ISSN: 1839-9940
J Genomics 2023; 11:37-39. doi:10.7150/jgen.83521 This volume Cite
Short Research Paper
Complete Genome Sequence of Floricoccus penangensis ML061-4 Isolated from Assam Tea Leaf [Camellia sinensis var. assamica (J.W.Mast.) Kitam.]
1. Department of Biology, Faculty of Science, Chiang Mai University, Chiang Mai, Thailand.
2. Graduate School, Chiang Mai University, Chiang Mai, Thailand.
3. Community Development Department, Ministry of Interior, Bangkok, Thailand.
4. School of Microbiology, University College Cork, Cork, Ireland.
5. APC Microbiome Ireland, University College Cork, Cork, Ireland.
6. Biological Sciences and ADAPT Research Centre, Munster Technological University, Cork, Ireland.
7. Darunsikkhalai School, King Mongkut's University of Technology Thonburi, Bangkok, Thailand.
8. Research Center in Bioresources for Agriculture, Industry and Medicine, Chiang Mai University, Chiang Mai, Thailand.
Received 2023-2-14; Accepted 2023-4-12; Published 2023-7-16
Abstract
Floricoccus penangensis is a Gram-positive coccoid organism that is a member of the lactic acid bacteria. F. penangensis ML061-4 was originally isolated from the surface of an Assam tea leaf, and its genome is herein shown to contain gene clusters predicted to be involved in complex carbohydrate metabolism and biosynthesis of secondary metabolites.
Floricoccus is a component genus of the lactic acid bacteria (LAB), and has been classified as a member of the family Streptococcaceae [1,2]. The genus Floricoccus was first described by Chuah et al. [1] and comprises two species i.e., Floricoccus penangensis and Floricoccus tropicus.
The Pacific Biosciences (PacBio) sequencing platform which is based on single-molecule real-time sequencing offers long read lengths and facilitates the improved assembly of genomes [3]. A previous study by Chuah et al. [1] presented the genome sequences of F. penangensis and F. tropicus strains which were determined using an Illumina sequencing platform.
F. penangensis ML061-4 was originally isolated from the surface of an Assam tea leaf in the Sakat sub-district of Pua district, Nan province, Thailand (19°15′53.62″N, 101°0′30.22″E). Briefly, four square centimeters of the fresh leaf was swabbed using a sterile cotton swab moistened with 0.85% (v/w) NaCl (Sigma-Aldrich, MO, U.S.A.), which was then plated by streaking on tryptic soy agar (TSA; Merck, Darmstadt, Germany). The plate was incubated at 37°C for 24 h. A single colony of this strain was then re-streaked on TSA to obtain a pure culture [4]. ML061-4 was routinely cultivated in M17 broth (Oxoid, Basingstoke, England) supplemented with 0.5% (w/v) glucose (GM17; Sigma-Aldrich) at 30°C for 24 h [5]. Genomic DNA of F. penangensis ML061-4 was prepared using a NucleoBond DNA extraction kit (Macherey-Nagel, Dueren, Germany), and genome sequencing was carried out using a Pacific Bioscience (PacBio) SMRT RSII sequencing platform (PacBio, Menlo Park, CA, United States). The library was prepared by the sequencing facility (Macrogen NGS Services, Seoul, South Korea) using the SMRTbell template prep kit 1.0, following the guide for the PacBio RS System and selecting for inserts of approximately 10-15 kb. Size selection was performed by the third-party sequencing provider using the BluePippin system. Approximately 56k subreads were obtained and the approximate subread N50 of these reads was 7.1 kb.
All quality controls including adapter removal, reads trimming, and quality control were performed by Macrogen using SMRT Analysis portal v2.3.0 (https://smrt-analysis.readthedocs.io/en/latest/SMRT-Analysis-Software-Installation-v2.3.0). Filtered subreads were assembled using the Hierarchical Genome Assembly Process (HGAP) pipeline with the method RS_Assembly.2 implemented in SMRT Analysis portal v2.3.0. Automatic annotation of predicted open reading frames (ORFs) was performed using NCBI's Prokaryotic Genome Annotation Pipeline (PGAP) v5.2 [6] to assign annotation, using an E-value cut-off of 0.0001 for hits showing at least 50% of similarity across at least 50% of the sequence length) against a non-redundant protein database provided by the National Centre for Biotechnology Information (NCBI) portal. Functional prediction of genes and proteins was integrated using the Clusters of Orthologous Groups (COGs) [7,8] and protein family (Pfam) [9], respectively, as previously described by Martín et al. [10]. Ribosomal RNA (rRNA) and transfer RNA (tRNA) genes were detected using RNAmmer v1.2 [11] and tRNA-scanSE v2.0 [12], respectively.
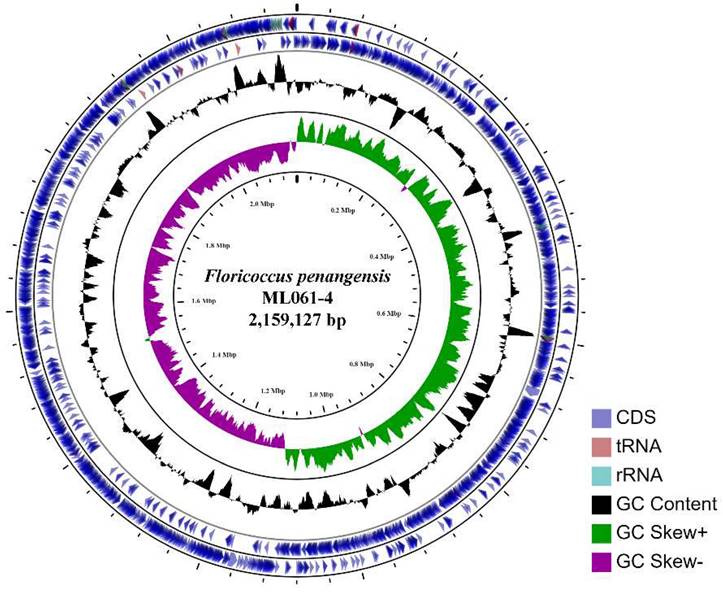
The assembled (at ~120× coverage) genome of F. penangensis ML061-4 is composed of a single contig of 2,159,127 bp, with a GC content of 33.2%, which is predicted to encode 2,134 genes. The circular genome map of ML061-4 is presented in Figure 1. The chromosome contains 19 rRNAs, 63 tRNAs, 3 ncRNAs, 16 pseudogenes and 2,049 protein-coding genes. The genome sequence was deposited in GenBank under accession number CP075561. Genome mapping of F. penangensis ML061-4 was evaluated using CGView online server v1.7 (http://cgview.ca/) [13].
Data availability
Genome sequence of F. penangensis ML061-4 was deposited in GenBank under accession number CP075561. The single-molecule real-time raw reads were deposited in SRA under the accession number SRX17727612.
The circular chromosome of F. penangensis ML061-4 using CGView. From outside to inside are the coding sequence (CDS, blue), tRNA (red), rRNA (blue-green), GC content (black) and GC skew (green/purple).
Acknowledgements
This research was funded by the National Research Council of Thailand (grant number PHD60I0089); the Graduate School; the Biology Department, Faculty of Science; the Research Center in Bioresources for Agriculture, Industry, and Medicine, Chiang Mai University, Thailand and the School of Microbiology; the APC Microbiome Ireland, University College Cork, Ireland through the financial support of Science Foundation Ireland under grant number 12/RC/2273-P2. F.B. is recipient of financial support from Science Foundation Ireland under Grant Agreement number 13/RC/2106_P2 at the ADAPT SFI Research Centre at Munster Technological University.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Chuah L-O, Yap K-P, Shamila-Syuhada AK. et al. Floricoccus tropicus gen. nov, sp. nov. and Floricoccus penangensis sp. nov. isolated from fresh flowers of durian tree and hibiscus. Int J Syst Evol Microbiol. 2017;67:4979-85
2. Rungsirivanich P, Inta A, Tragoolpua Y, Thongwai N. Partial rpoB gene sequencing identification and probiotic potential of Floricoccus penangensis ML061-4 isolated from Assam tea (Camellia sinensis var. assamica). Sci Rep. 2019;9:16561
3. Rhoads A, Au KF. PacBio sequencing and its applications. Genomics Proteomics Bioinformatics. 2015;13:278-89
4. Rungsirivanich P, Supandee W, Futui W, Chumsai-Na-Ayudhya V, Yodsombat C, Thongwai N. Culturable bacterial community on leaves of Assam tea (Camellia sinensis var. assamica) in Thailand and human probiotic potential of isolated Bacillus spp. Microorganisms. 2020;8:1585
5. Rungsirivanich P, Parlindungan E, O'Connor PM. et al. Simultaneous Production of Multiple Antimicrobial Compounds by Bacillus velezensis ML122-2 Isolated From Assam Tea Leaf [Camellia sinensis var. assamica (JW Mast.) Kitam.]. Front Microbiol. 2021;12:789362
6. Tatusova T, DiCuccio M, Badretdin A. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44:6614-24
7. Tatusov RL, Galperin MY, Natale DA, Koonin E V. The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000;28:33-6
8. Tatusov RL, Koonin E V, Lipman DJ. A genomic perspective on protein families. Science (80- ). 1997;278:631-7
9. Paysan-Lafosse T, Blum M, Chuguransky S. et al. InterPro in 2022. Nucleic Acids Res. 2023;51:D418-27
10. Martín R, Bottacini F, Egan M. et al. The infant-derived Bifidobacterium bifidum strain CNCM I-4319 strengthens gut functionality. Microorganisms. 2020;8:1313
11. Lagesen K, Hallin P, Rødland EA, Stærfeldt H-H, Rognes T, Ussery DW. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 2007;35:3100-8
12. Chan PP, Lowe TM. tRNAscan-SE: searching for tRNA genes in genomic sequences. In: Gene Prediction. Springer. 2019:1-14
13. Petkau A, Stuart-Edwards M, Stothard P, Van Domselaar G. Interactive microbial genome visualization with GView. Bioinformatics. 2010;26:3125-6
Author contact
![]() Corresponding authors: Narumol Thongwai, nthongwcom; Douwe van Sinderen, d.vansinderenie.
Corresponding authors: Narumol Thongwai, nthongwcom; Douwe van Sinderen, d.vansinderenie.

 Global reach, higher impact
Global reach, higher impact