ISSN: 1839-9940
J Genomics 2017; 5:58-63. doi:10.7150/jgen.20358 This volume Cite
Research Paper
CNV Analysis of Host Responses to Porcine Reproductive and Respiratory Syndrome Virus Infection
1. Animal Genomics and Improvement Laboratory, BARC, USDA-ARS, Beltsville, Maryland 20705, USA;
2. Animal Parasitic Diseases Laboratory, BARC East, USDA-ARS, Beltsville, Maryland 20705, USA.
3. Institute of Animal Science, Chinese Academy of Agricultural Science, Beijing, 100193, China;
4. College of Animal Science and Technology, Northwest A&F University, Shaanxi Key Laboratory of Agricultural Molecular Biology, Yangling, Shaanxi 712100, China.
5. Department of Diagnostic Medicine and Pathobiology, College of Veterinary Medicine, Kansas State University, Manhattan, KS 66506, USA.
* Co- first authors.
Abstract
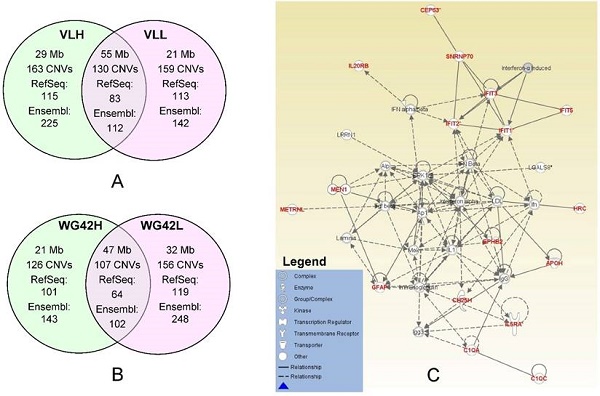
Porcine reproductive and respiratory syndrome (PRRS) is a devastating disease with a significant impact on the swine industry causing major economic losses. The objective of this study is to examine copy number variations (CNVs) associated with the group-specific host responses to PRRS virus infection. We performed a genome-wide CNV analysis using 660 animals genotyped with on the porcine SNP60 BeadChip and discovered 7097 CNVs and 271 CNV regions (CNVRs). For this study, we used two established traits related to host response to the virus, i.e. viral load (VL, area under the curve of log-transformed serum viremia from 0 to 21 days post infection) and weight gain (WG42 from 0 to 42 days post infection). To investigate the effects of CNVs on differential host responses to PRRS, we compared groups of animals with extreme high and low estimated breeding values (EBVs) for both traits using a case-control study design. For VL, we identified 163 CNVRs (84 Mb) from the high group and 159 CNVRs (76 Mb) from the low group. For WG42, we detected 126 (68 Mb) and 156 (79 Mb) CNVRs for high and low groups, respectively. Based on gene annotation within group-specific CNVRs, we performed network analyses and observed some potential candidate genes. Our results revealed these group-specific genes are involved in regulating innate and acquired immune response pathways. Specifically, molecules like interferons and interleukins are closely related to host responses to PRRS virus infection.
Keywords: PRRS, Copy Number Variation, SNP.

 Global reach, higher impact
Global reach, higher impact