ISSN: 1839-9940
J Genomics 2020; 8:30-36. doi:10.7150/jgen.43928 This volume Cite
Research Paper
Programmable CRISPR interference for gene silencing using Cas13a in mosquitoes
Department of Biology, New Mexico State University, PO Box 30001 MSC 3AF, Las Cruces NM, 88003, USA
Received 2020-1-13; Accepted 2020-2-17; Published 2020-3-1
Abstract
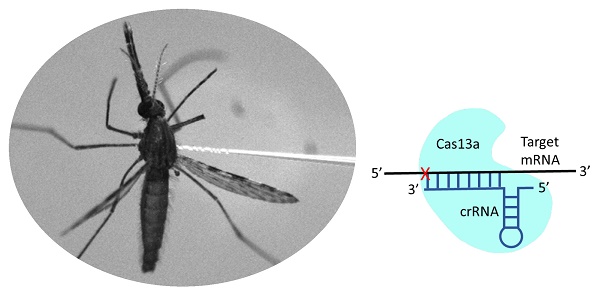
In the CRISPR-Cas systems, Cas13a is an RNA-guided RNA nuclease specifically targeting single strand RNA. We developed a Cas13a mediated CRISPR interference tool to target mRNA for gene silencing in mosquitoes. A Cas13a expressing plasmid was delivered to mosquitoes by intrathoracic injection, and Cas13a transcripts were detectable at least 10 days post-delivery. The target specific crRNA was synthesized in vitro using T7 RNA polymerase. The Cas13a plasmid and target crRNA can be delivered by intrathoracic injection together, or the Cas13a construct can be provided first, and then target crRNA can be given later when appropriate. The machinery was tested in two mosquito species. In Anopheles gambiae, vitellogenin gene was silenced by Cas13a/Vg-crRNA, which was accompanied by a significant reduction in egg production. In Aedes aegypti, the α- and δ-subunits of COPI genes were silenced by Cas13a/crRNA, which resulted in mortality and fragile midguts, reproducing a phenotype reported previously. Co-silencing genes simultaneously is achievable when a cocktail of target crRNAs is given. No detectable collateral cleavages of non-target transcripts were observed in the study. In addition to dsRNA or siRNA mediated RNA interference, the programmable CRISPR interference method offers an alternative to knock down genes in mosquitoes.
Keywords: CRISPR-Cas13a, RNA interference, Anopheles gambiae, Aedes aegypti, gene silencing, CRISPRi
Introduction
Characterization of mosquito life traits via functional genomics approaches can inform innovative control strategies through the identification of genes involved in various physiological processes, such as development, host-seeking, blood feeding, digestion, fecundity, immunity, xenobiotic metabolism and insecticide resistance. The last is perhaps the most immediately impactful, as, with increasing insecticide resistance, the array of options for vector control is shrinking and in dire need of replenishment. RNA interference (RNAi) based approaches have been widely used to identify genes that are relevant to vector competence, and RNAi-based effectors for mosquito control have been developed [1-5]. As an alternative or complement to RNAi-based tools, CRISPR-Cas9 based genome editing tools have been developed for gene function study in mosquito research field [6-8].
CRISPR-Cas systems are adaptive immune mechanisms used by prokaryotes to defend against invading DNA and RNA [9-12]. Cas9 is a RNA-guided DNA nuclease and once assembled with a CRISPR guide RNA (sgRNA), Cas9 is able to cleave target DNA in a highly specific fashion, and DNA-targeting Cas9 has been harnessed for genome editing [13, 14]. Furthermore, catalytically inactive Cas9 (dCas9) was adapted for manipulation of gene expression. The dCas9 can be fused with a gene repressor or transcription activator. Guided by CRISPR RNA, such dCas9 proteins are able to bind target promoter or exonic DNA sequence without cleavage, and either repress transcription (CRISPR interference, CRISPRi) or activate transcription of target genes (CRISPR activation, CRISPRa) [15-17]. Lately, Cas13 RNA nucleases [10, 18, 19], the new members in the CRISPR nuclease family, have been repurposed to specifically target endogenous RNAs as well as viral RNAs [11, 20-24]. Most Cas13 proteins are single “effector” proteins with two Higher Eukaryotes and Prokaryotes Nucleotide-binding (HEPN) domains [10, 25]. Once loaded with a target- specific crRNA, a Cas13 protein will locate target RNA and execute nuclease activity to degrade the target. Unlike Cas9, no Protospacer Adjacent Motif (PAM) sequences are required for Cas13 to function. Although a Protospacer Flanking Site (PFS), A, U, C, may be present for PspCas13b activity [26], no PFS is needed for LwaCas13a [11]. The Cas13 proteins tested for human RNA knockdown thus far have demonstrated high specificity and exhibited negligible, and significantly fewer, off-target effects than matched RNAi short hairpin RNAs (shRNA) using to trigger RNAi [11, 26]. In bacteria, Cas13 HEPN-nuclease is able to cleave not only the target-RNA in cis but also other non-target RNA present in trans [10]. Interestingly, no collateral effect was observed in three studies of CRISPR-Cas13 using human or plant cell lines [11, 26, 27], but Cas13a associated collateral RNA cleavage was reported in human glioma cancer cells [28]. Notably, Cas13 can effectively silence several transcripts in parallel [11, 20, 27, 29]. Taken together, CRISPR-Cas13 systems have become a new, exciting engine for CRISPRi [30].
In this paper, we present the evidence that the Cas13a mediated mRNA targeting is applicable in mosquitoes by demonstrating its efficacy in silencing genes in two mosquito species, Anopheles gambiae and Aedes aegypti, the major vectors of malaria and mosquito-borne viruses, respectively.
Materials and Methods
Plasmid construction
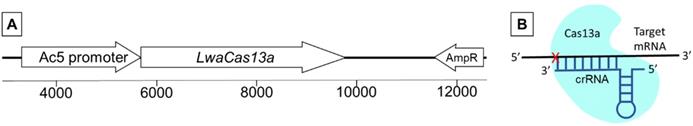
The Cas13a from Leptotrichia wadei belongs to the class 2 type VI RNA-guided RNA nucleases [11]. Its RNA targeting effect has been demonstrated in human and plant cells [11, 20, 28, 31]. Plasmid pAc-sgRNA-Cas9 was used as a template to engineer construct pAc-Cas13a (Fig. 1). Plasmid pAc-sgRNA- Cas9 was a gift from Ji-Long Liu (Addgene plasmid # 49330). The codons of LwaCas13a gene were optimized to Drosophila preference. The codon optimized Cas13a sequence was synthesized and cloned into the pAc-Cas13a plasmid at GenScript (https://www. genscript.com). The Cas13a was under the control of Drosophila actin (Ac5) promoter for constitutive Cas13a expression. The plasmid is replicable in E. coli host cells. The ampicillin resistance gene (AmpR) allows selection of positive clones (Fig.1A). The Cas13a sequence was confirmed by sequencing. The plasmid was extracted using a QIAprep Spin Miniprep kit (Cat No.27104, Qiagen). The sequence of plasmid pAc-LwaCas13a was deposited in NCBI under accession number MN812663.
Cas13a construct and function. (A) Map of pAc-LwaCas13a (partial). (B) Mechanism of Cas13a/crRNA mediated target RNA cleavage.
Synthesis of crRNA
A crRNA consists of a 36-nt direct repeat (DR) sequence and 28-nt target RNA specific sequence (N28), and the sequence of N28 is complementary to the target RNA sequence (Fig.1 B). The T7 promoter sequence (AGTTAATACGACTCACTATAGG) was added to the 5' end of the DR sequence (GATTTAGACTACCCCAAAAACGAAGGGGACTAAAAC) to enable crRNA synthesis using T7 RNA polymerase in vitro. The target specific sequences (N28) used in this study are shown in Table 1. The selection of target (N28) is straightforward as LwaCas13a does not require PFS for activity [11]. Please note, the target sequence for COPI-α was accidently designed as a-27nt sequence. For Vg, COPI-α and COPI-δ, one target crRNA was used for silencing, and for Caspar and Cactus, two target crRNAs were used. Template DNA duplexes of the crRNAs (T7-DR-N28) were synthesized at IDT Inc. (https://www.idtdna.com). The crRNAs were synthesized using T7-RNA polymerase (RPOLT7-RO ROCHE, Sigma-Aldrich). The crRNA synthesis reactions were set up in 40 μl containing template DNA duplex (1μg), 1 mM each of nucleotides ATP, GTP, CTP and UTP, 10X reaction buffer, T7 RNA polymerase 40U, and RNase inhibitor 20U. The reactions were incubated overnight at 37ºC and terminated by heating the mixture at 65ºC for 5 minutes. The crRNAs were treated with Turbo DNase I Kit (AM1907, ThermoFisher) to remove template DNA. The crRNA yield was quantified using a NanoDrop and stored at -20 ºC until use. Control crRNA (ctr crRNA) consisted of a randomly scrambled N28 nucleotide sequence, which had no homologous hit in the genomes of An. gambiae and Ae. aegypti.
The crRNA (N28) sequences.
| Gene | Gene ID | GenBank accession | crRNA sequence (N28) |
|---|---|---|---|
| Vg | AGAP004203 | XM_313104 | CACCTGCACCTTCACGCTGTCACCAGCC |
| COPI-δ | AAEL013230 | XM_001663354 | TGATAGACATACCGCACGGAATCTGTCT |
| COPI-α | AAEL015001 | XM_001663259 | ACCGCCTCTGCTTGTAGTTCCACACCT |
| Cactus-1 | AAEL000709 | XM_001650217 | ATCACCGTCGTCGTTCTGGTGGAAGTAC |
| Cactus-2 | AAEL000709 | XM_001650217 | TGATGCACAGGTCGTCCACCTTCATCGG |
| Caspar-1 | AAEL014738 | XM_021842247 | TCAACGCCGGACTCGGCCAGTGTCGTAC |
| Caspar-2 | AAEL014738 | XM_021842247 | TACCACTGCCACCGGCGGACGATCTCTG |
| Control | N/A | N/A | GACGCACATTCATAGTCTTCATCTGAGT |
Construct and crRNA Delivery
An. gambiae G3 strain and Ae. aegypti Puerto Rico strain were obtained from MR4 BEI and maintained using rearing conditions described previously [32, 33]. The pAc-Cas13a construct (0.5μg/μl) was delivered into one-day old adult female mosquitoes by intrathoracic injection. To aid construct delivery into cells, the plasmid was mixed with a transfecting agent FuGENE HD (E2311, Promega). FuGENE is a non-liposomal reagent containing lipids and other proprietary components [34]. The reagent has been used to facilitate delivery of Cas9 expressing plasmid to transform human cell lines [35] and Drosophila cell lines as well [36]. The construct solution was prepared with 1.6μl of FuGENE reagent and 10μg plasmid DNA in 20 μl volume for injection. The final concentration of construct DNA was 0.5 µg/µl in the mixture. Approximately, each An. gambiae mosquito received 100nl mixture, and each Ae. aegypti mosquito received 150nl mixture. Gene specific crRNAs were either delivered with the construct or separately at a later time point. For blood inducible genes, Vg and COPI, female mosquitoes were given a blood meal three days post construct injection. Corresponding crRNAs (0.5μg/μl, prepared in FuGENE as described above) were intrathoracically injected into mosquito hemocoel at two hours post blood meal. For co-delivery of the construct and crRNA, the construct (0.5µg/µl) and crRNA (0.5µg/µl) were mixed together with 1.6μl of FuGENE reagent in 20 μl. The mixture was injected into one-day-old female mosquitoes intrathoracically.
RNA isolation, cDNA synthesis and PCR
Total RNA from whole mosquitoes was isolated using Trizol (Invitrogen) following the manufacturer's instruction. The RNA was treated with Turbo DNase I Kit to remove genomic DNA contamination, and then 1μg RNA was converted to cDNA using Protoscript II RT (M0368S, New England Biolabs) following the manufacturer's instruction. The PCR assays were performed using 1μl 1:5 diluted cDNA as template, 0.2 μM primers (primer sequences are presented in Table S1) and 2 × PCR Master mix (M0482S, NEB), with the following cycling parameters: 35 cycles of denaturing at 95°C for 15 seconds, annealing at a temperature optimal for the amplicon (Table S1) for 15 seconds, and extension at 68°C for 20 seconds with an extra 5 min in the last cycle for final extension.
Statistical analysis
In the Vg gene knockdown experiment, eggs were dissected from ovaries at day 3 post blood meal. The egg counts were compared between the Vg crRNA and control crRNA cohorts. The non-parametric Mann-Whitney test was used for statistical comparison of the egg numbers. In the COPI gene knockdown experiment, a survival curve was plotted using GraphPad Prism, and a Mantel-Cox analysis was performed to compare the survival between the COPI crRNA and control crRNA cohorts.
Results
Cas13a expression in mosquitoes post intrathoracic delivery
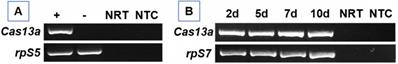
A construct was engineered to express Cas13a gene by modifying a plasmid that was successfully used to transfect Drosophila cells for targeted genetic mutagenesis previously by Bassett et al. (2014) [37]. As shown in Fig. 1, Cas13a coding sequence is under the control of constitutive promoter Ac5. The plasmid was prepared with transfection reagent FuGENE HD and injected into thorax of one-day-old mosquitoes. Cas13a transcription in whole mosquitoes was determined by RT-PCR. In An. gambiae, the RNA was sampled at day three post construct delivery, and Cas13a transcripts were detected (Fig. 2A). No Cas13a amplification was observed in the controls that did not receive the construct. In Ae. aegypti, the RNA samples were collected on day 2, 5, 7 and 10 post construct delivery, and Cas13a transcripts were detected in all time points (Fig. 2B).
Detection of Cas13a transcript in pAC-Cas13 injected mosquitoes. (A) RT-PCR of Cas13a in An. gambiae mosquitoes that received the construct (+) vs. control (-) at day 3 post plasmid delivery. NRT: No RT control; NTC: no template control. The rpS5 was used as a loading control. (B) Expression pattern of Cas13a transcript over a period of 10 days in Ae. aegypti, the rpS7 was used as a loading control.
Cas13a mediated Vitellogenin gene silencing in An. gambiae
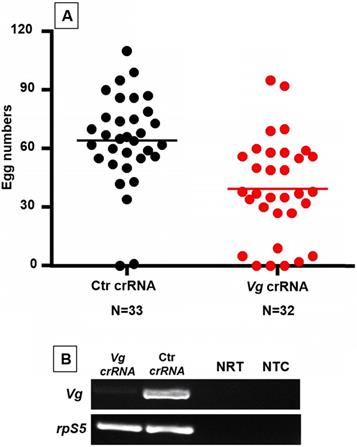
In mosquitoes, yolk protein precursor vitellogenins (Vg) are required for the vitellogenic stage in oogenesis after blood feeding [38]. To test Cas13a/crRNA mediated Vg silencing, the Cas13a construct was delivered to one-day-old An. gambiae (N = 120). Three days later, the mosquitoes were given a blood meal to induce Vg expression (N = 96). To enable Vg knockdown, Vg crRNA (N = 41) or control crRNA (N = 40) were injected into the blood engorged females at 2 hr post feeding. The Vg RT-PCR was used for verification of Vg knockdown. As shown in Fig. 3, the abundance of Vg transcript was reduced in females that received Vg-crRNA as compared to females that received control-crRNA. As expected, successful Vg knockdown resulted in reduction in egg production (Fig. 3). The Vg-crRNA treated mosquitoes produced on average 39 ± 25 (mean ± SD) eggs/female (N = 33), while control mosquitoes produced 64 ± 23 eggs/female (N = 32). The difference was statistically significant (Mann-Whitney test, P<0.001). A second experimental replicate generated data with similar pattern showing significant reduction in egg numbers (Mann-Whitney test, P<0.01, Fig. S1).
An. gambiae Vg knockdown reduced egg production. (A) Mosquitoes treated with Cas13a/Vg-crRNA had lower egg counts as compared to the cohort treated with Cas13a/ctr-crRNA (P<0.001). (B) Reduction of Vg transcripts was confirmed by RT-PCR. NRT: No RT control; NTC: no template control. The rpS5 was used as a loading control.
Cas13a mediated COPI gene silencing in Aedes aegypti
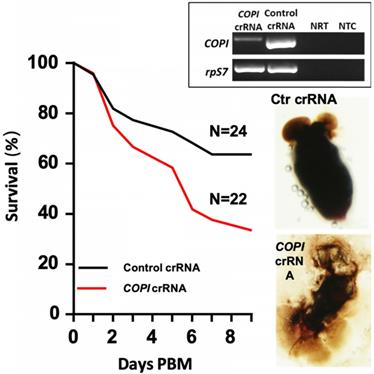
The coatomer complex I (COPI) proteins are involved in blood digestion in Ae. aegypti [39]. The COPI complex consists of α, β, β', γ, δ, ε and ζ subunits, encoded by separate genes. The production of COPI proteins is blood inducible between 18-36 hr post blood meal in fat body and 24-48 hr post blood meal in ovaries [39]. As shown by other investigators, dsRNA-mediated knockdown of the genes encoding all but the ε subunit led to blood meal-induced mortality [39]. Therefore, we targeted the COPI genes to test the gene silencing efficacy of the Cas13a machinery in Ae. aegypti. The Cas13a construct was injected into one-day-old mosquitoes (N = 90). At day 3 post construct delivery, the mosquitoes were given a blood meal. The engorged mosquitoes (N = 75) were split into two cohorts, one was injected with the crRNAs specific for α- and δ-COPI genes at 2 hr hours post blood meal (N = 27). The other cohort (N = 29) was injected with the control crRNA. A subset of mosquitoes (N = 5) in each cohort was sampled at 20 hr post injection to dissect the midguts to check their intactness. The carcasses from the same 5 mosquitoes were then used for RNA extraction and RT-PCR for knockdown validation. Reduction of abundance of the COPI transcripts was observed in the knockdown group (Fig. 4). The survival curves over 9 days post blood meal revealed a significantly lower survival of the COPI knockdown cohort (N = 22) than the control cohort (N = 24) (Fig. 4; Mantel-Cox test, P<0.001). It has previously been shown that COPI knockdown makes the mosquito midgut fragile [39]. Consistently, we observed that 5 out of 5 of midguts in the COPI knockdown mosquitoes were apt to break and leak during dissection, while the 5 of 5 midguts in the control mosquitoes were in good shape with intact blood bolus (Fig. 4). A second replicate experiment also showed a significant reduction in the survival of the COPI knockdown cohort (Mantel-Cox test, P<0.01, Fig. S2).
Phenotypes of Ae. aegypti COPI knockdown. Cas13a/COPI-crRNA resulted in mortality post blood meal (PBM) and fragile midgut. Reduction of COPI transcripts was confirmed by RT-PCR (insert panel). NRT: No RT control; NTC: no template control. The rpS7 was used as a loading control.
Cas13a mediated double gene knockdown
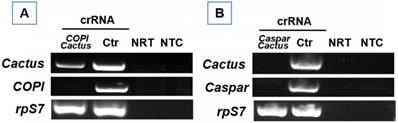
To determine the potential for silencing multiple genes, a cocktail of the Cas13a construct and crRNAs against the genes Cactus and Caspar or Cactus and COPI were prepared, respectively. Each cocktail was delivered into 20 one-day-old females of Ae. aegypti. The RNA samples were extracted and subject to RT-PCR for determining transcript abundance at day 6 post-delivery. As expected, the abundance of target gene transcripts was reduced by the respective treatment (Fig. 5), suggesting that multiple gene silencing is achievable and the Cas13a machinery can be effective for at least 6 days.
RT-PCR verification of double gene knockdown in Ae. aegypti that received a cocktail of Cas13a construct with (A) Cactus-and Caspar- or (B) Cactus- and COPI-crRNAs. NRT: No RT control; NTC: no template control. The rpS7 was used as a loading control.
Absence of detectable collateral cleavage of non-target RNA
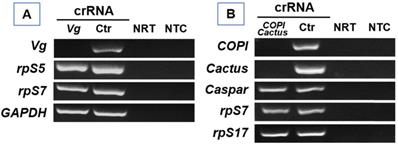
In bacteria, activated Cas13a displays cleavage activity of non-target RNA[10]. To determine if Cas13a has such collateral activity in mosquitoes, we examined the abundance of arbitrarily selected non-target transcripts in Cas13a/Vg crRNA treated An. gambiae as well as Cas13a/COPI-Cactus crRNA treated Ae. aegypti. As shown in Fig. 6, in An. gambiae, Cas13a/Vg crRNA treatment resulted in Vg knockdown, but non-target transcripts rpS5, rpS7 and GAPDH (glyceraldehyde 3-phosphate dehydrogenase) were not affected by the activated Cas13a. Likewise, in Ae. aegypti, COPI/Cactus crRNA activated Cas13a to co-silence COPI and Cactus, but did not affect non-target transcripts Caspar, rpS7 and rpS17 (Fig. 6). The data indicate that in mosquitoes the Cas13a may not execute collateral cleavage activity on non-target RNA.
RT-PCR of non-target transcripts. (A) In Vg silenced An. gambiae, the abundance of non-target transcripts rpS5, rpS7, and GAPDH was not affected by Cas13a/Vg crRNA. (B) In COPI and Cactus co-silenced Ae. aegypti, the abundance of non-target transcripts Caspar, rpS7, and rpS17 was not affected by Cas13a/COPI-Cactus crRNA. NRT: No RT control; NTC: no template control.
Discussion
In this proof of concept study, we demonstrate the effectiveness of CRISPRi mediated by CRISPR-Cas13a/crRNA machinery. The LwaCas13a was derived from L. wadei, and under control of Drosophila Actin promoter (Fig.1). The construct was delivered into the hemocoel of adult mosquitoes by intrathoracic injection, and Cas13a was transcribed constitutively (Fig. 2). Likely, the construct enters into the nucleus where the Cas13a gene is transcribed, and then the mRNA comes into the cytoplasm where the protein is synthesized. The construct remained active to transcribe Cas13a for at least 10 days post-delivery in Ae. aegypti (Fig. 2), which makes it temporally flexible to administer crRNAs targeting various genes that are expressed at different time points during a mosquito's life span. Target-specific crRNAs can be delivered either with the construct together or after the construct delivery at a time as appropriate to the experimental design. The system can silence highly abundant transcripts, as demonstrated by targeting Vg transcripts in An. gambiae (Fig. 3) and COPI transcripts in Ae. aegypti (Fig. 4). In both cases, the target genes are induced by blood meal to a high transcriptional level. In addition, we have tested the system on silencing genes Cactus and Caspar or Cactus and COPI simultaneously in Ae. aegypti. The co-knockdown activity can be effective at least for 6 days (Fig. 5). Taken together, these data conclusively demonstrate that the Cas13a-CRISPRi machinery is functional in An. gambiae and Ae. aegypti mosquitoes. This tool may work well in other mosquito species. We have not tested knockdown effect of the Cas13a system on genes that are mainly expressed in midgut, salivary glands and ovaries, which needs further studies.
The Cas13a-CRISPRi has certain advantages over the dCas9-CRISPRi for repressing gene expression [16, 40]. First, the dCas9-CRISPRi machinery acts at DNA level while Cas13a targets mRNA directly. RNA-guided binding of dCas9 to a specific promoter or coding sequence can block transcription. This mode of action is efficient in bacteria, but often is not very efficient in eukaryotic cells [16]. The dCas9 fusion proteins with a repressive domain have been developed for transcriptional repression, such as dCas9-KRAB (Krüppel associated box) in mammalian cells [15], but it is challenging to develop a fusion dCas9 with universal applicability. In addition, target specific sgRNA selection may be limited by the PAM that is required for the Cas9 activity. On the contrary, no PFS is required for target RNA cleavage by LwaCas13a in eukaryotic cells [23]. The mode of action of Cas13 is simple and programmable, and the efficacy has proven high in mammalian cells [11, 26] and in mosquitoes in the current study. RNAi mediated gene silencing is a very common practice in mosquito gene function studies. Recently, application versions have been developed for mosquito vector control [4, 5]. In these application cases, dsRNA is used to trigger RNAi machinery. In dsRNA mediated RNAi, the effective siRNA sequences sometimes are difficult to predict, therefore, a large dsRNA fragment is often used to increase chances to generate effective siRNA by Dicer. However, this strategy is accompanied with a higher chance to produce siRNA with off-target potentials [41, 42]. In addition, efficacy of dsRNA mediated RNAi varies case by case, and an optimal outcome is often a result of an empirical process [43]. In addition to the RNAi method, the CRISPRi approach mediated by RNA targeting Cas13 proteins offers an alternative for gene silencing in mosquitoes.
There is a concern about the potential of collateral cleavage with Cas13a, in which non-target RNA sequences can be cleaved by Cas13a in bacteria [10, 11]. Interestingly, this promiscuous RNA degradation activity was not observed in several studies in mammalian and plant cells [11, 26, 27]. These data have warranted its safety to be used as an effector to target against RNA viruses that infect humans [23]. However, a Cas13a/crRNA associated collateral cleavage was recently shown in human U87 glioblastoma cells [28]. In the study, exogenous gene GFP or EGFRVIII were overexpressed and targeted by Cas13a/crRNA. In this context, a partial degradation of ribosomal RNA profile was observed, and the abundance of non-target transcripts of GAPDH, HOTHAIR and L3MTL1 was reduced as well [28]. Furthermore, the RNA integrity was compared between the LN229 glioma cell line and HEK293T cells after treatment with the Cas13a/crRNA, the LN229 cells tended to be more sensitive to the collateral effect than the HEK293T cells [28]. In our study, we examined arbitrarily selected non-target transcripts, three in Vg silenced An. gambiae and three in COPI and Cactus co-silenced Ae. aegypti; no detectable reduction of these non-target mRNAs was observed (Fig. 6). The data suggest that collateral effect of Cas13a may not be a concern in mosquitoes, although we cannot completely rule out the possibility of collateral cleavage. Additional studies with large scale examination of non-target RNA would help confirm the absence of non-target cleavage by the Cas13/crRNA system in mosquitoes.
The Cas13a-CRISPRi system holds promise for robust and flexible programming to silence one or more genes simultaneously in mosquitoes, with potential applications in other arthropods.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
Authors thank Dr. Michelle Riehle in Medical College of Wisconsin for her insightful comments on the manuscript. This research is supported by an Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the National Institutes of Health under grant number P20GM103451, the National Institutes of Health SC1AI112786 and the National Science Foundation [No. 1633330]. The content is solely the responsibility of the authors.
Authorship Confirmation Statement
JX conceived the idea for the project and devised the study. JX, AK, KAH designed experiments, AK, WY, ASM, AP conducted experiments. AK, JX, KAH wrote manuscript. The authors confirm that all co-authors have reviewed and approved the manuscript. The authors affirm that the paper is original with unpublished findings, not under consideration by any other journals.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Lopez SBG, Guimaraes-Ribeiro V, Rodriguez JVG. et al. RNAi-based bioinsecticide for Aedes mosquito control. Sci Rep. 2019;9:4038
2. Whitten MM. Novel RNAi delivery systems in the control of medical and veterinary pests. Curr Opin Insect Sci. 2019;34:1-6
3. Zhang H, Li HC, Miao XX. Feasibility, limitation and possible solutions of RNAi-based technology for insect pest control. Insect Sci. 2013;20:15-30
4. Airs PM, Bartholomay LC. RNA Interference for Mosquito and Mosquito-Borne Disease Control. Insects. 2017;8:1-21
5. Balakrishna Pillai A, Nagarajan U, Mitra A. et al. RNA interference in mosquito: understanding immune responses, double-stranded RNA delivery systems and potential applications in vector control. Insect Mol Biol. 2017;26:127-139
6. Chaverra-Rodriguez D, Macias VM, Hughes GL. et al. Targeted delivery of CRISPR-Cas9 ribonucleoprotein into arthropod ovaries for heritable germline gene editing. Nat Commun. 2018;9:3008
7. Li M, Bui M, Yang T. et al. Germline Cas9 expression yields highly efficient genome engineering in a major worldwide disease vector, Aedes aegypti. Proc Natl Acad Sci U S A. 2017;114:E10540-E10549
8. Kistler KE, Vosshall LB, Matthews BJ. Genome engineering with CRISPR-Cas9 in the mosquito Aedes aegypti. Cell Rep. 2015;11:51-60
9. Jinek M, Chylinski K, Fonfara I. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816-821
10. Abudayyeh OO, Gootenberg JS, Konermann S. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science. 2016;353:aaf5573
11. Abudayyeh OO, Gootenberg JS, Essletzbichler P. et al. RNA targeting with CRISPR-Cas13. Nature. 2017;550:280-284
12. Makarova KS, Wolf YI, Alkhnbashi OS. et al. An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol. 2015;13:722-736
13. Zhang F, Wen Y, Guo X. CRISPR/Cas9 for genome editing: progress, implications and challenges. Hum Mol Genet. 2014;23:R40-46
14. Doudna JA, Charpentier E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science. 2014;346:1258096
15. Gilbert LA, Larson MH, Morsut L. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell. 2013;154:442-451
16. Qi LS, Larson MH, Gilbert LA. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell. 2013;152:1173-1183
17. Bikard D, Jiang W, Samai P. et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Res. 2013;41:7429-7437
18. Shmakov S, Abudayyeh OO, Makarova KS. et al. Discovery and Functional Characterization of Diverse Class 2 CRISPR-Cas Systems. Mol Cell. 2015;60:385-397
19. Zetsche B, Gootenberg JS, Abudayyeh OO. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell. 2015;163:759-771
20. Aman R, Ali Z, Butt H. et al. RNA virus interference via CRISPR/Cas13a system in plants. Genome Biol. 2018;19:1-9
21. Ali Z, Mahas A, Mahfouz M. CRISPR/Cas13 as a Tool for RNA Interference. Trends Plant Sci. 2018;23:374-378
22. Terns MP. CRISPR-Based Technologies: Impact of RNA-Targeting Systems. Mol Cell. 2018;72:404-412
23. Freije CA, Myhrvold C, Boehm CK. et al. Programmable Inhibition and Detection of RNA Viruses Using Cas13. Mol Cell. 2019;76:826-837
24. Mahas A, Neal Stewart C Jr, Mahfouz MM. Harnessing CRISPR/Cas systems for programmable transcriptional and post-transcriptional regulation. Biotechnol Adv. 2018;36:295-310
25. Liu L, Li X, Ma J. et al. The Molecular Architecture for RNA-Guided RNA Cleavage by Cas13a. Cell. 2017;170:714-726 e710
26. Cox DBT, Gootenberg JS, Abudayyeh OO. et al. RNA editing with CRISPR-Cas13. Science. 2017;358:1019-1027
27. Konermann S, Lotfy P, Brideau NJ. et al. Transcriptome Engineering with RNA-Targeting Type VI-D CRISPR Effectors. Cell. 2018;173:665-676 e614
28. Wang Q, Liu X, Zhou J. et al. The CRISPR-Cas13a Gene-Editing System Induces Collateral Cleavage of RNA in Glioma Cells. Adv Sci (Weinh). 2019;6:1901299
29. Gootenberg JS, Abudayyeh OO, Lee JW. et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 2017;356:438-442
30. Pei Y, Lu M. Programmable RNA manipulation in living cells. Cell Mol Life Sci. 2019;76:4861-4867
31. Zhao X, Liu L, Lang J. et al. A CRISPR-Cas13a system for efficient and specific therapeutic targeting of mutant KRAS for pancreatic cancer treatment. Cancer Lett. 2018;431:171-181
32. Kandel Y, Vulcan J, Rodriguez SD. et al. Widespread insecticide resistance in Aedes aegypti L. from New Mexico, U.S.A. PLoS One. 2019;14:e0212693
33. Wang Y, Gilbreath TM 3rd, Kukutla P. et al. Dynamic gut microbiome across life history of the malaria mosquito Anopheles gambiae in Kenya. PLoS One. 2011;6:e24767
34. Jacobsen L, Calvin S, Lobenhofer E. Transcriptional effects of transfection: the potential for misinterpretation of gene expression data generated from transiently transfected cells. Biotechniques. 2009;47:617-624
35. Kennedy EM, Kornepati AV, Goldstein M. et al. Inactivation of the human papillomavirus E6 or E7 gene in cervical carcinoma cells by using a bacterial CRISPR/Cas RNA-guided endonuclease. J Virol. 2014;88:11965-11972
36. Bassett AR, Liu JL. CRISPR/Cas9 and genome editing in Drosophila. J Genet Genomics. 2014;41:7-19
37. Bassett AR, Tibbit C, Ponting CP. et al. Mutagenesis and homologous recombination in Drosophila cell lines using CRISPR/Cas9. Biol Open. 2014;3:42-49
38. CLEMENTSM AN, BOOCOCK R. Ovarian development in mosquitoes: Stages of growth and arrest, and follicular resorption. Physiological Entomology. 2008;9:1-8
39. Isoe J, Collins J, Badgandi H. et al. Defects in coatomer protein I (COPI) transport cause blood feeding-induced mortality in Yellow Fever mosquitoes. Proc Natl Acad Sci U S A. 2011;108:E211-217
40. Xu X, Qi LS. A CRISPR-dCas Toolbox for Genetic Engineering and Synthetic Biology. J Mol Biol. 2019;431:34-47
41. Naito Y, Yamada T, Matsumiya T. et al. dsCheck: highly sensitive off-target search software for double-stranded RNA-mediated RNA interference. Nucleic Acids Res. 2005;33:W589-591
42. Jackson AL, Bartz SR, Schelter J. et al. Expression profiling reveals off-target gene regulation by RNAi. Nat Biotechnol. 2003;21:635-637
43. Scott JG, Michel K, Bartholomay LC. et al. Towards the elements of successful insect RNAi. J Insect Physiol. 2013;59:1212-1221
Author contact
Corresponding author: Jiannong Xu, jxuedu, (575) 646-7713, Department of Biology, New Mexico State University, PO Box 30001 MSC 3AF, Las Cruces NM, 88003, USA

 Global reach, higher impact
Global reach, higher impact