ISSN: 1839-9940
J Genomics 2020; 8:53-61. doi:10.7150/jgen.44650 This volume Cite
Research Paper
Draft genome sequence of Actinomadura sp. K4S16 and elucidation of the nonthmicin biosynthetic pathway
1. Biological Resource Center, National Institute of Technology and Evaluation (NBRC), Kisarazu, Chiba 292-0818, Japan.
2. Biotechnology Research Center and Department of Biotechnology, Toyama Prefectural University, Imizu, Toyama 939-0398, Japan.
3. NBRC, Shibuya-ku, Tokyo 151-0066, Japan.
4. Faculty of Science, Kasetsart University, Bangkok, Thailand.
Received 2020-2-7; Accepted 2020-4-21; Published 2020-5-17
Abstract
Actinomadura sp. K4S16 (=NBRC 110471) is a producer of a novel tetronate polyether compound nonthmicin. Here, we report the draft genome sequence of this strain together with features of the organism and assembly, annotation and analysis of the genome sequence. The 9.6 Mb genome of Actinomadura sp. K4S16 encoded 9,004 putative ORFs, of which 7,701 were assigned with COG categories. The genome contained four type-I polyketide synthase (PKS) gene clusters, two type-II PKS gene clusters, and three nonribosomal peptide synthetase (NRPS) gene clusters. Among the type-I PKS gene (t1pks) clusters, a large t1pks cluster was annotated to be responsible for nonthmicin synthesis based on bioinformatic analyses. We also performed feeding experiments using labeled precursors and propose the biosynthetic pathway of nonthmicin.
Keywords: Actinomadura, biosynthesis, nonthmicin, polyether, polyketide, tetronate
Introduction
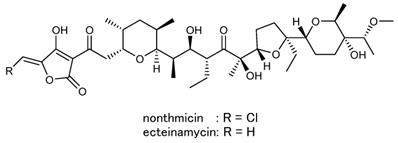
Actinomycetes are well known as a promising source for diverse bioactive secondary metabolites. Especially, members of Streptomyces have attracted attention as the most useful screening sources for new drug leads and a large number of bioactive compounds have been identified from cultures of this genus [1,2]. Consequently, the chance of finding novel secondary metabolites from Streptomyces members has recently dwindled. Thus, the focus of screening has recently moved to less exploited genera of rare actinomycetes [3]. In our screening for novel bioactive compounds from rare actinomycetes, Actinomadura sp. K4S16 was isolated from rice field soil in Thailand and found to produce a tetronate polyether designated nonthmicin along with ecteinamycin (Fig. 1) [4]. Nonthmicin shows inhibitory activity against tumor cell invasion and protective activity for neuronal cell damage. This new polyether compound is characterized by the tetronic acid functionality modified by a chlorine atom. Halogenated tetronic acids are not known from nature except nonthmicin. In this study, we conducted whole genome shotgun sequencing of the strain to elucidate the biosynthetic pathway of nonthmicin. We herein present the draft genome sequence of Actinomadura sp. K4S16, together with the taxonomical identification of the strain, description of its genome properties and annotation of the gene cluster for nonthmicin biosynthesis. Biosynthetic pathway for nonthmicin was predicted by bioinformatics analysis and confirmed by precursor-incorporation experiments.
Chemical structures of nonthmicin and ecteinamycin.
Materials and Methods
Sequenced strain
In the course of screening for novel bioactive substances from rare actinomycetes, Actinomadura sp. K4S16 was isolated from rice field soil collected in Thailand and found to produce a novel polyketide compound named nonthmicin and its known congener ecteinamycin (Fig. 1) [4]. Actinomadura sp. K4S16 was preserved as TP-A0891 at the Toyama Prefectural University, deposited into the NBRC culture collection, and publicly available from the collection as NBRC 110471.
Chemotaxonomic analyses
The isomer of diaminopimelic acid in the whole-cell hydrolysate was analyzed according to the method described by Hasegawa et al. [5]. Isoprenoid quinones and cellular fatty acids were analyzed as described previously [6].
Phylogenetic analysis based on 16S rRNA gene sequences
PCR template was prepared according to the protocol for Gram-positive bacteria of DNeasy Blood & Tissue kit (Qiagen). The gene encoding 16S rRNA was amplified by PCR using two universal primers, 9F and 1541R. After purification of the PCR product by AMPure (Beckman Coulter), the sequencing was carried out according to an established method [7]. Homology search of the sequence was conducted using EzBioCloud [8]. A phylogenetic tree was reconstructed by on the basis of the 16S rRNA gene sequence together with taxonomically close type strains showing more than 98% similarities by ClustalX2 [9].
Growth conditions and genomic DNA preparation
A monoisolate of Actinomadura sp. K4S16, isolated as single colony, was grown on polycarbonate membrane filter (Advantec) on double-diluted NBRC 227 agar medium (0.2% yeast extract, 0.5% malt extract, 0.2% glucose, 2% agar, pH 7.3) at 28°C. High quality genomic DNA for sequencing was extracted and isolated from the mycelia with an EZ1 DNA Tissue Kit and a BioRobot EZ1 (Qiagen) according to the manufacturer's protocol for extraction of nucleic acid from Gram-positive bacteria. The size, purity, and double-strand DNA concentration of the genomic DNA were measured by pulsed-field gel electrophoresis, ratio of absorbance values at 260 nm and 280 nm, and Quant-iT PicoGreen dsDNA Assay Kit (Life Technologies), respectively, to assess the quality of genomic DNA.
Genome sequencing and assembly
Shotgun and paired-end libraries were prepared and subsequently sequenced using 454 pyrosequencing technology and MiSeq (Illumina) paired-end technology, respectively (Table 1). The 82 Mb shotgun sequences and 707 Mb paired-end sequences were assembled using Newbler v2.8 and subsequently finished using GenoFinisher [10] to yield 43 scaffolds larger than 500 bp. The draft genome sequence has been deposited in the INSDC database under the accession number BDDE01000001-BDDE01000043. The project information and its association with MIGS version 2.0 compliance are summarized in Table 1 [11].
Project information
| MIGS ID | Property | Term |
|---|---|---|
| MIGS 31 | Finishing quality | Improved-high-quality draft |
| MIGS-28 | Libraries used | 454 shotgun library, Illumina paired-end library |
| MIGS 29 | Sequencing platforms | 454 GS FLX+, Illumina MiSeq |
| MIGS 31.2 | Fold coverage | 8.6 x, 73 x, respectively |
| MIGS 30 | Assemblers | Newbler v2.8, GenoFinisher |
| MIGS 32 | Gene calling method | Progidal |
| Locus tag | K4S16 | |
| GenBank ID | BDDE00000000 | |
| GenBank date of release | Aug, 2019 | |
| GOLD ID | Not registered | |
| BioProject | PRJDB4748 | |
| MIGS 13 | Source material identifier | NBRC 110471 |
| Project relevance | Industrial |
Genome annotation
Coding sequences were predicted with Prodigal [12] and tRNA-scanSE [13]. The gene functions were assigned using an in-house genome annotation pipeline, and domains related to polyketide synthase (PKS) and nonribosomal peptide synthetase (NRPS) were searched using the SMART and PFAM domain databases. PKS and NRPS gene clusters and their domain organizations were determined as reported previously [7]. Substrates of adenylation (A) and acyltransferase (AT) domains were predicted using antiSMASH [14]. Protein-protein BLAST search against the NCBI Non-redundant protein sequences (nr) database was also used for predicting function of proteins encoded in the nonthmicin biosynthetic gene cluster.
Digital DNA-DNA hybridization
Digital DNA-DNA hybridization (DDH) between Actinomadura sp. K4S16 and A. mexicana DSM 44485T (FZNP01000001-FZNP01000053) was conducted using Formula 2 of Genome-to-Genome Distance Calculator 2.1 [15].
Feeding experiments using labeled precursors
Inoculation, cultivation, extraction, and purification were performed in the same manner as previously reported [4]. Supplementation of sodium [1-13C]acetate or [1-13C]propionate (20 mg/100 ml medium/flask, 10 flasks) was initiated at 48 h after inoculation and periodically carried out every 24 h for four times. After further incubation for 24 h, the whole culture broths were extracted with 1-butanol and several steps of purification yielded 55 mg and 100 mg of 13C-labeled nonthmicin, respectively.
Results and Discussion
Feature, classification, and genome properties
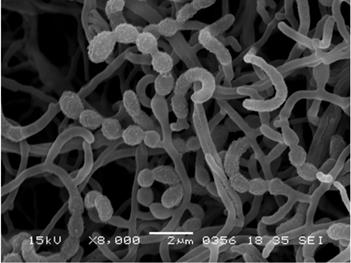
The general feature of Actinomadura sp. K4S16 is shown in Table 2. This strain grew well on ISP 2 and ISP 4 agar media, but poorly on ISP 5 and ISP 7. The color of aerial mycelia was white and that of the reverse side was pale orange on ISP 2 agar medium. Strain K4S16 formed extensively branched substrate mycelium. The aerial mycelium formed short chains of arthrospores. A scanning electron micrograph of this strain (Fig. 2) shows that spore chains are hooked or spiral (1 turn) and the spore surface is rugose. Growth occurred at 20-45 °C (optimum 28 °C) and pH 5-8 (optimum pH 7). Strain K4S16 exhibited growth with 0-2 % (w/v) NaCl (optimum 0 % NaCl) and the strain utilized arabinose, fructose, glucose, mannitol, rhamnose, sucrose, and xylose as sole carbon source for energy and growth, but not raffinose (all at 1%, w/v).
Classification and general features of Actinomadura sp. K4S16 [11]
| MIGS ID | Property | Term | Evidence codea |
|---|---|---|---|
| Classification | Domain Bacteria | TAS [25] | |
| Phylum Actinobacteria | TAS [26] | ||
| Class Actinobacteria | TAS [27] | ||
| Order Actinomycetales | TAS [27-30] | ||
| Suborder Streptosporangineae | TAS [27] | ||
| Family Thermomonosporaceae | TAS [27,30,31] | ||
| Genus Actinomadura | TAS [29,32] | ||
| Species undetermined (a new genomospecies) | This study | ||
| strain: K4S16 | TAS [4] | ||
| Gram stain | Not tested, likely positive | NAS | |
| Cell shape | Branched mycelia | IDA | |
| Motility | Not reported | ||
| Sporulation | Sporulating | IDA | |
| Temperature range | 20 °C to 45 °C | IDA | |
| Optimum temperature | 28 °C | IDA | |
| pH range; Optimum | 5 to 8; 7 | IDA | |
| Carbon source | Arabinose, fructose, glucose, mannitol, rhamnose, sucrose, xylose | IDA | |
| MIGS-6 | Habitat | Rice field soil | NAS |
| MIGS-6.3 | Salinity | 0 % to 2 % NaCl | IDA |
| MIGS-22 | Oxygen requirement | Aerobic | IDA |
| MIGS-15 | Biotic relationship | Free-living | IDA |
| MIGS-14 | Pathogenicity | Not reported | |
| MIGS-4 | Geographic location | Thailand | TAS [4] |
| MIGS-5 | Sample collection | March 13, 2010 | NAS |
| MIGS-4.1 | Latitude | Not reported | |
| MIGS-4.2 | Longitude | Not reported | |
| MIGS-4.4 | Altitude | Not reported |
a Evidence codes - IDA: Inferred from Direct Assay; TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). These evidence codes are from the Gene Ontology project [33].
Scanning electron micrograph of Actinomadura sp. K4S16 grown on double-diluted ISP 2 agar for 7 days at 28 °C. Bar, 2 µm.
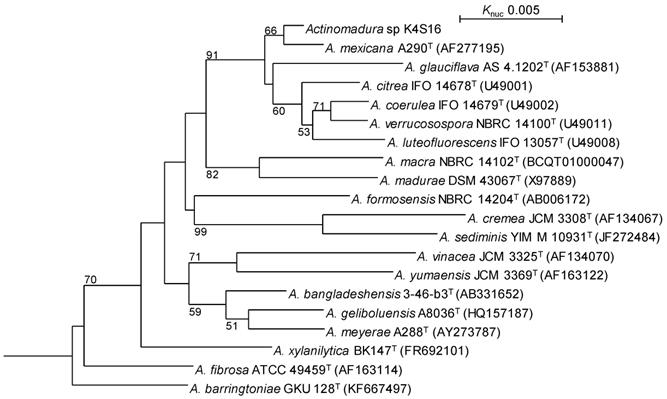
The whole-cell hydrolysate of strain K4S16 contained meso-diaminopimelic acid as its diagnostic peptidoglycan diamino acid. The predominant menaquinones were identified as MK-9(H4) and MK-9(H6); in addition, MK-9(H2) and MK-9(H8) were also detected as minor components. The major cellular fatty acids (>10%) were C16:0 and C18:1 ω9c. The 16S rRNA gene sequence of the strain indicated the highest similarity (99.58 %, 1415/1421) to Actinomadura mexicana A290T (AF277195) as the closest type strain. The phylogenetic analysis confirmed that the strain belongs to the genus Actinomadura (Fig. 3).
Phylogenetic tree based on 16S rRNA gene sequences of Actinomadura sp. K4S16 and its phylogenetically close type strains showing over 98.0 % sequence similarities. Accession numbers for 16S rRNA genes are shown in parentheses. The tree uses sequences aligned by ClustalX2 [9] and constructed by the neighbor-joining method [24]. All positions containing gaps were eliminated. The building of the tree also involves a bootstrapping process repeated 1,000 times to generate a majority consensus tree, and only bootstrap values above 50% are shown at branching points. Streptosporangium roseum DSM 43021T was used as an outgroup.
A draft genome size of Actinomadura sp. K4S16 was 9,647,292 bp and the G+C content was 72.4 % (Table 3). Of the total 9,068 genes, 9,004 were protein-coding genes and 64 were RNA genes. The classification of genes into COGs functional categories is shown in Table 4. Digital DDH between Actinomadura sp. K4S16 and the type strain of the closest species, A. mexicana DSM 44485T suggested that the DNA-DNA relatedness was 49.0 %, which is below 70 %, the cut-off point for the assignment of bacterial strains to the same species [16]. This suggests that Actinomadura sp. K4S16 is a novel independent genomospecies.
Genome statistics of Actinomadura sp. K4S16
| Attribute | Value | % of Total |
|---|---|---|
| Genome size (bp) | 9,647,292 | 100 |
| DNA coding (bp) | 8,684,283 | 90.0 |
| DNA G+C (bp) | 6,982,736 | 72.4 |
| DNA scaffolds | 43 | - |
| Total genes | 9,068 | 100 |
| Protein coding genes | 9,004 | 99.3 |
| RNA genes | 64 | 0.7 |
| Pseudogenes | - | - |
| Genes in internal clusters | 4,198 | 46.6 |
| Genes with function prediction | 5,358 | 59.5 |
| Genes assigned to COGs | 7,701 | 85.5 |
| Genes with Pfam domains | 2,655 | 29.5 |
| Genes with signal peptides | 618 | 6.8 |
| Genes with transmembrane helices | 2,022 | 22.5 |
| CRISPR repeats | 1 | - |
Number of genes associated with general COG functional categories
| Code | Value | %age | Description |
|---|---|---|---|
| J | 275 | 3.1 | Translation, ribosomal structure and biogenesis |
| A | 4 | 0.0 | RNA processing and modification |
| K | 1,158 | 12.9 | Transcription |
| L | 444 | 4.9 | Replication, recombination and repair |
| B | 3 | 0.0 | Chromatin structure and dynamics |
| D | 52 | 0.6 | Cell cycle control, cell division, chromosome partitioning |
| V | 202 | 2.2 | Defense mechanisms |
| T | 627 | 7.0 | Signal transduction mechanisms |
| M | 365 | 4.1 | Cell wall/membrane biogenesis |
| N | 46 | 0.5 | Cell motility |
| U | 87 | 1.0 | Intracellular trafficking and secretion |
| O | 232 | 2.6 | Posttranslational modification, protein turnover, chaperones |
| C | 540 | 6.0 | Energy production and conversion |
| G | 709 | 7.9 | Carbohydrate transport and metabolism |
| E | 851 | 9.5 | Amino acid transport and metabolism |
| F | 131 | 1.5 | Nucleotide transport and metabolism |
| H | 260 | 2.9 | Coenzyme transport and metabolism |
| I | 435 | 4.8 | Lipid transport and metabolism |
| P | 465 | 5.2 | Inorganic ion transport and metabolism |
| Q | 453 | 5.0 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 1,459 | 16.2 | General function prediction only |
| S | 614 | 6.8 | Function unknown |
| - | 1,303 | 14.5 | Not in COGs |
The total is based on the total number of protein coding genes in the genome.
PKS and NRPS gene clusters in the genome
We analyzed biosynthetic gene clusters for polyketides and nonribpsomal peptides in the genome. Actinomadura sp. K4S16 harbored four type-I PKS gene (t1pks) clusters, two type-II PKS gene (t2pks) clusters, and three NRPS gene (nrps) clusters, as shown in Table 5. T1pks-1 cluster encoded only a PKS composed of ACP-KS/AT/DH/KR/ACP/ACP-TE domains, which showed 87% sequence identity to phenolpthiocerol synthesis type-I polyketide synthase PpsD of Mycobacterium tuberculosis 401416 (CND43678), suggesting it may synthesize phenolpthiocerol-like compounds. T1pks-2 cluster encoded two PKSs whose domain organizations are KS/AT/KR and KS/AT, respectively. Since these PKSs did not show sequence similarities to PKSs whose products are identified and the domain organization is unusual, we are not able to predict the product. T1pks-3 cluster encoded a PKS composed of KS/AT/KR/DH domains. Because such domain organization is specific to iterative PKSs for enediyne syntheses, this gene cluster likely synthesizes enediyne-type polyketide compounds. T1pks-4 cluster is responsible for nonthmicin synthesis as stated in the following section. T2pks-1 cluster might synthesize aromatic compounds similar to tetarimycin A or mithramycin, because its KSα showed 70 to 71 % sequence identities to TamM (AFY23044) and MtmP (CAA61989). T2pks-2 cluster did not show high sequence similarities (less than 55 % identities) to any PKSs registered in GenBank, suggesting that the product will be unique. Nrps-1 gene cluster harbored six NRPS modules and the products were predicted to be peptides containing amino dihydroxybenzoic acid, cysteine, glycine, and methyl ornithine. Nrps-2 gene cluster encoded four modules and the products will be composed of starter molecule-Cys-Cys-methyl Cys. Nrps-3 gene cluster had seven modules and the products are likely hexapeptides including amino acid residues such as alanine and threonine. The presence of these PKS and NRPS gene clusters suggests that this strain has the potential to produce diverse polyketide- and nonribosomal peptide-compounds as the secondary metabolites.
PKSs and NRPSs encoded in each PKS or NRPS gene cluster of Actinomadura sp. K4S16
| Cluster | ORF (K4S16_) | Domain | Predicted product |
|---|---|---|---|
| t1pks-1 | 13_05730 | ACP-KS/AT/DH/KR/ACP/ACP-TE | phenolpthiocerol-like polyketide |
| t1pks-2 | 18_02330 | KS/AT/KR | unpredictable |
| 18_02340 | KS/AT | ||
| t1pks-3 | 23_01060 | KS/AT/KR/DH | enediyne-type polyketide |
| t1pks-4 | 09_00520 | KS/AT/ACP-KS/AT/DH/KR/ACP-KS/AT/DH/ER/KR/ACP | nonthmicin, ecteinamycin |
| 09_00630 | KS/AT/DH/KR/ACP-KS/AT/DH/ER/KR/ACP | ||
| 09_00640 | KS/AT/KR/ACP-KS/AT/KR/ACP | ||
| 09_00660 | KS/AT/ACP | ||
| 09_00750* | KS/AT/KR/ACP-KS/AT/DH/ER/KR/ACP-KS/AT/DH/KR/ACP | ||
| 09_00770* | KS/AT/KR/ACP | ||
| t2pks-1 | 11_01150 | KSα | aromatic polyketide |
| 11_01140 | KSβ (CLF) | ||
| 11_01130 | ACP | ||
| t2pks-2 | 09_01590 | KSα | aromatic polyketide |
| 09_01600 | KSβ (CLF) | ||
| 09_01610 | ACP | ||
| nrps-1 | 11_07800 | T-C/ACys/T | peptide containing DHB, Cys, Gly, & mOrn |
| 11_07770 | ADHB | ||
| 11_07630 | C/AGly/T-C/A/T-C/AOrn/MT/T-C/T | ||
| nrps-2 | 16_01040 | T-C/ACys/T-C/ACys/T | St-Cys-Cys-mCys |
| 16_01050 | C/ACys/MT/T-TE | ||
| nrps-3 | 24_00530 | C/AAla/T-C/A/T-C/T-C/AThr/T/E | Ala-x-Thr-x-Thr-x |
| 24_00520 | C/A/T-C/AThr/T/E | ||
| 24_00510 | C/A/T-TE |
Substrates of A domains are shown by subscripts. DHB, dihydroxybenzoic acid; mOrn, methyl ornithine; St, starter molecule; mCys, methyl cysteine; x, unpredicted amino acid. *encoded in the complementary strand.
Nonthmicin biosynthetic pathway
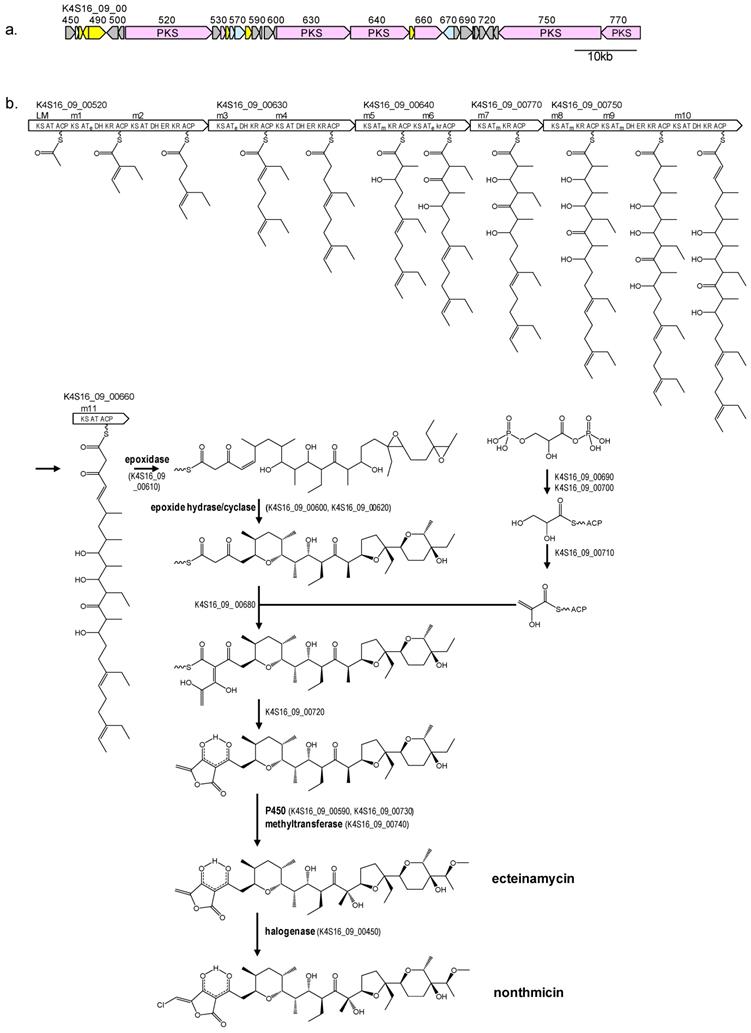
The chemical structure of nonthmicin suggested that their carbon skeletons are assembled from five malonyl-CoA, four methylmalonyl-CoA, and three ethylmalonyl-CoA molecules by a type-I PKS pathway. We therefore searched for a t1pks cluster consisting of 12 PKS modules. Among all of the four t1pks clusters present in Actinomadura sp. K4S16 (Table 5), t1pks-4 cluster encoded six large PKSs and several enzymes related to secondary metabolite syntheses (Table 6, Fig 4a) and its assembly line contains 12 PKS modules. Substrates of AT domains in modules 1, 3 and 6 were predicted to be ethylmalonyl-CoA, whereas those in modules 5, 7, 8 and 9 were methymalonyl-CoA. According to the collinearity rule of type-I PKS pathways [17] and the chemical structure of nonthmicin, the polyketide backbone synthesized by the PKS assembly line was predicted as shown in Fig. 4b. The predicted structure is in good accordance with the nonthmicin backbone. The elongated polyketide chain is then converted to form three polyether moieties by an epoxidase and epoxide hydrase/cyclase(s) in a similar manner for the nanchangmycin biosynthesis [18]. The tetronic acid part may be synthesized by ORFs K4S16_09_00680 to K4S16_09_00720 as proposed for tetronic acid-containing polyketides such as tetrocarcin A, chlorothricin, abyssomicin, and quatromicin [19-22], because these ORFs are orthologues of TcaDs, ChlM and ChlDs, AbyAs and QunDs. Two cytochrome P450s (K4S16_09_00590 and K4S16_09_00730) and a methyltransferase (K4S16_09_00740) are probably responsible for the introduction of one hydroxy group and one methoxy group to produce ecteinamycin. Chlorination to the tetronate moiety is presumably catalyzed by a halogenase (K4S16_09_00450) to yield nonthmicin. On the basis of these bioinfomatic evidences, we here propose the biosynthetic pathway of nonthmicin and ecteinamycin as shown Fig. 4b.
ORFs of t1pks-4 gene cluster responsible for the synthesis of nonthmicin
| ORFa | Size (aa) | Deduced function | Closest protein homolog [origin] | Id/Si (%)b | Accession number |
|---|---|---|---|---|---|
| 00450 | 551 | halogenase | halogenase B [Actinoplanes sp. ATCC 33002] | 56/72 | AAQ04685 |
| 00460 | 188 | flavin reductase | flavin reductase-like, FMN-binding [Saccharopolyspora erythraea NRRL 2338] | 47/62 | CAM04194 |
| 00470 | 220 | two-component system response regulator | response regulator receiver protein, partial [Microbispora rosea] | 65/78 | WP_030509695 |
| 00480c | 353 | two-component system histidine kinase | hypothetical protein [Herbidospora cretacea] | 43/63 | WP_030450128 |
| 00490 | 906 | transcriptional regulator | ATPase [Microbispora sp. ATCC PTA-5024] | 41/54 | ETK35445 |
| 00500c | 576 | 3-hydroxybutyryl-CoA dehydrogenase | 3-hydroxybutyryl-CoA dehydrogenase [Streptomyces rapamycinicus NRRL 5491] | 56/65 | AGP52928 |
| 00510c | 340 | 3-oxoacyl-ACP synthase | 3-oxoacyl-ACP synthase III [Streptomyces sp. C] | 61/70 | EFL20209 |
| 00520 | 4,859 | polyketide synthase | polyketide synthase [Streptomyces albus] | 54/64 | AEZ53945 |
| 00530 | 442 | crotonyl-CoA reductase | NADPH:quinone reductase [Streptomyces albulus PD-1] | 74/83 | EXU89989 |
| 00540 | 258 | type-II thioesterase | thioesterase [Streptomyces sp. C] | 59/69 | EFL20221 |
| 00550 | 224 | transcriptional regulator | hypothetical protein [Actinomadura madurae] | 54/70 | WP_021595170 |
| 00560 | 310 | ABC transporter ATP-binding protein | hypothetical protein [Lechevalieria aerocolonigenes] | 77/86 | WP_030471487 |
| 00570 | 531 | ABC transporter permease protein | hypothetical protein [Actinopolymorpha alba] | 60/74 | WP_020580181 |
| 00580 | 388 | transcriptional regulator | LuxR-family transcriptional regulator [Actinokineospora sp. EG49] | 42/52 | EWC63761 |
| 00590 | 398 | cytochrome P450 | cytochrome P450 [Streptomyces bingchenggensis BCW-1] | 52/68 | ADI04501 |
| 00600c | 136 | epoxide hydrolase/cyclase | epoxide hydrolase [Streptomyces longisporoflavus] | 53/67 | ACR50776 |
| 00610 | 467 | epoxidase | hypothetical protein SBI_01389 [S. bingchenggensis BCW-1] | 58/70 | ADI04510 |
| 00620c | 183 | epoxide hydrolase/cyclase | putative epoxide hydrolase/cyclase [Streptomyces albus subsp. albus] | 53/66 | CCD31907 |
| 00630 | 3,941 | polyketide synthase | lasalocid modular polyketide synthase [Streptomyces sp. C] | 54/64 | EFL20211 |
| 00640 | 3,165 | polyketide synthase | polyketide synthase [Streptomyces graminofaciens] | 50/61 | BAJ16467 |
| 00650 | 263 | transcriptional regulator | putative pathway specific activator [S. longisporoflavus] | 55/67 | ACR50789 |
| 00660 | 1,563 | polyketide synthase | Beta-ketoacyl synthase [Streptomyces violaceusniger Tu 4113] | 50/62 | AEM87323 |
| 00670c | 576 | ABC transporter permease protein | Putative exporter of polyketide antibiotics- like protein [Thermomonospora curvata DSM 43183] | 45/60 | ACZ00124 |
| 00680 | 342 | 3-oxoacyl-ACP synthase | 3-oxoacyl-ACP synthase [Streptomyces sp. CNQ865] | 65/77 | WP_027767626 |
| 00690 | 637 | glyceroyl-ACP biosynthesis protein | methoxymalonyl-ACP biosynthesis protein FkbH [Streptomyces monomycini] | 60/69 | WP_033040494 |
| 00700 | 75 | ACP | ACP [Amycolatopsis orientalis] | 72/81 | AFI57025 |
| 00710 | 265 | 2-oxoglutarate dehydrogenase | acyltransferase [Streptomyces olindensis] | 68/81 | KDN76174 |
| 00720 | 365 | hydrolase or acyltransferase | 2-oxoacid dehydrogenase/acyltransferase [Micromonospora chalcea] | 54/65 | ACB37748 |
| 00730c | 398 | cytochrome P450 | cytochrome P450 [S. bingchenggensis BCW-1] | 53/71 | ADI04501 |
| 00740c | 288 | methyltransferase | SAM-dependent methyltransferase [Streptomyces sp. NRRL F-2890] | 48/59 | WP_030734046 |
| 00750c | 5,524 | polyketide synthase | Acyl transferase [S. violaceusniger Tu 4113] | 50/61 | AEM84952 |
| 00770c | 2,121 | polyketide synthase | PieA2 [Streptomyces piomogenus] | 51/62 | AEZ54375 |
a locus tag number after K4S16_09_; b identity/similarity; c encoded in the complementary strand.
Genetic map of nonthmicin biosynthetic gene cluster of Actionomadura sp. K4S16 (a) and the predicted biosynthetic pathway (b). Pink, PKS; yellow, transcriptional regulator; light blue, transporter; gray, genes related to secondary metabolite synthesis. AT, acyltransferase for malonyl-CoA; ATm, acyltransferase for methylmalonyl-CoA; ATe, acyltransferase for ethylmalonyl-CoA; kr, inactive KR.
Feeding experiments using labeled precursors
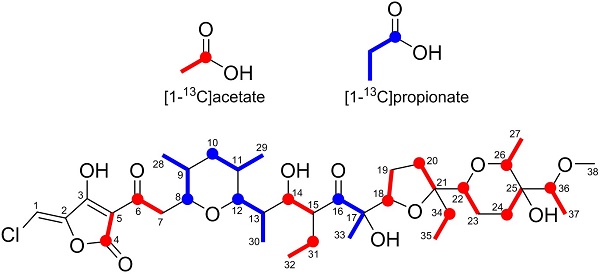
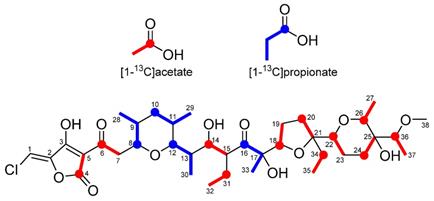
To verify the predicted biosynthetic pathway for nonthmicin, feeding experiments were carried out using 13C-labeled precursors such as [1-13C]acetate and [1-13C]propionate. The signal intensities in 13C NMR spectrum of these labeled nonthmicin is shown in Table 7. Feeding of sodium [1-13C]acetate gave enrichments at twelve carbons at C4, C6, C14, C18, C20, C22, C24, C25, C26, C31, C34, and C36. [1-13C]propionate feeding enriched four methyl carbons at C28, C29, C30, and C33 (Fig. 5). These results unambiguously established that the polyether polyketide structure of nonthmicin is assembled from five malonyl-CoA, four methylmalonyl-CoA, and three ethylmalonyl-CoA. Labeling of C4 and C5 by acetate and non-labeling of C1, C2, and C3 by any precursors indicated that tetronic acid moiety is derived from one acetate and one glycerate units [23]. These results also supported by annotated ORFs of t1pks-4 cluster (K4S16_09_00690, K4S16_09_00700, K4S16_09_00710) (Fig. 4b, Table 6).
Incorporation of 13C-labeled precursors into nonthmicin
| Position | δC | Relative enrichmentsa | |
|---|---|---|---|
| [1-13C]acetate | [1-13C]propionate | ||
| 1 | 98.9 | 0.78 | 0.80 |
| 2 | 147.8 | 0.79 | 0.64 |
| 3 | 181.4 | 0.78 | 0.89 |
| 4 | 169.6 | 2.56 | 0.98 |
| 5 | 96.4 | 0.82 | 0.62 |
| 6 | 199.8 | 2.28 | 0.84 |
| 7 | 35.4 | 0.94 | 1.24 |
| 8 | 77.7 | 1.33 | 4.25 |
| 9 | 27.7 | 0.86 | 0.87 |
| 10 | 36.2 | 1.09 | 4.85 |
| 11 | 28.8 | 1.10 | 0.74 |
| 12 | 69.7 | 1.60 | 4.70 |
| 13 | 36.4 | 1.06 | 0.95 |
| 14 | 73.7 | 2.25 | 0.89 |
| 15 | 48.8 | 0.87 | 0.80 |
| 16 | 222.8 | 1.28 | 4.49 |
| 17 | 78.6 | 0.80 | 0.64 |
| 18 | 83.9 | 2.41 | 0.88 |
| 19 | 24.8 | 1.13 | 0.95 |
| 20 | 29.0 | 2.14 | 0.95 |
| 21 | 88.8 | 0.71 | 0.84 |
| 22 | 70.2 | 2.64 | 0.90 |
| 23 | 19.9 | 0.96 | 0.98 |
| 24 | 24.8 | 2.19 | 1.15 |
| 25 | 73.0 | 2.49 | 1.15 |
| 26 | 74.6 | 2.47 | 0.89 |
| 27 | 14.1 | 1.00 | 0.90 |
| 28 | 17.5 | 1.01 | 0.86 |
| 29 | 11.1 | 0.93 | 0.91 |
| 30 | 7.2 | 0.94 | 0.75 |
| 31 | 21.3 | 3.11 | 0.87 |
| 32 | 12.6 | 0.93 | 0.72 |
| 33 | 20.5 | 0.86 | 1.00 |
| 34 | 30.0 | 2.19 | 0.96 |
| 35 | 9.1 | 0.91 | 1.05 |
| 36 | 78.9 | 3.25 | 0.96 |
| 37 | 11.1 | 0.82 | 0.87 |
| 38 | 56.2 | 0.80 | 0.93 |
a 13C signal intensity of each peak in the labeled 1 divided by that of the corresponding signal in the unlabeled 1, respectively, normalized to give an enrichment ratio of 1 for the unenriched peak of C27 and C33. The numbers in bold type indicate 13C-enriched atoms from 13C-labeled precursors.
Incorporation of 13C-labeled precursors into nonthmicin.
Precursor-directed biosynthesis of bromo-analogue of nonthmicin
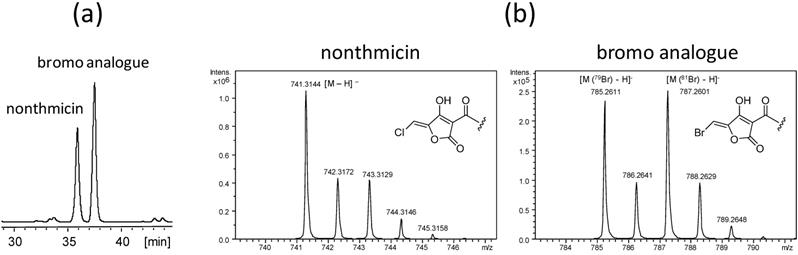
A putative halogenase gene (K4S16_09_00450), showing 56% identity and 72% similarity of amino acid sequence to HalB from Actinoplanes sp. ATCC 33002, present in the nonthmicin biosynthetic gene cluster was expected to be responsible for the halogenation (Table 6). If this gene product is also active for bromine, it can be used for the precursor-directed biosynthesis of a brominated analogue. In fact, supplementation of sodium bromide into the culture resulted in the production of a new nonthmicin congener (Fig. 6a) in which the chlorine atom was replaced by the bromine atom. The structure of the bromo analogue was confirmed analysing data by MS (Fig. 6b) and NMR (data not shown).
HPLC chromatogram of the bromine supplemented culture broth (a) and MS spectra of nonthmicin and bromine analogue (b).
Conclusion
We successfully found the type-I PKS gene cluster for nonthmicin biosynthetic and proposed a plausible biosynthetic pathway by the genome analysis of Actinomadura sp. K4S16, a producer of nonthmicin. Incorporation experiments of 13C-labeled precursors also suggested that nonthmicin is biosynthesized by PKS pathway. These findings will provide significant information not only for the biosynthetic mechanism but also for the genetic engineering to synthesize more potential bioactive molecules based on the nonthmicin structure.
Abbreviations
A: adenylation; ABC: ATP-binding cassette; ACP: acyl carrier protein; Ala: alanine; AT: acyltransferase; ATP: adenosine triphosphate; BLAST: Basic Local Alignment Search Tool; C: condensation; CLF: chain length factor; CoA: coenzyme A; COG: Clusters of Orthologous Groups; Cys: cysteine; DH: dehydratase; DHB: dihydroxybenzoic acid; E: epimerase; ER: enoylreductase; Gly: glycine; ISP: International Streptomyces project; KS: ketosynthase; KR: ketoreductase; kr: inactive KR; MIGS: minimum information about a genome sequence; MT: methyltransferase; NBRC: Biological Resource Center, National Institute of Technology and Evaluation; NMR: nuclear magnetic resonance; NRPS: nonribosomal peptide synthetase; nrps: NRPS gene; PKS: polyketide synthase; t1pks: type-I PKS gene; t2pks: type-II PKS gene; T: thiolation; TE: thioesterase; Thr: threonine.
Acknowledgements
This research was supported by a Grant-in-aid for Scientific Research from the Ministry of Education, Culture, Sports, and Technology of Japan to YI. We also thank Dr. Akane Kimura, Ms. Tomoko Hanamaki, Ms. Aya Uohara, Ms. Satomi Miura, Ms. Satomi Saitou and Ms. Chiyo Shibata for assistance to this study.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Berdy J. Bioactive microbial metabolites. J Antibiot. 2005;58:1-26
2. Watve MG, Tickoo R, Jog MM, Bhole BD. How many antibiotics are produced by the genus Streptomyces? Arch Microbiol. 2001;176:386-390
3. Lazzarini A, Cavaletti L, Toppo G, Marinelli F. Rare genera of actinomycetes as potential producers of new antibiotics. Antonie van Leeuwenhoek. 2000;78:399-405
4. Igarashi Y, Matsuoka N, In Y, Kataura T, Tashiro E, Saiki I, Sudoh Y, Duangmal K, Thamchaipenet A. Nonthmicin, a polyether polyketide bearing a halogen-modified tetronate with neuroprotective and antiinvasive activity from Actinomadura sp. Org Lett. 2017;19:1406-1409
5. Hasegawa T, Takizawa M, Tanida S. A rapid analysis for chemical grouping of aerobic actinomycetes. J Gen Appl Microbiol. 1983;29:319-322
6. Hamada M, Yamamura H, Komukai C, Tamura T, Suzuki K, Hayakawa M. Luteimicrobium album sp. nov, a novel actinobacterium isolated from a lichen collected in Japan, and emended description of the genus Luteimicrobium. J Antibiot. 2012;65:427-431
7. Komaki H, Ichikawa N, Hosoyama A, Fujita N, Igarashi Y. Draft genome sequence of marine-derived Streptomyces sp. TP-A0598, a producer of anti-MRSA antibiotic lydicamycins. Stand Genomic Sci. 2015;10:58
8. Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J. Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int System Evol Microbiol. 2017;67:1613-1617
9. Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG. Clustal W and Clustal X version 2.0. Bioinformatics. 2007;23:2947-2948
10. Ohtsubo Y, Maruyama F, Mitsui H, Nagata Y, Tsuda M. Complete genome sequence of Acidovorax sp. strain KKS102, a polychlorinated-biphenyl degrader. J Bacteriol. 2012;194:6970-6971
11. Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, Ashburner M, Axelrod N, Baldauf S, Ballard S, Boore J, Cochrane G, Cole J, Dawyndt P, De Vos P, DePamphilis C, Edwards R, Faruque N, Feldman R, Gilbert J, Gilna P, Glockner FO, Goldstein P, Guralnick R, Haft D, Hancock D, Hermjakob H, Hertz-Fowler C, Hugenholtz P, Joint I, Kagan L, Kane M, Kennedy J, Kowalchuk G, Kottmann R, Kolker E, Kravitz S, Kyrpides N, Leebens-Mack J, Lewis SE, Li K, Lister AL, Lord P, Maltsev N, Markowitz V, Martiny J, Methe B, Mizrachi I, Moxon R, Nelson K, Parkhill J, Proctor L, White O, Sansone SA, Spiers A, Stevens R, Swift P, Taylor C, Tateno Y, Tett A, Turner S, Ussery D, Vaughan B, Ward N, Whetzel T, San Gil I, Wilson G, Wipat A. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol. 2008;26:541-547
12. Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics. 2010;11:119
13. Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997;25:955-964
14. Weber T, Blin K, Duddela S, Krug D, Kim HU, Bruccoleri R, Lee SY, Fischbach MA, Muller R, Wohlleben W, Breitling R, Takano E, Medema MH. antiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 2015;43:W237-243
15. Meier-Kolthoff JP, Auch AF, Klenk HP, Goker M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics. 2013;14:60
16. Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WE, Murray RGE, Stackebrandt E, Starr MP, Trüper HG. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int Journal Syst Bacteriol. 1987;37:463-464
17. Fischbach MA, Walsh CT. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem Rev. 2006;106:3468-3496
18. Sun Y, Zhou X, Dong H, Tu G, Wang M, Wang B, Deng Z. A complete gene cluster from Streptomyces nanchangensis NS3226 encoding biosynthesis of the polyether ionophore nanchangmycin. Chem Biol. 2003;10:431-441
19. Fang J, Zhang Y, Huang L, Jia X, Zhang Q, Zhang X, Tang G, Liu W. Cloning and characterization of the tetrocarcin A gene cluster from Micromonospora chalcea NRRL 11289 reveals a highly conserved strategy for tetronate biosynthesis in spirotetronate antibiotics. J Bacteriol. 2008;190:6014-6025
20. Gottardi EM, Krawczyk JM, von Suchodoletz H, Schadt S, Muhlenweg A, Uguru GC, Pelzer S, Fiedler HP, Bibb MJ, Stach JE, Süssmuth RD. Abyssomicin biosynthesis: formation of an unusual polyketide, antibiotic-feeding studies and genetic analysis. ChemBioChem. 2011;12:1401-1410
21. He HY, Pan HX, Wu LF, Zhang BB, Chai HB, Liu W, Tang GL. Quartromicin biosynthesis: two alternative polyketide chains produced by one polyketide synthase assembly line. Chem Biol. 2012;19:1313-1323
22. Jia XY, Tian ZH, Shao L, Qu XD, Zhao QF, Tang J, Tang GL, Liu W. Genetic characterization of the chlorothricin gene cluster as a model for spirotetronate antibiotic biosynthesis. Chem Biol. 2006;13:575-585
23. Sun Y, Hong H, Gillies F, Spencer JB, Leadlay PF. Glyceryl-S-acyl carrier protein as an intermediate in the biosynthesis of tetronate antibiotics. ChemBioChem. 2008;9:150-156
24. Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406-425
25. Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Nat Acad Sci USA. 1990;87:4576-4579
26. Garrity GM, Holt JG. The road map to the manual. In: Garrity GM, Boone DR and Castenholz RW (eds), Bergey's Manual of Systematic Bacteriology, 2nd ed. Volume 1, New York: Springer. 2001:119-169
27. Stackebrandt E, Rainey FA, Ward-Rainey NL. Proposal for a new hierarchic classification system, Actinobacteria classis nov. Int System Evol Microbiol. 1997;47:479-491
28. Buchanan RE. Studies in the nomenclature and classification of the bacteria: II. The primary subdivisions of the Schizomycetes. J Bacteriol. 1917;2:155-164
29. Skerman VBD, McGowan V, Sneath PHA. Approved lists of bacterial names. Int System Evol Microbiol. 1980;30:225-420
30. Zhi XY, Li WJ, Stackebrandt E. An update of the structure and 16S rRNA gene sequence-based definition of higher ranks of the class Actinobacteria, with the proposal of two new suborders and four new families and emended descriptions of the existing higher taxa. Int System Evol Microbiol. 2009;59:589-608
31. Zhang Z, Kudo T, Nakajima Y, Wang Y. Clarification of the relationship between the members of the family Thermomonosporaceae on the basis of 16S rDNA, 16S-23S rRNA internal transcribed spacer and 23S rDNA sequences and chemotaxonomic analyses. Int System Evol Microbiol. 2001;51:373-383
32. Lechevalier HA, Lechevalier MP. A critical evaluation of the genera of aerobic actinomycetes. In The Actinomycetales (edited by Prauser H). Jena: Gustav Fischer Verlag. 1970:95-405
33. Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25-29
Author contact
Corresponding author: Hisayuki Komaki, NBRC, 2-5-8 Kazusakamatari, Kisarazu, Chiba 292-0818, Japan. E-mail: komaki-hisayukigo.jp.

 Global reach, higher impact
Global reach, higher impact