ISSN: 1839-9940
J Genomics 2024; 12:6-13. doi:10.7150/jgen.87156 This volume Cite
Research Paper
Whole genome sequencing of Streptomyces antnestii sp. nov. with a potency to become an industrial strain
1. Department of Botany, University of North Bengal, Raja Rammohanpur, Siliguri-734013, India.
2. Bioinformatics Facility, University of North Bengal, Raja Rammohanpur, Siliguri-734013, India.
3. Dept. of Molecular, Cellular and Biomedical Sciences, University of New Hampshire, Durham, NH 03824, USA.
4. Hubbard Center for Genomic Studies, University of New Hampshire, Durham, NH 03824 USA.
5. Leibniz Institute DSMZ - German Collection of Microorganisms and Cell Cultures: Braunschweig, Germany.
6. Midnapore College, Midnapore, West Bengal 721101, India.
7. Biswa Bangla Genome Center, University of North Bengal, Raja Rammohanpur, Siliguri-734013, India.
8. Present address: HM Clause, Davis, California, USA.
Received 2023-6-14; Accepted 2023-10-12; Published 2024-1-1
Abstract
Streptomyces Strain San01 is isolated from the soil of ant-nest found in the tea estate of Darjeeling, India. The morphology, biochemical, as well as the molecular characteristics, proved that San01 belonged to the genus Streptomyces. The average nucleotide identity (ANI) value between the genome sequence of the studied strain and its closest phylogenetic neighbors were very low and also could be distinguished from its closest neighbour with broad range of phenotypic data. The draft genome sequence of isolate San01 (NZ_RZYA00000000.1) was estimated to be 9.12 Mbp in size with 71.2% of GC content and it encompasses 39 biosynthetic gene clusters that emphasize the biotechnological potential of this isolate.Based on the phenotypic, genetic and genomic data, isolate San01 (=JCM 34633 = NCTC 14543) merits to be recognized as a type strain of a novel species and hereby propose the name Streptomyces antnestii sp. nov. Incidentally, this is the first report on Streptomyces genomes from Darjeeling, India.
Keywords: Streptomyces, Eastern Himalaya, Whole genome sequencing, ANI, MLSA
Introduction
Actinobacteria is a phylum of gram-positive, filamentous bacteria with high GC content that are distributed in a wider range of habitats including terrestrial and aquatic ecosystems [1-3]. This phylum is an important player in ecological, agricultural, medicinal, and industrial fields due to its efficient production of secondary metabolites [4-6]. Within the Actinobacteria, the genus Streptomyces of the family Streptomycetaceae [7] and order Streptomycetales [8], is highly valuable in both medical and industrial aspects as a major bio-resource for antibiotics and other natural products including antivirals, antifungals, antitumoral, anti-parasitic, etc. [9]. Streptomyces are mainly soil-dwelling saprophytes that have a genome size of 6-12 Mbp [10] and maintain a life cycle with three morphological forms: vegetative hyphae, aerial hyphae, and spores [9]. Ecologically, Streptomyces can degrade organic compounds, circulate nutrients, and play a pivotal role in maintaining biogeochemical cycles [11].
Darjeeling is part of the sub-Himalayan region in India, is renowned for its tea gardens [12]. This region has not been exploited for actinobacterial population studies and no previous report on actinobacterial isolation has been found from this region. Here, we report the draft genome sequence of the Streptomyces sp. strain San01 from an ant nest soil found in the tea garden of Darjeeling. The studied strain was further subjected to polyphasic and genomic analyses followed by construction of the 16S and the comparison of the ANI scores for delineation of species. The resultant data confirms its taxonomic status as new species. Therefore, we propose Streptomyces sp. San01 as novel species for which the name Streptomyces antnestii sp. nov. is proposed.
Materials and Methods
Isolation and Maintenance of the organism
Soil samples from the ant (Solenopsis sp.) nest were collected from Dhajea tea garden situated in Darjeeling (26.9079° N, 88.2230° E), West Bengal, India in April 2016. The collected soil samples were air-dried for three to four days at 28 ℃. Serial dilution of dried soil sample was carried out with sterile saline solution (0.85% NaCl in distilled water) [13] up to 10-6 dilution. It was followed by inoculation of 0.1 ml of each dilution onto the sterile growth media by standard spread plate technique. The following growth media were used for isolation: Starch nitrate agar [14], Inorganic Salts, Starch Agar and Nutrient agar media [15]. All media were amended with Fluconazole (25µg/ml) and Rifampicin (5µg/ml) before pouring, to minimize the fungal and bacterial contamination [16, 17]. The plates were incubated at 30 ºC for five days. Typical colony of actinobacteria was selected and purified by growing on the selective ISP4 medium and incubated for 7 days at 30 °C [18]. After two rounds of purification, the isolate was sub-cultured on ISP4 slants and preserved at 4°C for further use [19]. Storage of the culture for the long term was carried out at 20% glycerol at -20 °C.
Morphological, Cultural and Physiological Characterization
Cultural characteristics which include the parameters such as aerial and substrate mycelium formation along with diffusible pigment production were determined according to the International Streptomyces Project [20]. The isolate San01 was inoculated and incubated for 7 days at 30 °C on various growth media which includes International Streptomyces Project (ISP) Medium 1: Tryptone-yeast extract broth [21], ISP Medium 2: Yeast extract-malt extract agar [22], ISP Medium 3: Oatmeal agar, ISP Medium 4: Inorganic salts-starch agar [24], ISP Medium 5: Glycerol-asparagine agar [23], ISP Medium 6: Peptone-yeast extract iron agar [26], ISP Medium 7: Tyrosine agar [27] and Bennet agar [28] obtained from Hi-Media, Mumbai, India. The gram nature of the strain was confirmed by the gram staining reaction [29]. Spore chain morphology of the isolate was studied using ISP4 media by coverslip method [30] following the International Streptomyces Project [20]. Cellular morphology of the strain was observed using a light microscope and scanning electron microscopy (SEM) [31].
The following biochemical tests were carried out: melanin formation (Shirling and Gottlieb 1966), xanthine, tyrosine and casein decomposition [32], esculin hydrolysis [33], citrate utilization [34], amylase production [35], cellulase production [36], gelatinase production [37], catalase production [38], Lipase production [39], Tween 80 degradation [40], Nitrate reduction test [41], Hydrogen sulphide production [25] and Carbohydrate utilization test [20].
Growth at different pH (5-10), temperature (4°C to 45°C) and NaCl (1 to 10%) tolerance was evaluated on Bennet agar medium. Antibiotic susceptibility was assayed on Bennet Agar (BA) medium by the Kirby-Bauer disc diffusion method [42] using the following antibiotics: Amikacin (30mcg), Ampicillin (10mcg), Amoxycillin (10mcg), Cefadroxil (30mcg), Cefoperazone (75mcg), Ceftazidime (30mcg), Chlorampheniol (30mcg), Ciprofloxacin (5 mcg), Cloxacillin (1 mcg), Cotrimoxazole (25mcg), Erythromycin (15mcg), Gentamicin (10mcg), Nalidixic Acid (10mcg), Netillin (10mcg), Nitrofurantoin (300mcg), Norfloxacin (10mcg), Penicillin (10 units), Tobramycin (10mcg) and Vancomycin (30mcg). Antibiotic discs (Hi-Media Pvt. Ltd., Mumbai, India) impregnated with antibiotics of known amount were placed on the surface of San01 inoculated plates maintaining aseptic condition and incubated at 30 °C for 5 days.
MALDI-TOF and FAME analysis
Microbial identification of isolate San01 was performed using MALDI-TOF MS [43]. Fatty acid Methyl esters (FAME) analysis was carried out for the identification of the bacterial species. The isolate Streptomyces sp. San01 cultivated in Inorganic Salt Starch Agar medium (ISP4) for 7 days at 30°C, fatty acids extracted and methylated followed by analysis by Microbial Identification (MIDI) system and the resultant peaks identified using the actino6 database [44].
Genome Characterization and phylogenetic studies
The genomic DNA of isolate San01 was extracted and purified by CTAB method [30]. The 16S rDNA sequence was amplified by Polymerase Chain Reaction (PCR) using actino-specific primers: ACT235F (5'CGC GGC CTA TCA GCT TGT TG3') and ACT 878R (3'CCG TAC TCC CCA GGC GGG G5' [45]. The amplified 16S rRNA gene fragments of the Streptomyces strain were then sequenced. The sequence similarity was studied through NCBI BLAST program online and similarity-based searches via the ezBioCloud server. Sanger sequencing of the partial 16S rRNA helped us in genus-level identification before proceeding with whole-genome sequencing.
Whole-genome sequencing was performed at the Hubbard Center for Genome Studies (University of New Hampshire, Durham, NH) using Illumina sequencing technology [46]. Along with this, a 16S phylogeny was also generated. Average nucleotide identity (ANI) has considered as a potent genome-based criterion for establishing species identity among genetically related micro-organisms [47]. The ANI score was calculated by ANI calculator (http://enve-omics.ce.gatech.edu/).
CodonW software (http://codonw.sourceforge.net/) was used for the thorough codon usage analysis. This includes GC composition, Frequency of Optimal codons (Fop), Effective number of codons (Enc), Aromaticity, Hydrophobicity, Codon Adaptation Index (CAI) and Relative Synonymous Codon Usage (RSCU). DAMBE was used for CAI calculation [48]. Spearman's Rank correlation coefficient among considered codon usage indices was calculated through SPSS ver 26 software. Genome mining was carried out for the search of biosynthetic gene clusters responsible collectively for bioactive molecule production in the newly sequenced genome using the antiSMASH software version 5.1.0.
Results and Discussion
Morphological, Cultural and Physiological Characterization
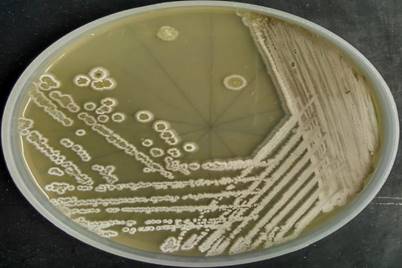
The colonies of Streptomyces sp. San01 are slow-growing, chalky with typical morphology of Streptomycetes [49] and sporulated well in ISP4 media (Fig. 1). The strain was gram-positive and filamentous and the spore chains were of flexibilis to looped type (Table 4). The growth of strain at various pH ranges proves the adaptability of the strain at both acidic and alkaline pH. The growth of the strain was observed at the temperature range of 21°C-37°C and showed salt tolerant ability with maximum growth at NaCl concentration of 7% (w/v).
Morphological characteristics of the strain San01- Colony morphology and aerial mycelium of strain San01 after incubation on ISP4 on 7 days.
Template table that can be used for the purpose of protologue description of a novel species.
| Name of a Genus | Streptomyces |
|---|---|
| Name of a Species | antnestii |
| Specific epithet | antnestii |
| Species status | sp.nov |
| New taxon description | Streptomyces antnestii (. ant nestii L. gen. n. antnestii from ant nest referring to the source of the organism) is aerobic, gram-stain positive, gelatinase-positive actinobacterium which forms an extensively branched substrate mycelium that bears aerial hyphae which is of flexibilis to looped type chains of spores in ISP4 media. The growth of strain at various pH ranges proves the adaptability of the strain at both acidic and alkaline pH. The growth of the strain was observed at temperature range of 21°C-37°C and showed salt tolerant ability with maximum growth at NaCl conc of 7% (w/v). The strains produce industrially significant enzymes cellulase, amylase, gelatinase, lipase and could utilize esculin. The culture isolate showed growth in presence of xanthine but could not utilize citrate and is catalase negative. Nitrate reduction and hydrogen sulfide production is absent. It utilized sugars, fructose, xylose, arabinose, raffinose, rhamnose, mannitol, glucose, sucrose, cellulose, galactose and lactose but showed no growth or only traces of growth with inositol. The predominant fatty acids are anteiso-C15:0, C16:0, iso-C16:0, anteiso-C17:1, C19:0. |
| Origin country | India |
| Origin region or where the strain belong to | Dhajea, Darjeeling, West Bengal, India |
| Isolation source | Soil |
| Date of Sampling | April 2016 |
| Location of site of Sampling | 26.9079° N, 88.2230° E |
| Accession number of 16S rRNA gene | MZ947234 |
| Accession number of Genome | NZ_RZYA00000000.1 |
| Status of Genome | Draft Genome |
| Size of Genome | 9.12 Mbp |
| G+C % | 71.2% |
| Strain numbers in study | 1 |
| Type strain designation | San01 |
| Collection numbers of Strains | JCM 34633, NCTC 14543 |
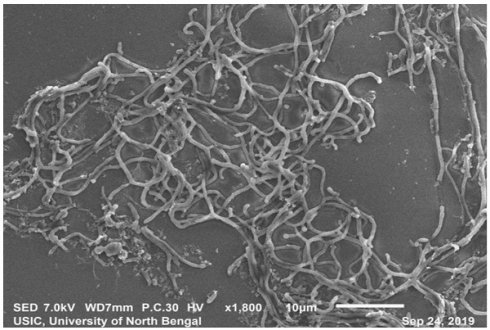
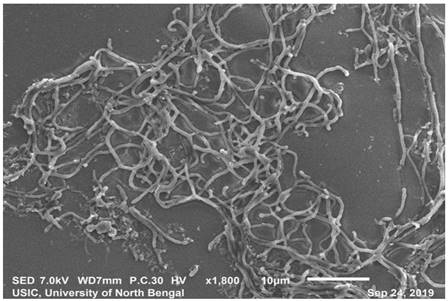
SEM analysis revealedthe strain was found to be filamentous with the breadth of the filament in the range of 0.455 µm to 0.645 µm. (Fig. 2). The strain San01 showed variation from S.aureus B7319T with Grey substrate mycelium color in contrast with Reddish orange substrate mycelium in the case of S. aureus and the spore chain arrangement found to be flexibilis to looped in San01 and Open looped in case of. S.aureus B7319T.The San01 produced pink pigment on oatmeal agar and S.aureus B7319T produced golden pigment. The San01 failed to utilize citrate but S. aureus B7319T utilized citrate. The results of the biochemical assays are summarized in Table 1. The isolate showed growth in all the media with good growth observed in Medium: ISP 1, ISP 2, ISP 3, ISP 4, ISP 7 and Bennet agar. The diffusible pink pigment production in ISP 3, ISP 4 and Bennet Agar was observed (Supplementary Table 1). Isolate San01 grew well over a broad range of pH (acidic to alkaline) values and showed optimum growth at 21°C to 37°C temperature range. The isolate was halo-tolerant, exhibiting a maximum growth at 7% (w/v) NaCl. (Supplementary Table 2a, 2b and 2c). The isolate was resistant to Ampicillin (10 mcg), Amoxicillin (10 mcg), Ceftazidime (30mcg), Cloxacillin (1 mcg), Cotrimoxazole (25mcg), Norfloxacin (10mcg), Penicillin (10 units) (Supplementary Table 3). The strains produce industrially significant enzymes Cellulase, Amylase, Gelatinase, and Lipase and could utilize Esculin. The culture isolate showed growth in the presence of Xanthine but could not utilize Citrate and is Catalase negative. Nitrate reduction and Hydrogen sulfide production is absent. It could utilize sugars, fructose, xylose, arabinose, raffinose, rhamnose, mannitol, glucose, sucrose, cellulose, galactose, and lactose but showed no growth or only traces of growth with inositol.
Scanning electron microscopy of San01 after 5 days of incubation in Bennet broth.
Comparison of the biochemical and morphological characteristics of isolate San 01 with the S.aureus B7319 T (data taken from Atlan et al. 2000 and Manfio et al. 2003).
| Characteristics Studied | Isolate San01 | S.aureus B7319T |
|---|---|---|
| Morphology and pigmentation Aerial spore mass colour on oatmeal agar (ISP 3) | Grey | Grey |
| Pigments on oatmeal agar | Pink | golden |
| Substrate mycelium colour on oatmeal agar | Grey | Reddish orange |
| Substrate mycelium colour on Bennett's | Off White to slightly pinkish and greyish | Reddish orange |
| Substrate mycelium colour on glycerol asparagine | Off White | Reddish orange |
| Substrate mycelium colour on inorganic salts - starch | grey | Reddish orange |
| Substrate mycelium colour on yeast extract - malt extract agars. | Off White | Reddish orange |
| Spore-chain arrangement | flexibilis to looped | Open looped |
| Melanin pigments production on peptone yeast extract iron agar | - | - |
| Pigment on inorganic salt starch agar | Pink | Orange yellow |
| Degradation of: | ||
| Starch | + | ND |
| Xanthine | + | + |
| Casein | + | ND |
| Tyrosine | + | ND |
| Gelatin | + | ND |
| Esculin | + | ND |
| Tween 80 | + | ND |
| tributyrin | + | ND |
| Nitrate reduction test | - | + |
| Hydrogen sulphide production | - | ND |
| Growth on sole carbon source | ||
| Arabinose | + | ND |
| Dextrin | + | ND |
| Fructose | + | ND |
| Meso-Inositol | + | + |
| Sucrose | + | ND |
| Xylose | No growth or only traces of growth | ND |
| Glucose | + | ND |
| Mannitol | + | + |
| Rhamnose | + | ND |
| Cellulose | + | ND |
| Raffinose | + | ND |
| Galactose | + | ND |
| Lactose | + | ND |
| Utilization of citrate | - | + |
| Growth at 7% NaCl | + | + |
| Temperature range for growth | 21 to 37 | 10 to 35 |
+=positive, - =negative, w =weakly positive,ND=no data available
MALDI-TOF and FAME analysis
MALDI-TOF is one of the analytical techniques with good accuracy for identifying microbes. It revealed the best match with Streptomyces hirsuitus with a similarity score 1.717. This score indicates that the isolated strain is a candidate type strain for novel species. FAME analysis using MIDI Sherlock has given a similarity index 0.0 suggests that there is no species in the database that could be identified with isolate San01 (Supplementary Table 4). The major fatty acids of isolate San01 (>15%) was iso-C16:0 while the type strain of S.S.aureus 41785Tareiso-C16:0 and anteiso-C15:0 as [50] shown in Supplementary Table 4.
Genome Characterization and phylogenetic studies
The whole genome sequencing of the isolate revealed that the number of contigs in the final draft assembly of Streptomyces sp. San01 is 524 with an N50 contig size of 314,067, a total sequence length of 9,124,249 bps, an average coverage of 280x, and a GC content of 71.2%. From the genome sequence of San01 the complete 16S rRNA (1531 bp) gene sequence extracted and were submitted to NCBI with the accession number MZ947234. The whole genome sequences have also been submitted to NCBI with the accession number NZ_RZYA00000000.1.
The annotation of the assembled Streptomyces sp. San01 genome was done via the NCBI Prokaryotic Genome Annotation Pipeline (PGAP) [51], which resulted in 7,693 protein-coding genes, 68 tRNA, 10 rRNA (five 5S rRNA, two 16S rRNA and three 23S rRNA), 3 non-coding RNA and 298 pseudogenes. This strain contained 7693 protein-coding genes, 68 tRNA, 10 rRNA (five 5S rRNA, two 16S rRNA and three 23S rRNA), 3 non-coding RNA and 298 pseudogenes. A comparison of the feature annotations produced by the PGAP of Streptomyces sp San01 (NZ_RZYA00000000.1.) and the publicly available Streptomycesaureus strain NRRL B- 2808 [52] (Genebank accession no of LIPQ00000000.1) genome annotations can be seen in Table 2. In the codon usage analysis of the newly sequenced genome of Streptomyces sp. San 01 it was revealed that 91.8% GC3 count indicated a stringent effect of compositional constrain on this genome. Further, C3 was predominating over G3. The parameters GC, GC3, CAI and Fop were positively correlated with each other with P<0.01 (Table 3).
Comparison of the genomic data between Streptomyces sp. San01 and S.aureus NRRL B-2808
| Genomic data | Streptomyces sp. San01 | Streptomyces aureus strain NRRL B-2808 |
|---|---|---|
| Size | 9.12 Mbp | 7.92 Mb |
| GC | 71.2% | 71.8% |
| Protein coding genes | 7,693 | 6769 |
| RNA | 81 | 79 |
| tRNA | 68 | 72 |
| rRNA | 10 rRNA (five 5S rRNA, two 16S rRNA and three 23S rRNA) | 4 |
| Noncoding RNA | 3 | 3 |
| Pseudogenes | 298 | 529 |
Spearman rank correlation among different codon usage indices. The positive correlation among GC, GC3, CAI and Fop revealed the persistence of compositional constrain on the Streptomyces sp. San01. ** indicates P<0.01
| GC | GC3 | CAI | ENc | Fop | Aromo | |
|---|---|---|---|---|---|---|
| GC | 1.00 | 0.85** | 0.87** | -0.75** | 0.82** | 0.58 |
| GC3 | 1.00 | 0.79** | -0.78** | 0.85** | 0.49 | |
| CAI | 1.00 | -0.72** | 0.82** | 0.44 | ||
| ENc | 1.00 | -0.51 | 0.32 | |||
| Fop | 1.00 | 0.42 | ||||
| Aromo | 1.00 |
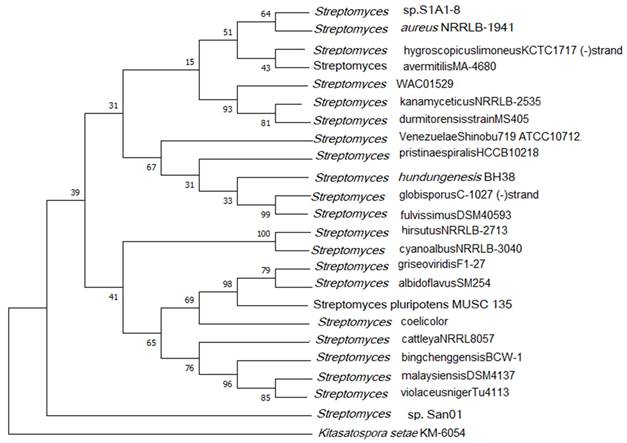
The similarity-based searches via ezbiocloud we found that San 01 showed high similarity percentage with Streptomyces aureus NRBC100912, Streptomyces dumitorensis MS405T and Streptomyces kanamyceticus NRBC 13414T. ANI is the similarity index between a pair of given genomes and a score of >95% is an indicator that both the genomes are from the same species [53]. Thus the ANI scores were calculated between genomes of San 01 and the above-mentioned strains and the ANI scores obtained were 82.50%, 83.67% respectively for Streptomyces aureus NRBC100912, and Streptomyces kanamyceticus NRBC 13414T. However, no genome sequence was found in the database for Streptomyces dumitorensis MS405T for ANI calculation. The 16S phylogeny was based on the NJ algorithm with 1000 bootstrap values and the out-group used was Kitasatospora setae (Fig 3). Based on this phylogeny it was observed that the studied strain San01 did not cluster with any of the formed groups indicating the uniqueness of this strain.
16S rRNA based phylogenetic tree.
Bioinformatic analysis revealed a total number of 123 CAZymes related genes for Streptomyces sp. San01. CAI (codon adaptation index) values of those genes grouped them under PHX (Potentially highly expressed) genes. This clearly explained the pivotal role of CAZymes related genes among the newly sequenced genome and thereby confirming the role of San01 in the production of important enzymes such as amylase and cellulase. The cluster analysis by antiSMASH software helped in exploring the bioactive potential of the strains [54-56]. San01 harbored 32 secondary metabolite biosynthetic gene clusters (BGCs) with more than 70% similarity (Supplementary Figure 1, Supplementary Table 5, and Supplementary Table 6). The presence of numerous BGCs with less similarity to the BGC database provided evidence for the potentiality of the strain to produce diverse secondary metabolites of importance.
The present study is the pioneer work, where the Darjeeling hills are being explored for the isolation of Streptomyces species. In this work, the actinobacteria strain isolated from ant nests of tea garden soils is identified as belonging to the genus Streptomyces based on morphological, biochemical, microscopic, and 16S rDNA as well as whole genome sequencing studies. The ability of the isolates to produce industrial enzymes like lipase, gelatinase, amylase, and cellulase makes them industrially important. The carbohydrate-activated enzymes (CAZymes) related genes within this genome support the role of the isolate in the production of amylase and cellulase. This emphasizes the importance of the isolate for the exploration of secondary metabolites for industrial applications in the future. The type strain San01 (JCM - 34633 and NCTC -14543) is isolated from the ant nest soil of the tea garden of Darjeeling Hills, India. The GenBank accession number of the assembled draft genome of strain San01 is NZ_RZYA00000000.1.
The Genbank accession number for the 16S rRNA gene sequence of strain San01 is MZ947234.
The Genebank accession number for the whole genome sequence is NZ_RZYA00000000.1.
Abbreviations
ISP: International Streptomyces Project; BA: Bennet Agar; SEM: Scanning Electron Microscopy; MALDI-TOF MS: Matrix-Assisted Laser Desorption Ionization Mass Spectrometry; FAME: Fatty acid Methyl esters; CAZymes: Carbohydrate- activated enzymes; NRPS: Nonribosomal peptide synthetase; PKS: Polyketide synthase (PKSs).
Supplementary Material
Supplementary tables.
Acknowledgements
This work was partially supported by USDA National Institute of Food and Agriculture Hatch 02281 (L.S.T.), National Institutes of Health 5R25GM125674 “Innovative Programs to Enhance Research Training (IPERT)” (W.K.T.), National Institutes of Health P20GM103506 IeDA program (W.K.T.), and the College of Life Science and Agriculture at the University of New Hampshire. Sequencing was performed on an Illumina HiSeq2500 instrument purchased with NSF MRI grant DBI-1229361 (to W.K.T.).
AS would like to acknowledge the Bioinformatics Facility, Department of Botany, and Biswa Bangla Genome Centre at the University of North Bengal for providing the facilities. We would like to acknowledge Dr. Tilak Saha, Department of Zoology, University of North Bengal, Darjeeling, India for helping us with Scanning Electron Microscopy.
Author contribution
AS and SC conceived the idea. SC collected the samples, isolated the DNA, and did all the wet lab works. JS, CP, WKT, LT did the whole genome sequencing of the organism. IS, SC, and GS did the Bioinformatics analysis of the genomic data. SC, SJ, GS, AS, IN, LT, CP, and IS prepared the manuscript. All authors have approved this version of the manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Amin DH, Abolmaaty A, Borsetto C. et al. In silico genomic mining reveals unexplored bioactive potential of rare actinobacteria isolated from Egyptian soil. Bull Natl Res Cent. 2019;43:1-9.doi.org /10.1186/s42269-019-0121y
2. Elbendary AA, Hessain AM, El-Hariri MD. et al. Isolation of antimicrobial producing actinobacteria from soil samples. Saudi J Biol Sci. 2018;25:44-46 doi.org/10.1016/j.sjbs
3. Rego A, Raio F, Martins TP. et al. Actinobacteria and cyanobacteria diversity in terrestrial antarctic microenvironments evaluated by culture-dependent and independent methods. Front Microbiol. 2019;10:1018. doi.org/10.3389/fmicb
4. Ferraiuolo SB, Cammarota M, Schiraldi C, Restaino OF. Streptomycetes as platform for biotechnological production processes of drugs. Appl Microbiol Biotechnol. 2021;105:551-568 doi.org/10.1007/s00253-020-11064-2
5. Olanrewaju OS, Babalola OO. Streptomyces: implications and interactions in plant growth promotion. Appl Microbiol Biotech. 2019;103:1179-1188 doi.org/10.1007/s00253-018-09577-y
6. Harir M, Bendif H, Bellahcene M. et al. Streptomyces secondary metabolites. Basic Biology and Applications of actinobacteria. 2018;6:99-122
7. Waksman SA, Henrici AT. The nomenclature and classification of the actinomycetes. J Bacteriol Res. 1943;46(4):337-41
8. Cavalier-Smith T. The phagotrophic origin of eukaryotes and phylogenetic classification of Protozoa. Int J Syst Evol. Microbiol. 2002;52(2):297-354
9. Jones SE, Elliot MA. Streptomyces exploration: competition, volatile communication and new bacterial behaviours. Trends Microbiol. 2017;25:522-531 doi.org/10.1016/j.tim.2017.02.001
10. Tidjani AR, Lorenzi JN, Toussaint M. et al. Massive gene flux drives genome diversity between sympatric Streptomyces conspecifics. MBi. 2019;10:e01533-19 doi.org/10.1128/mBio.01533-19
11. Siupka P, Piński A, Babicka D, Piotrowska-Seget Z. Genome Mining Revealed a High Biosynthetic Potential for Antifungal Streptomyces sp. S-2 Isolated from Black Soot. Int J Mol Sci. 2020;21:2558.doi.org /10.3390/ijms21072558
12. Sharma N, Phan HT, Chikae M. et al. Black tea polyphenol theaflavin as promising antioxidant and potential copper chelator. J Sci Food Agric. 2020;100(7):3126-3135
13. Taddei A, Rodríguez MJ, Márquez-Vilchez E, Castelli C. Isolation and identification of Streptomyces spp. from Venezuelan soils: Morphological and biochemical studies. I Microbiol Res. 2006;161:222-231 doi.org/10.1016/j.micres.2005.08.004
14. Korayem AS, Abdelhafez AA, Zaki MM, Saleh EA. Optimization of biosurfactant production by Streptomyces isolated from Egyptian arid soil using Plackett-Burman design. Ann Agric Sci. 2015;60:209-17 doi.org/10.1016/j.aoas
15. Singh V, Haque S, Singh H. et al. screening, and identification of novel isolates of actinomycetes from India for antimicrobial applications. Front Microbiol. 2016;7:1921. doi.org/10.3389/fmicb.2016.01921
16. Rao KV, Rao TR. Molecular characterization and its antioxidant activity of a newly isolated Streptomyces coelicoflavus BC 01 from mangrove soil. J Young Pharm. 2013;5:121-126 doi.org/10.1016/j.jyp
17. Dholakiya RN, Kumar R, Mishra A, Mody KH, Jha B. Antibacterial and antioxidant activities of novel actinobacteria strain isolated from Gulf of Khambhat, Gujarat. Front Microbiol. 2017;8:2420. doi.org/10.3389/fmicb.2017.02420
18. Rashad FM, Fathy HM, El-Zayat AS, Elghonaimy AM. Isolation and characterization of multifunctional Streptomyces species with antimicrobial, nematicidal and phytohormone activities from marine environments in Egypt. Microbiol Res. 2015;175:34-47 doi.org/10.1016/j.micres.2015.03.002
19. Shepherd MD, Kharel MK, Bosserman MA, Rohr J. Laboratory maintenance of Streptomyces species. Curr Protoc Microbiol. 2010;18:10E-1 https://doi.org/10.1002/9780471729259.mc10e01s18
20. Shirling ET, Gottlieb D. Methods for characterization of Streptomyces species1. Int J Syst Evol Microbiol. 1966;16:313-340
21. Pridham TG, Gottlieb D. The utilization of carbon compounds by some Actinomycetales as an aid for species determination. J Bacteriol. 1948;56:107
22. Pridham TG, Anderson P, Foley C. et al. selection of media for maintenance and taxonomic study of Streptomycetes. Antibiotics Annual. 1957;1:947-953
23. Pridham TG, Lyons Jr AJ. Streptomyces albus (Rossi-Doria) Waksman ET Henrici: taxonomic study of strains labeled Streptomyces albus. J Bacteriol. 1961;8:431
24. Kuster E. Outline of a comparative study of criteria used in characterization of the actinomycetes. Int J Syst and Evol Microbiol. 1959;9:97-104
25. Kuster E, Williams ST. Production of hydrogen sulfide by Streptomycetes and methods for its detection Appl Microbiol. 1964; 12: 46-52 https://doi.org/10.1128/am.
26. Tresner HD, Danga F. Hydrogen sulfide production by Streptomyces as a criterion for species differentiation. J Bacteriol. 1958;76:239-244
27. Shinobu R. Physiological and cultural study for the identification of soil actinomycetes species. Osaka University. 1958
28. Jones KL. Fresh isolates of actinomycetes in which the presence of sporogenous aerial mycelia is a fluctuating characteristic. J Bacteriol. 1949;57:141-145
29. Hucker George J, Conn JH. Methods of Gram staining. New work agriculture experimental station. 1923;93:1-37
30. Williams ST, Cross T. Chapter XI actinomycetes. In methods in microbiology. Academic Press. 1971;4:295-334 )
31. Kumar V. et al. Scanning electron microscopy of Streptomyces without use of any chemical fixatives. Scanning. 2011;33(6):446-449
32. Berd D. Laboratory identification of clinically important aerobic actinomycetes. Appl Microbiol. 1973;25:665-681 doi.org/10.1128/am.25.4.665-681
33. Gordon RE, Barnett DA, Handerhan JE, Pang CH. Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Evol Microbiol. 1974;24(1):54-63
34. Simmons JS. A culture medium for differentiating organisms of typhoid-colon aerogenes groups and for isolation of certain fungi. J Infect Dis. 1926;39:209-214
35. Gopinath SC, Anbu P, Arshad MM. et al. Biotechnological processes in microbial amylase production. BioMed research international. 2017:1-3
36. Kasana RC, Salwan R, Dhar H. et al. A rapid and easy method for the detection of microbial cellulases on agar plates using Gram's iodine. Curr Microbiol. 2008;57(5):503-507
37. Smith Jr HL, Goodner K. Detection of bacterial gelatinases by gelatin-agar plate methods. J Bacteriol Res. 1958;76(6):662-665
38. Reiner K. Catalase test protocol. American society for microbiology. 2010:1-6
39. Aly MM, Al-Shareef HA, Quri HA, Al-seeni MN. Spore forming bacterium from oil contaminated soil as a source of a lipase enzyme with exogeneous lipolytic activity. IOSR J Pharm Biol Sci. 2017;12(1):105-14
40. Kumar A, Kanwar R, Mehta SK. Development of Phosphatidylcholine/Tween 80 based biocompatible clove oil-in-water nanoemulsion as a green nanocarrier for controlled herbicide delivery. Environ Pollut. 2022;293:118558
41. Maciejewska M, Adam D, Naome A. et al. Assessment of the potential role of Streptomyces in cave moonmilk formation. Front Microbiol. 2017;8:1181. doi.org/10.3389/fmicb.2017.01181
42. Bauer AW. Antibiotic susceptibility testing by a standardized single disc method. Am J clin pathol. 1966;45:149-158
43. Rahi P, Vaishampayan P. MALDI-TOF MS application in microbial ecology studies. Front Microbiol. 2020;10:2954
44. Sasser M. Identification of Bacteria by Gas Chromatography of Cellular Fatty Acids. MIDI Inc Technical Notes. 1990;101:1-7
45. Stach JE, Maldonado LA, Ward AC, Goodfellow M, Bull AT. New primers for the class Actinobacteria: application to marine and terrestrial environments. Environ Microbiol. 2003;5:828-841 doi.org/10.1046/j.1462-2920.2003.00483.x
46. Bennett S. Solexa ltd. Pharmacogenomic. 2004;5:433-438
47. Li X, Tambong J, Yuan KX. et al. Re-classification of Clavibacter michiganensis subspecies on the basis of whole genome and multi-locus sequence analyses. Int J Syst Evol Microbiol. 2018;68:234-240.doi.org /10.1099%2Fijsem.0.002492
48. Xia X, Xie Z DAMBE. software package for data analysis in molecular biology and evolution. J Hered. 2001;92:371-373 doi.org/10.1093/jhered/92.4.371
49. Locci R. Streptomyces and related genera. Bergey's manual of systematic bacteriology. 1989;4:2451-2508
50. Cortés Albayay CJ. Bioprospection and peptidogenomics of novel Streptomyces species from Salar de Huasco. 2019.
51. Tatusova T, DiCuccio M, Badretdin A. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44:6614-6624 doi.org/10.1093/nar/gkw569
52. Manfio GP, Atalan E, Zakrzewska-Czerwinska. et al. Classification of novel soil Streptomycetes as Streptomyces aureus sp. nov, Streptomyces laceyi sp. nov. and Streptomyces sanglieri sp. nov. Anton Leeuw Int. 2003;83:245-255
53. Figueras MJ, Beaz-Hidalgo R, Hossain MJ, Liles MR. Taxonomic affiliation of new genomes should be verified using average nucleotide identity and multilocus phylogenetic analysis. Genome announc. 2014;2:e00927-14
54. Medema MH, Blin K, Cimermancic P. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 2011;39:339-346.doi.org /10.1093/nar/gkr466
55. Weber T, Blin K, Duddela S, Krug D. et al. antiSMASH 3.0—a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 2015;43:237-243 doi.org/10.1093/nar/gkv437
56. Jakubiec-Krzesniak K, Rajnisz-Mateusiak A. et al. Secondary metabolites of actinomycetes and their antibacterial, antifungal and antiviral properties. Pol J Microbiol. 2018;67:259. doi.org/10.21307%2Fpjm
Author contact
Corresponding author: Arnab.nbucom.

 Global reach, higher impact
Global reach, higher impact