ISSN: 1839-9940
J Genomics 2013; 1:5-12. doi:10.7150/jgen.3769 This volume Cite
Review
Cellular and Molecular Implications of Mature Adipocyte Dedifferentiation
1. College of Animal Science and Technology, Northwest A&F University, Yangling, Shaanxi Province 712100, China.
2. Department of Animal Sciences, Washington State University, Pullman, WA 99164, USA.
3. Department of Animal Sciences, Federal University of Viçosa, Viçosa, MG 3670-000, Brazil.
4. Department of Agricultural, Food and Nutritional Science, University of Alberta, Edmonton, Alberta T6G 2P5, Canada.
5. College of Animal Science and Technology, Nanjing Agricultural University, Nanjing 210095, China.
6. United States Department of Agriculture, Agriculture Research Services, Athens, GA 30605, USA.
* Contributed equally to this effort.
Published 2013-10-15
Abstract
There is a voluminous amount of scientific literature dealing with the involvement of adipocytes in molecular regulation of carcass composition, obesity, metabolic syndrome, or diabetes. To form adipocytes (process termed adipogenesis) nearly all scientific papers refer to the use of preadipocytes, adipofibroblasts, stromal vascular cells or adipogenic cell lines, and their differentiation to form lipid-assimilating cells containing storage triacylglyceride. However, mature adipocytes, themselves, possess ability to undergo dedifferentiation, form proliferative-competent progeny cells (the exact plasticity is unknown) and reinitiate formation of cells capable of lipid metabolism and storage. The progeny cells would make a viable (and alternative) cell system for the evaluation of cell ability to reestablish lipid assimilation, ability to differentially express genes (as compared to other adipogenic cells), and to form other types of cells (multi-lineage potential). Understanding the dedifferentiation process itself and/or dedifferentiated fat cells could contribute to our knowledge of normal growth processes, or to disease function. Indeed, the ability of progeny cells to form other cell types could turn-out to be important for processes of tissue reconstruction/engineering and may have implications in clinical, biochemical or molecular processes.
Keywords: mature adipocytes, dedifferentiation, review.
Introduction
As a consequence of cellular adipogenesis, lipid metabolism, gene and protein expression and adipokine secretion, mature adipocytes containing a single large lipid droplet were generally considered to be in the terminal stage of differentiation [1-4]. Recent studies, however, have indicated that mature adipocytes possess the ability to dedifferentiate into a population of proliferation-competent progeny cells known as adipofibroblasts or dedifferentiated fat (DFAT) cells [5-13]. This ill-defined phenomenon challenges the traditional view of the terminality of mature adipocytes [1, 14].
Mature adipocyte dedifferentiation
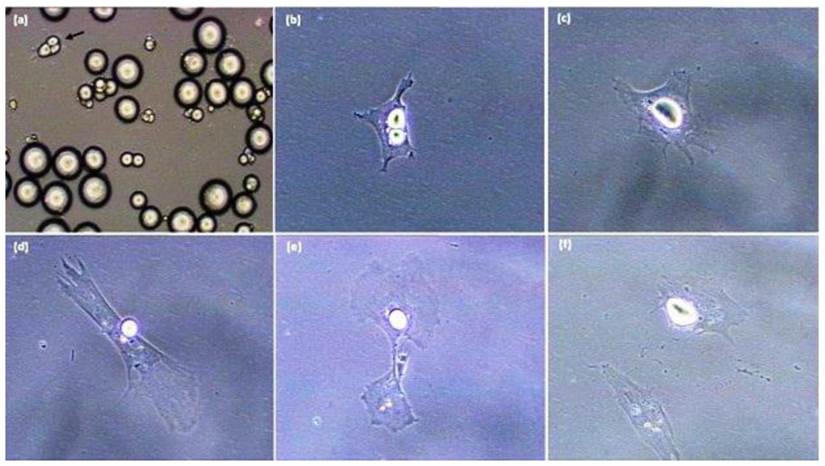
Adipocyte dedifferentiation is readily seen in vitro [5-10, 15-33]. The key to obtaining knowledge with progeny of mature adipocytes is to insure that initial cell cultures are pure, and do not possess contamination with spindle-shaped preadipocytes. In order to accomplish this, during the dedifferentiation process mature adipocytes attach to the surface of a culture flask (Figure 1) and change morphology from a round cell into an irregular shaped cell. Subsequently, the lipid droplet is sectioned into smaller droplets and the cell divides into two progeny cells processing different amounts of lipid [7, 15-17, 34]. Chen et al. (2009 a,b) [15,18] indicated that cultured mature pig-derived adipocytes expel lipid directly into the medium prior to becoming capable of cell division, rather than retaining lipids and transferring the lipid into progeny cells symmetrically or asymmetrically as do mature adipocytes from other animals [5,7, 19,20,35]. These results suggest that lipid-processing by individual cells may be different among species, while the cellular and molecular regulation of the dedifferentiation process remains unclear. Although dedifferentiation may result in cellular/molecular markers being expressed in the diametrically-opposite direction of traditional adipogenesis [21], these checkpoints have not been defined, even though the phenomenon of mature adipocytes forming proliferative-competent progeny cells was first reported over 30 years ago by Adebonojo (1975a,b) while performing research on human adipocytes [22, 23].
Sequence of pig-derived (mature) adipocyte dedifferentiation in vitro. A. Initial ceiling culture, B. Mature adipocyte following purification by differential plating methods. C-F. Sequence whereby the mature cell (in B) undergoes lipid reorganization/extrusion and divides with asymmetric cell division. All photomicrographs were taken with a Sony RGB digital camera married to a Nikon Diaphot phase contrast microscope and Image Pro Plus image analysis software [20 x magnification].
Are there other explanations about these cells? Could DFAT cells that have been described as originating from mature adipocytes actually be cells like fibroblasts that possess the potential for extreme levels of lipid metabolism [5, 7, 36]? Alternatively, could these cells be a completely new type of cell which presents both differentiation and proliferation abilities [5, 7, 36]?
While our studies with mature adipocyte dedifferentiation involved pure cultures of cells derived from large meat animals, similar cell systems from other animals have been established. To clarify the molecular mechanisms of mature adipocyte dedifferentiation, Ono et al., [24] using porcine cells and microarray analysis, identified differentially expressed genes during dedifferentiation.
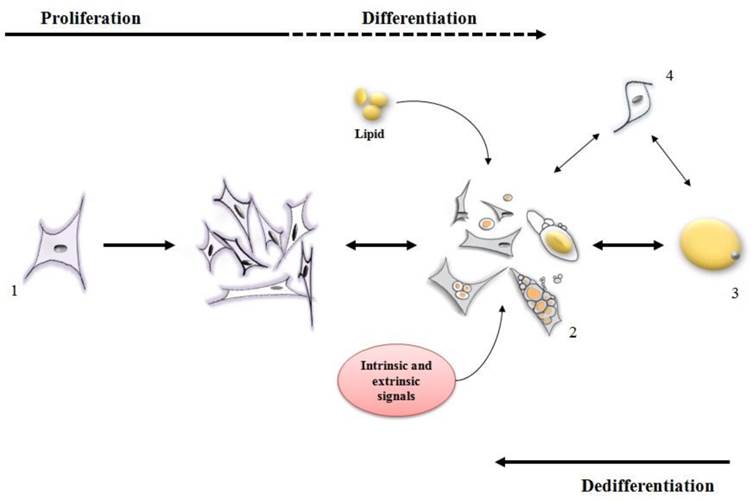
Ono et al. [24] reported that dedifferentiation resulted in 308 genes being down-regulated [many of which were involved in metabolism of lipid such as ADIPOQ, LIPE, PDK4, LPL, FASN, PPARG and FABP4]. Concomitantly, 368 genes were clearly up regulated during dedifferentiation [up regulated genes were those related to proliferation and differentiation, such as SERPINE1, TIMP1, PLAU, SFRP2, AEBP1, PRRX2, PEG10, IGFBP5 and ID2]. A variety of other studies are available to indicate that during dedifferentiation mature adipocytes lose their functional phenotype, reenter the cell cycle, gain a fibroblast-like appearance and eventually acquire a multipotent capacity for lineage differentiation [8-10,12,13,19] (Figure 2).
Stages of differentiation and dedifferentiation of fat cells. (1) adipofibroblasts, preadipocytes or stromal vascular cells proliferate until the cell population is expanded to the correct number. Under intrinsic and extrinsic signals, these cells (2) initiate the metabolism/accumulation of lipids where some of them can become a mature adipocyte (3) while some of them retain the capacity to lose lipid and return to a proliferative stage. Other types of cells (4) may also display both proliferative and lipid accumulative ability contributing for an increase in the mature adipocyte population.
DFAT cells as a model for adipogenesis
DFAT cells apparently possess a strong ability to proliferate (up to 30 passages) without detectable lipid deposition [25], which suggests the availability of DFAT cells could be used as a novel model for adipocyte differentiation and tissue formation. Whereas a variety of different cell culture systems have been used to address questions dealing with adipogenesis, including the use of lines of Swiss mouse embryo 3T3-L1 and stromal vascular (SV) cells [5], some significant differences between these cells and DFAT cells have been reported. Markers such as ANPEP, ITGB1, CD34, CD44 and THY1, ENG, MCAM, HLA-A, HLA-B, HLA-C were found on both SV and DFAT cells [28, 37-39]. However, the surface immuno-phenotype partially changes in different passages of adipose-derived SV cells. At the early passages (primary to 4th) of adipose-derived SV cells, the hematopoietic-associated markers (ITGAL, CD14, PTPRC, CD86 and HLA-DR) decreased and the MSCs(Mesenchymal stem cells)-associated markers (ANPEP, ITGB1, CD34, CD44, CD63, NT5E, THY1 and ALCAM) increased significantly [28,39]. Conversely, the phenotype of DFAT cells remained considerably stable from the early 2nd to as long as the 30th passage [25].
Cellular origin, homogeneity of cultures, whether the cells were previously transformed, repeatability of obtaining the same proportions of cells for culture, markers of adipogenesis and lipid metabolism, and viability of cells are all physiologies that must be considered when evaluating SV cells, cell lines and progeny from DFAT adipocytes [3,5-7,11-14,19,32,40]. For example, the doubling time of DFAT cells was approximately 18 h whereas the time for the 3T3-L1 cells was 22 h [26], DFAT cells apparently possess higher GPDH (Glycerol-3-phosphate dehydrogenase) activity than 3T3-L1 cells, and expression of SREBPF1 and LPL mRNA, which is related to lipid metabolism, has been detected during the growth phase in DFAT cells but not in 3T3-L1 pre-adipocytes [5]. The expression of PPARGC1A and CEBPA mRNA shows an earlier peak during differentiation in DFAT cells than in 3T3-L1 cells [26, 27]. Moreover, gene expression of re-differentiating DFAT progeny cells showed some differences to published reports using beef-derived SV cells [14], and unlike SV cells confluency is not required for DFAT cells to undergo differentiation in vitro [31]. Data from in vivo studies also suggest that DFAT cells possess greater ability to differentiate, not only into mature adipocytes, but also into vascular endothelial cells, forming a normal adipose tissue [27]. From these results, it could be inferred that DFAT cells could serve as an alternative model for studies of the mechanism of adipose tissue differentiation and formation for both in vitro and in vivo studies.
In meat animals, understanding variables regarding regulation, cellularity, growth and/or development of adipose tissue in different depots are important since fat deposition has an economic meaning in livestock production [19]. Different markers interact functionally to direct several regulatory process involved in adipogenesis or lipid metabolism, such as PPAR and SREBP [41, 42]. However, the regulatory compounds of adipogenesis are not necessarily similar between ruminants and monogastric animals [19]. It has been reported that fat deposition follows the order in which adipocyte number and volume is achieved firstly in the visceral adipose depot and lastly in the intramuscular depot [36,43]. Chen et al [16] have indicated that that PPARG and CEBPA mRNA were higher in DFAT cells from intramuscular than visceral adipose tissue derived from finishing pigs after differentiation, which means intramuscular DFAT cells may be more active in adipogenesis than visceral DFAT cells. This may imply that even if DFAT cells originated from the same animal, different depots present different rates of redifferentiation and molecular markers. Additional evidence to support this concept includes: high RNA abundance of FABP4 found in adipofibroblasts originated from dedifferentiation of bovine perimuscular mature adipocytes, suggesting that progeny preadipocytes may potentially have an ability to redifferentiate at early time point due to inherited characteristics of molecular components in the cytoplasm of progeny cells derived from the mature adipocytes, although shape of the cells remains fibroblast-like form [14]. Studies that provide knowledge of the dynamic of accumulation and metabolism of lipids may lead to the adoption of strategies in the livestock production in order to enhance fat deposition in specific (desired) depots [34, 35].
Multilineage potential of DFAT cells
A slightly different focus of DFAT cells studies is to understand the mechanisms that allow these cells to transdifferentiate in other types of cells [28-30]. Under specific in vitro conditions, beef-derived DFAT cells redifferentiate in a protracted manner, of which some cells contain lipid and some did not (only possessing the storage vesicles; [31]). DFAT cells are capable of transdifferentiating to other types of cells such as osteoblasts, chondrocytes, skeletal myocytes, smooth muscle cells, cardiomyocytes, vascular endothelial cells, and neural cells under appropriate culture conditions in vitro or in vivo [24]. A particularly elegant study found that RUNX2, SPP1, SP7, BGLAP, PTH1R and SOX9 were expressed in DFAT cells originating from mature adipocytes suggesting the osteogenic and chondrogenic potentials of these cells are enhanced [28]. Moreover, the transdifferentiation of DFAT cells into osteoblasts was observed in vivo by the implantation of collagen-based scaffolds [32]. Osteogenic differentiation of DFAT cells could also be induced by an analog of retinol which interacts with bone morphogenetic proteins to inhibit adipogenesis and enhance osteogenesis [32].
Although DFAT and SV cells share similarity with regard to multipotential lineage, a distinct aspect of the differentiation capacities between these cells exist in their angiogenesis behavior [25,44]. Previous studies have shown that adipose tissue-derived SV cells can differentiate into endothelial lineage and express endothelial markers and incorporate acetylated low density lipoprotein (Dil-ac-LDL) when agents such as fibroblast growth factor (FGF), epidermal growth factor (EGF) and vascular endothelial growth factor (VEGF) were applied [45-49]. On the other hand, DFAT cells with or without induction have negative expression of endothelial cell or progenitor markers such as such VWF, PECAM1 and CD34 [28].
So, do DFAT cells possess multipotent potential [11], or might the manipulation of the daughter cells simply be something manifested through cell culture manipulation [44, 50, 51]? Recently, it was demonstrated that the regulatory region of MYOD1, a master regulator for skeletal myogenesis, in DFAT cells shows a high degree of methylation [29]. Treatment of 5-azacytidine, a demethylating agent, led to the expression of MYOD1 and MYOG in DFAT cells, as well as the formation of multinucleated cells expressing MUTYH, although MYF5 was still absent after induction [29]. Another study found expression of specific cardiomyocyte-specific markers was observed in DFAT cells derived from mature adipocytes when co-cultured with neonatal cardiomyocytes, suggesting that DFAT cells can convert into cardiomyocyte phenotype under appropriate microenvironment [33]. Moreover, it was found that DFAT cells expressed cardiac phenotypic markers when grown on semisolid methylcellulose medium in the absence of cardiomyocytes, indicating that DFAT cells have the potential to spontaneously convert to cardiomyocyte phenotype [33]. Sakuma et al., [30] using immunohistochemical analysis revealed that more than 50% of the DFAT cells were successfully transformed into α-smooth muscle actin-positive cells under specific culture condition. The expressions of TAGLN2, α-smooth muscle actin and smooth muscle-myosin heavy chain were increased in DFAT cells during the first week of differentiation culture.
Collectively, these studies suggest that the DFAT cells have potential to differentiate into several lineages, including adipose cells, osteoblasts, chondrocytes, skeletal myocytes, smooth muscle cells, cardiomyocytes, endothelial cells and neuronal cells [44, 50]. Moreover, although much of the observed multilineage potential was dependent on environmental conditions, such as the type of culture media used [51] once cells begin to express a phenotype for a different tissue form they can be utilized for regimens of tissue reconstruction or tissue regeneration/engineering requiring those specific cells. Further studies with DFAT cells should yield a number of uses for clinical, biochemical, metabolism and molecular application.
Summary
Dedifferentiation of mature adipocytes, and proliferation/transdifferentiation abilities of their daughter progeny cells provide us with a cellular system for exploring adipocyte involvement in fat-related diseases or improving animal productivity. Moreover, DFAT cells could be used as a unique adipogenesis model, and new information may be gained on comparative differences among these cells and SV cells, and cell lines such as 3T3-L1 cells. Mature adipocytes that possess the ability to lose lipid and form proliferative-competent daughter cells may be (in fact) a different type of cell than has ever been categorized. Continued studies may uncover this potentially exciting possibility. Finally, the multipotential lineage of DFAT cells may lead to the use of these cells in tissue reconstruction/ engineering. Collectively, there are cellular and molecular implications of mature adipocyte dedifferentiation, including the potential uses in clinical, biochemical and molecular applications.
Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. Dodson MV, Mir PS, Hausman GJ, Guan LL, Du M, Jiang Z, Fernyhough ME, Bergen WG. Obesity, metabolic syndromes and adipocytes. Journal of Lipids. 2011;2011:721686
2. Kokta TA, Dodson MV Gertler A, Hill RA. Intercellular signaling between adipose tissue and muscle tissue. Domestic Animal Endocrinology. 2004;27:303-331
3. Dodson MV, Hausman GJ, Guan LL, Du M, Jiang Z. Potential impact of mature adipocyte dedifferentiation in terms of cell numbers. International Journal of Stem Cells. 2011;4:76-78
4. Du M, Dodson MV. Advanced techniques to enhance marbling in meat. Control of Meat Quality (Joo S, ed). 2011
5. Fernyhough ME, Helterline DI, Vierck JL, Hausman GJ, Hill RA, Dodson MV. Dedifferentiation of mature adipocytes to form adipofibroblasts: More than a possibility. Adipocytes. 2005;1(1):17-24
6. Dodson MV, Fernyhough ME, Vierck JL, Hausman GJ. Adipocytes may not be a terminally differentiated cell type: Implications for animal production. Animal Science. 2005;80(3):239-240
7. Fernyhough ME, Bucci L, Hausman GJ, Antonio J, Vierck JL, Dodson MV. Gaining a solid grip on adipogenesis. Tissue and Cell. 2005;37(4):335-338
8. Sugihara H, Funatsumaru S Yonemitsu N, Miyabara Toda S, Hikichi Y. A simple culture method of fat cells from mature fat tissue fragments. Journal of Lipid Research. 1989;30:1987-1995
9. Shigematsu M, Watanabe H, Sugihara H. Proliferation and differentiation of unilocular fat cells in the bone marrow. Cell Structure Function. 1999;24:89-100
10. Tholpady SS, Aojanepong C, Lull R, Jeong JH, Mason AC, Futrell JW, Ogle RC, Katz AJ. The cellular plasticity of human adipocytes. Annal of Plastic Surgery. 2005;54:651-656
11. Dodson MV, Fernyhough ME. Mature adipocytes: Are there still novel things that we can learn from them? Tissue and Cell. 2008;40:307-308
12. Vierck JL, McNamara JP Dodson MV. Proliferation and differentiation of progeny of ovine unilocular fat cells (adipofibroblasts). In Vitro Cellular and Development Biology-Animal. 1996;32:564-572
13. Vierck JL, McNamara JP, Dodson MV. Two alternative procedures to isolate adipofibroblasts from sheep skeletal muscle. Method in Cell Science. 1996;18:309-314
14. Taniguchi M, Guan le L, Zhang B, Dodson MV, Okine E, Moore SS. Adipogenesis of bovine perimuscular preadipocytes. Biochemical and Biophysical Research Communication. 2008;366:54-59
15. Chen J, Guridi M, Fernyhough ME, Jiang Z, Guan LL, Hausman GJ, Dodson MV. Initial differences in lipid processing leading to pig- and beef-derived mature adipocyte dedifferentiation. Basic and Applied Myology (European Journal of Translational Myology). 2009;19(5):243-246
16. Sugihara H, Yonemitsu N, Miyabara S, Toda S. Proliferation of unilocular fat cells in the primary culture. Journal of Lipid Research. 1987;28(9):1038-1045
17. Chen J, Dodson MV, Jiang Z. Cellular and molecular comparison of redifferentiation of intramuscular- and visceral-adipocyte derived progeny cells. International Journal of Biological Sciences. 2010;6(1):80-88
18. Chen J, Guridi M, Fernyhough ME, Jiang Z, Guan L, Hausman GJ, Dodson MV. Clonal Mature Adipocyte Production of Proliferative-competent Daughter Cells Requires Lipid Export Prior to Cell Division. International Journal of Stem Cells. 2009;2:76-79
19. Fernyhough ME, Vierck JL, Hausman GJ, Mir PS, Okine EK, Dodson MV. Primary adipocyte culture: adipocyte purification methods may lead to a new understanding of adipose tissue growth and development. Cytotechnology. 2004;46:163-172
20. Zhang HH, Kumar S Barnett AH, Eggo MC. Ceiling culture of mature human adipocytes: use in studies of adipocyte functions. Journal of Endocrinology. 2000;164:119-128
21. Fernyhough ME, Vierck JL, Dodson MV. Assessing a non-traditional view of adipogenesis: adipocyte dedifferentiation -mountains or molehills? Cells, Tissues, Organs. 2006;182(3-4):226-228
22. Adebonojo FO. Studies on human adipose cells in culture: relation of cell size and multiplication to donor age. Yale Journal of Biology and Medicine. 1975;48:9-16
23. Adebonojo FO. Monolayer cultures of disaggregated human adipocytes. In Vitro. 1975;11:50-54
24. Ono H, Oki Y, Bono H, Kano K. Gene expression profiling in multipotent DFAT cells derived from mature adipocytes. Biochemical Biophysical Research Communication. 2011;407(3):562-567
25. Shen J, Sugawara A, Yamashita J, Ogura H, Sato S. Dedifferentiatiated fat cells: and alternative source of adult multipotent cells from the adipose tissues. International Journal of Oral Science. 2011;3:117-124
26. Yagi K, Kondo D, Okazaki Y, Kano K. A novel preadipocyte cell line established from mouse adult mature adipocytes. Biochemical Biophysical Research Communications. 2004;321:967-974
27. Nobusue H, Endo T, Kano K. Establishment of a preadipocyte cell line derived from mature adipocytes of GFP transgenic mice and formation of adipose tissue. Cell Tissue Research. 2008;332:435-446
28. Matsumoto T, Kano K, Kondo D, Fukuda N, Iribe Y, Tanaka N, Matsubara Y, Sakuma T, Satomi A, Otaki M, Ryu J, Mugishima H. Mature adipocyte-derived dedifferentiated fat cells exhibit multilineage potential. Journal of Cell Physiology. 2008;215:210-222
29. Kazama T, Fujie M, Endo T, Kano K. Mature adipocytes-derived dedifferentiation fat cells can transdifferentiate into skeletal myocytes in vitro. Biochemical and Biophysical Research Communications. 2008;337:780-785
30. Sakuma T, Matsumoto T, Kano K, Fukuda N, Obinata D, Yamaguchi K, Yoshida T, Takahashi S, Mugishima H. Mature adipocyte derived, dedifferentiated fat cells can differentiate into smooth muscle-like cells and contribute to bladder tissue regeneration. Journal of Urology. 2009;182(1):355-365
31. Fernyhough ME, Hausman GJ, Dodson MV. Progeny from dedifferentiated bovine adipocytes display protracted adipogenesis. Cells, Tissues, Organs. 2008;188:359-372
32. Oki Y, Watanabe S, Endo T, Kano K. Mature adipocyte-derived dedifferentiated fat cells can trans-differentiate into osteoblasts in vitro and in vivo only by all-trans retinoic acid. Cell Structure Function. 2008;33(2):211-222
33. Jumabay M, Matsumoto T, Yokoyama S, Kano K, Kusumi Y, Masuko T, Mitsumata M, Saito S, Hirayama A, Mugishima H, Fukuda N. Dedifferentiated fat cells convert to cardiomyocyte phenotype and repair infarcted cardiac tissue in rats. Journal of Molecular and Cell Cardiology. 2009;47(5):565-575
34. Hausman GJ, Dodson MV, Ajuwon K, Azain M, Barnes K, Guan LL, Jiang Z, Poulos SP, Sainz RD, Smith S, Spurlock M, Novakofski J, Fernyhough ME, Bergen WG. Board Invited Re-view: The biology and regulation of preadipocytes and adipocytes in meat animals. Journal of Animal Science. 2009;87:1218-1246
35. Dodson MV, Jiang Z, Chen J, Hausman GJ, Guan LL, Novakofski J, Thompson DP, Lorenzen CL, Fernyhough ME, Mir PS, Reecy JM. Allied industry approaches to alter intramuscular fat content and composition in beef animals. Journal of Food Science. 2010;75(1):226-228
36. Allen CE. Cellularity of adipose tissue in meat animals. Federal Procedure. 1976;35(11):2302-2307
37. Katz AJ, Tholpady A, Tholpady SS, Shang H, Ogle RC. Cell surface and transcriptional characterization of human adipose-derived adherent stromal (hADAS) cells. Stem Cells. 2005;23:412-423
38. Gronthos S, Franklin DM, Leddy HA, Robey PG, Storms RW, Gimble JM. Surface protein characterization of human adipose tissue-derived stromal cells. Journal of Cellular Physiology. 2001;189(1):54-63
39. Schaffler A, Buchler C. Concise review: adipose tissue- derived stromal cells - basic and clinical implications for novel cell-based therapies. Stem Cells. 2007;25:818-827
40. Poulos SP, Dodson MV, Hausman GJ. Cell line models for differentiation: preadipocytes and adipocytes. Experimental Biology and Medicine. 2010;235(10):1185-1193
41. Ntambi JM, Kim YC. Adipocyte differentiation and gene expression. Journal of Nutrition. 2000;130:3122-3126
42. Shimano H. Sterol regulatory element-binding proteins (SREBPs): transcriptional regulators of lipid synthetic genes. Progress in Lipid Research. 2001;40:439-452
43. Sainz RD, Hasting E. Simulation of the development of adipose tissue in beef cattle. Mcnamara JP, France J, Beever DE, Editors. Modeling Nutrient Utilization in Farm Animals. 2000
44. Malgorzata WZ, Walenko K. Stem cells from adipose tissue. Cellular & Molecular Biology Letters. 2011;16:236-257
45. Urbich C, Dimmeler S. Cell biology of Leydig cells in the testis. Circulation Research. 2004;95:343-353
46. Gehling UM, Ergu¨n S, Schumacher U, Wagener C, Pantel K, Otte M, Schuch G, Schafhausen P, Mende T, Kilic N, Kluge K, Schäfer B, Hossfeld DK, Fiedler W. In vitro differentiation of endothelial cells from AC133-positive progenitor cells. Blood. 2000;95:3106-3112
47. Hebert TL, Wu X, Yu G, Goh BC, Halvorsen YC, Wang Z, Moro C, Gimble JM. Culture effects of epidermal growth factor (EGF) and basic fibroblast growth factor (bFGF) on cryopreserved human adipose-derived stromal/ stem cell proliferation and adipogenesis. Journal of Tissue Engineering and Regenerative Medicine. 2009;3:553-561
48. Froehlich H, Gulati R, Boilson B, Witt T, Harbuzariu A, Kleppe L, Dietz AB, Lerman A, Simari RD. Carotid repair using autologous adipose-derived endothelial cells. Stroke. 2009;40:1886-1891
49. Miranville A. Improvement of postnatal neovascularization by human adipose tissue-derived stem cells. Circulation. 2004;110:349-355
50. Tallone T, Realini C, Böhmler A, Kornfeld C, Vassalli G, Moccetti T, Bardelli S, Soldati G. Adult Human adipose tissue contains several types of multipotent cells. Journal of Cardiovascular Translational Research. 2011;4:200-210
51. Balwierz A, Czech U, Polus A, Filipkowski R. K, Mioduszewska B, Proszynski T, Kolodziejczyk P, Skrzeczynska-Moncznik J, Dudek W, Kaczmarek L, Kulig J, Pryjma J, Dembinska-Kiec A. Human adipose tissue stromal vascular fraction cells differentiate depending on distinct types of media. Cell Proliferation. 2008;41:441-459
Author Biography
Shengjuan Wei is a Ph.D. candidate in the college of Animal Science and Technology at Northwest A&F University and a visiting scholar in Animal Sciences Department at Washington State University. Her research focuses on biotechnical breeding, gene function and adipocyte development.
Marcio S. Duarte is a Ph.D. student in the Department of Animal Sciences at Federal University of Viçosa-Brazil. He has additional research training from USDA - Meat Animal Research Center and the Department of Animal Sciences at Washington State University. His research interests focus on regulation of skeletal muscle and adipose tissue development during the pre- and postnatal phase and the factors underlying meat tenderness.
Dr. Linsen Zan is a professor in the College of Animal Science and Technology at Northwest A&F University, and Director of the National Beef Cattle Improvement Center in China. His research interests focus on genetic improvement and germplasm innovation for beef and dairy cattle; high efficiency breeding and industrialization of beef and dairy cattle; regulation and control of animal growth and development and biotechnology breeding.
Dr. Min Du is an endowed (funded) professor in the Department of Animal Sciences at Washington State University. His research focuses on the physiological conditions regulating skeletal muscle growth and development, and mechanisms governing differentiation of mesenchymal stem cells into myocytes, adipocytes and fibroblasts during fetal development. He has been publishing actively and currently serving as reviewers for a number of funding agencies and journals.
Dr. Zhihua Jiang is an Associate Professor in the Animal Sciences Department at Washington State University. Since the early 1990's, Dr. Jiang's research has focused on comparative genome biology with aims at understanding how orthologous genes have evolved and shuffled during evolution. The overall objective of his research program is to develop comparative genomic tools and reagents for determining gene sequence, location, expression and function, thus advancing genome sciences and their applications in agriculture and biomedicine. Dr. Jiang has published more than 110 publications that were well cited in the field. He was editor of the book "Reproductive Genomics in Domestic Animals" Published by Wiley-Blackwell in 2010. Dr. Jiang has received eight US patent awards.
Dr. LeLuo Guan is an Assistant Professor in Bovine Functional Genomics with the Department of Agricultural, Food and Nutritional Science at the University of Alberta. Her research in bovine functional genomics involves establishing links between omics results and economically important traits in livestock species including elucidation of molecular mechanisms of livestock production traits using gene expression profiling (currently using a set of 24,000 oligos for high through-put DNA microarray); identification of association of rumen microbial structures with cattle economically important traits at the genetic and transcriptomic levels; characterization of genes involved in microbe-microbe/host-microbe interaction. Dr. Guan has over 10 years of experience in the area of environmental microbiology and molecular biology. Dr. Guan has extensive experience in the field of microbial ecology and the characterization of complex bacterial communities and bacterial interspecies interactions. Dr. Guan has been one of the key contributors to the full-length cDNA project in the International Bovine Genome Sequencing Project. She has extensive knowledge and experience in developing research approaches to study bovine functional genomics to link the omics and/or microbes to important traits in beef cattle. She also has an excellent track record for the commercialization of research results.
Dr. Jie Chen is a professor in the college of Animal Science and Technology at Nanjing Agricultural University in China. His research focuses on genetic improvement of the pig, and the molecular mechanisms underlying pig economical traits. For the last decade, Dr. Chen has studied the molecular regulation of pork (meat) quality, especially intramuscular fat content, and several molecular markers have been identified for marker-assisted selection of pork intramuscular fat content.
Dr. Gary J. Hausman attended Iowa State University and graduated with a major in Animal Science. He then enrolled in graduate school at the University of Wisconsin-Madison and completed the Ph. D. in meat science and muscle biology in 1977. Dr. Hausman has been a Research Physiologist in the Animal Physiology Research Unit since 1980 and is an Adjunct Professor of the Human Foods and Nutrition and Animal Science Departments and is on the Graduate Faculty at the University of Georgia (UGA). For over 30 years, Gary Hausman has added to our understanding of embryonic and postnatal preadipocyte/adipocyte development. Dr. Hausman has authored or co-authored over 200 scientific articles published in refereed journals,(receiving >4300 citations; h-index = 33) 119 abstracts, 10 book chapters miscellaneous publications and has given 23 invited presentations at scientific meetings. Dr. Hausman's national and international reputation is reflected in invitations to present and discuss research data at national and international meetings and requests to consult with colleagues in academia, in industry, and in other governmental institutions throughout the world. Dr. Hausman was the recipient of the Outstanding Young Animal Scientist of the American Society of Animal Science (ASAS)-Southern Section, the 1993 ASAS Animal Growth and Development Research Award and was named an ASAS Fellow in 2011.
Dr. Michael V. Dodson is a Professor in the Animal Sciences Department at Washington State University (WSU), and affiliate faculty of the Center for Integrated Biotechnology (WSU) and affinity member of the Washington Center for Muscle Biology (collective effort between WSU and the University of Washington). For nearly three decades Dr. Dodson has studied skeletal muscle-derived stem cells. This work has resulted in nearly 150 papers on new methods for satellite cell- and adipocyte-derived stem cell isolation, culture, analysis and interpretation. Dr. Dodson has supported team research for his entire career, including a variety of efforts related to the USDA-sponsored research projects: NC1131 Molecular mechanisms regulating skeletal muscle growth and differentiation and NCCC201 Regulation of adipose tissue accretion in meat-producing animals (NCCC210).
Corresponding authors: Dr. M.V. Dodson, E-mail: dodsonedu, Dr. L. Zan, E-mail: zanlscom.cn or Dr. J. Chen, E-mail: jiechenedu.cn.

 Global reach, higher impact
Global reach, higher impact