenetic footprinting analysis. Journal of Molecular Evolution. 2010;71(5-6):427-436
12. Matsunami M, Saitou N. Vertebrate paralogous conserved noncoding sequences may be related to gene expressions in brain. Genome Biology and Evolution. 2013;5(1):140-150
Author contact
Corresponding author: E-mail: takashi.kitano.evolutionibaraki.ac.jp.

 Global reach, higher impact
Global reach, higher impact