ISSN: 1839-9940
J Genomics 2023; 11:1-8. doi:10.7150/jgen.77880 This volume Cite
Research Paper
Draft Genomes of Frankia strains AiPa1 and AiPs1 Retrieved from Soil with Monocultures of Picea abies or Pinus sylvestris using Alnus incana as Capture Plant
1. Université Claude-Bernard Lyon 1, Université de Lyon, UMR 5557 CNRS Ecologie Microbienne, Villeurbanne, France.
2. Texas State University, Department of Biology, 601 University Drive, San Marcos, TX 78666, USA.
Received 2022-8-9; Accepted 2022-10-12; Published 2023-1-1
Abstract
The genomes of two nitrogen-fixing Frankia strains, AiPa1 and AiPs1, are described as representatives of two novel candidate species. Both strains were isolated from root nodules of Alnus incana, used as capture plants in bioassays on soils from a reforested site at Karttula, Finland, that was devoid of actinorhizal plants but contained 25 year-old monocultures of spruce (Picea abies (L.) Karsten) or pine (Pinus sylvestris L.), respectively. ANI analyses indicate that each strain represents a novel Frankia species, with genome sizes of 6.98 and 7.35 Mb for AiPa1 and AiPs1, respectively. Both genomes harbored genes typical for many other symbiotic frankiae, including genes essential for nitrogen-fixation, for synthesis of hopanoid lipids and iron-sulfur clusters, as well as clusters of orthologous genes, secondary metabolite determinants and transcriptional regulators. Genomes of AiPa1 and AiPs1 had lost 475 and 112 genes, respectively, compared to those of other cultivated Alnus-infective strains with large genomes. Lost genes included one hup cluster in AiPa1 and the gvp cluster in AiPs1, suggesting that some genome erosion has started to occur in a different manner in the two strains.
Keywords: Frankia, Actinorhizal symbiosis, genome, nitrogen-fixing, biosynthetic gene clusters
Introduction
Technological advances in whole genome sequencing, in single-cell metagenomics and in comparative bioinformatics have revolutionized the description of microbial genera, species and subspecies [1-3]. Comparative sequence analyses of whole genomes and the ANI metric [4] are now used as foundations for the classification of both cultured and uncultured microbes [5-7]. Members of the genus Frankia are soil and nodule actinobacteria that have resisted isolation attempts for a long time. The first isolate was described only in 1978 [8], after which many more followed [9-12]. Differentiation of isolates has also been hampered by the limited availability of distinguishing phenotypic features between populations [13]. Consequently, species of the genus Frankia have been scantily described for many years [14, 15].
Whole genome sequencing techniques have permitted to overcome these difficulties, resulting in the description of twelve species in the genus Frankia so far, with type strains deposited in international culture collections [16-18]. Five candidate species have also been described using whole genome analyses of uncultured Frankia populations in root nodules [6, 18-20]. The number of available whole genome sequences for Frankia strains has increased significantly during the last years, with many strains potentially representing new species [21-23]. These data indicate that the genus Frankia is probably much more diverse than the twelve species and five candidate species described so far [13, 24-27]. This statement is supported by recent genome analyses of Frankia strains isolated from nodules of Alnus glutinosa as representatives of three yet undescribed nitrogen-fixing symbiotic species [22], and by the identification of two additional species of non-nitrogen-fixing and non-symbiotic frankiae [21].
Comparative sequence analyses of amplicons of an actinobacteria-specific insertion in the 23S rRNA genes of frankiae identified strains AiPa1 and AiPs1 as additional candidates for the description of new species [28]. Strains AiPa1 and AiPs1 have been isolated from root nodules of Alnus incana that was used as capture plant in bioassays aiming to determine the effects of 25 year-old monocultures of spruce (Picea abies (L.) Karsten) and pine (Pinus sylvestris L.) at a reforested site at Karttula, Finland (62° 53′, 26° 58′) on the nodulation capacity and diversity of frankiae in soils devoid of actinorhizal plants [29]. Basic soil characteristics were virtually identical for both sites, characterized as a fine silty sand with 12 to 14% organic matter and a pH of 5.3 to 5.5 [29, 30]. The aim of this study was to use whole genome sequence analyses in order to evaluate and corroborate the potential of strains AiPa1 and AiPs1 for the description of new Frankia species.
Materials and Methods
Sample preparation
Defined Propionate Medium (DPM) containing propionate and NH4Cl as C and N source [31], respectively, was used to grow Frankia strains AiPa1 and AiPs1. Cells of both strains had been preserved in 20% v/v glycerol at -80°C since 2003. After two weeks of growth at 30°C, cells were harvested by centrifugation (15,000 x g, 5 min). After a brief sonication to disrupt aggregates of cell filaments (10 s at 20% output in a S-450 sonifier, Branson Ultrasonics, Danbury, CT) [32] followed by an additional centrifugation step, DNA was extracted from cell pellets using the SurePrepTM Soil DNA Isolation Kit (Fisher Scientific, Houston, TX) [33]. DNA was sent to the Microbial Genomics Sequencing Center, Pittsburgh, PA, USA, for library preparation and sequencing using standard protocols for the Illumina tagmentation and the NextSeq Illumina platform (2 x 150 bp).
Genome assembly
Fastp was used with default settings to filter and trim sequence reads [34], where reads with average %GC<54 were removed using bbduk (https://jgi.doe.gov/data-and-tools/software-tools/bbtools/bb-tools-user-guide/). Genomes were assembled using SPAdes 3.13.0 [35] and their quality checked with QUAST [36]. Default values in the lineage workflow (lineage_set) in CheckM v1.0.18 were finally used to assess genome completeness and contamination [37].
Comparative genomic analysis
We computed Average Nucleotide Identity (ANI) [4] of the assembled genomes of AiPa1 and AiPs1 with Frankia genomes of type strains of all described species and other selected genomes using the pyani platform with the b (Blast) setting ([38]; https://pyani.readthedocs.io). Clusters of orthologous genes (COGs) [39], secondary metabolite clusters and genes specific to or lost in the new genomes were identified through antiSMASH [40] on the Mage platform [41]. An MLSA with AtpD, DnaA, FtsZ, Pgk, and RpoB was used to compute an AA distance matrix as done previously [42] to construct a phylogenetic tree using a rapid Neighbour Joining algorithm [43] and a bootstrap analysis [44].
Results
Characteristics of the two Frankia genomes
CheckM scores of 98.6% and 98.5% indicated that the genomes of strains AiPa1 and AiPs1, respectively, could be considered complete, while contamination indices of 0.27 and 4.05 demonstrated that they were pure. Genome sizes of AiPa1 and AiPs1 were 6.98 Mb and 7.35 Mb with GC contents of 71.12 and 72.13%, respectively, and were made up of 165 and 1,203 contigs with the largest contig being 310,535 and 128,115, respectively (Table 1).
Basic genome characteristics of Frankia strains AiPa1 and AiPs1 compared to those of strains and type strains of Frankia species in cluster 1. Additional comparisons with strains and type strains of clusters 2, 3 and 4 are provided in Supplemental Table S1.
| Cluster 1 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Strain | ACN14aT | ARgP5T | CpI1T | QA3 | AgB32 | AgKG'84/4 | Ag45/Mut15 | AgPM24 | AiPa1 | AiPs1 | CcI3T |
| COG1 | alni | canadensis | torreyi | casuarinae | |||||||
| G+C content (mol%) | 72.8 | 72.4 | 72.4 | 72.6 | 72.22 | 72.13 | 71.37 | 71.35 | 71.12 | 72.13 | 70.1 |
| Genome length (Mb) | 7.50 | 7.73 | 7.62 | 7.59 | 6.71 | 6.51 | 6.44 | 6.67 | 6.98 | 7.35 | 5.43 |
| # CDS | 6,714 | 7,500 | 7,201 | 7,307 | 6,364 | 6,122 | 6,088 | 6,370 | 7,601 | 6,847 | 5,593 |
| secondary metabolite clusters* | 27 | 33 | 28 | 33 | 42 | 30 | 29 | 38 | 33 | 43 | 26 |
| nifH** | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| shc | 2 | 2 | 2 | 2 | 1 | 1 | 2 | 2 | 2 | 2 | 1 |
| hupL | 2 | 2 | 2 | 2 | 1 | 1 | 1 | 1 | 2 | 1 | 2 |
| sufD | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| celA1 | 2 | 2 | 2 | 0 | 2 | 2 | 2 | 2 | 2 | 2 | 0 |
| glxA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 |
| bcsA | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 0 |
| gvpJ | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| sodF | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| geoA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| argG | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| accA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| can | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 2 | 2 |
| rhbE | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| lac | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| phdA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| dctA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 |
| tgsA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 1 |
| ddnB | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 2 | 0 |
| mopB | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 |
| qorB | 1 | 2 | 1 | 1 | 1 | 0 | 0 | 0 | 2 | 1 | 0 |
| glbN | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 2 | 2 | 1 |
| # contigs | 1 | 568 | 153 | 120 | 274 | 342 | 157 | 230 | 165 | 1,203 | 1 |
| Accession | NC_008278.1 | OESX01000001 | JYFN00000000 | WGS NZ_AJWA.1 | JALKFX000000000 | JALKFU000000000 | JALKFT000000000 | JALKFW000000000 | JALKFW000000000 | JAMQQF000000000 | CP000249.1 |
| Reference | (46) | (62) | (63) | (64) | (23) | (23) | (22) | (22) | This study | This study | (46) |
* indicates the number of clusters identified by AntiSMASH
** indicates the number of hits (>50%) following a BlastP. nif is nitrogenase, shc is squalene hopene cyclase, hup is hydrogenase uptake, suf is sulfur-iron cluster, cel is cellulase, glx is glucose oxidase, bcs is cellulose synthase, gvp is gas vesicle cluster, sodF is superoxide dismutase iron, geoA is geosmine synthase, arG is arginine, acc is acetate carboxylase, can is carbonic anhydrase, rhb is rhizobactin, lac is laccase, phd is a phytoene desaturase, dct is a dicarboxylate transporter, tgs is diacylglycerol O-acyltransferase. ddn is F420H(2)-dependent quinone nitroreductase, mop is molybdenum transport, qor is quinone oxydoreductase, glb is hemoglobin.
Phylogenetic analysis of Frankia spp
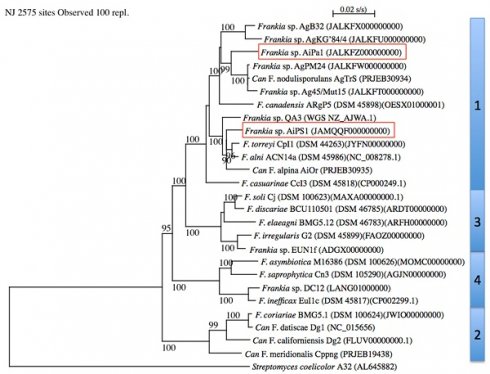
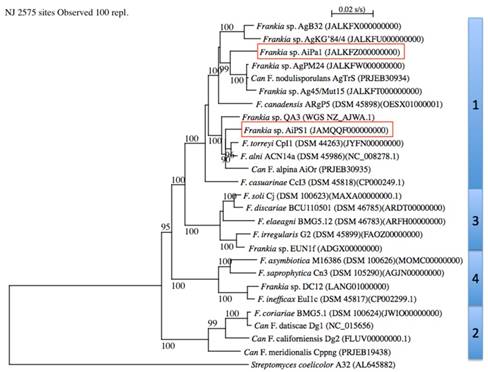
The MLSA with Frankia type strains revealed that strains AiPa1 and AiPs1 were members of Frankia cluster 1 (Figure 1). AiPa1 and AiPs1 represent two distinct lineages within the genus Frankia: strain AiPa1 has an ANI of 82% to its closest relatives (Ag45/Mut15 and AgPM24), with ANI percentages to other Frankia strains between 76 and 80%, while AiPs1 is closely related to, but distant from Frankia alni ACN14a and Frankia torreyi CpI1 with ANI values of 90 and 91%, well below the threshold of 95 proposed to delineate species (Figure 2). Both strains belong to cluster 1 frankiae, with ANI values ranging from 80% to 91%, while 76-77% values were obtained with cluster 2 genomes, and 77-78% with cluster 3 and 4 genomes (Figure 2).
Phylogenetic tree of complete genomes using Streptomyces coelicolor A32 (AL645882) as outgroup. Frankia clusters are indicated on the right. Bootstrap results above 90% are given at nodes. The bar indicates 0.02%substitution/site. The two genomes described in the present study are framed.
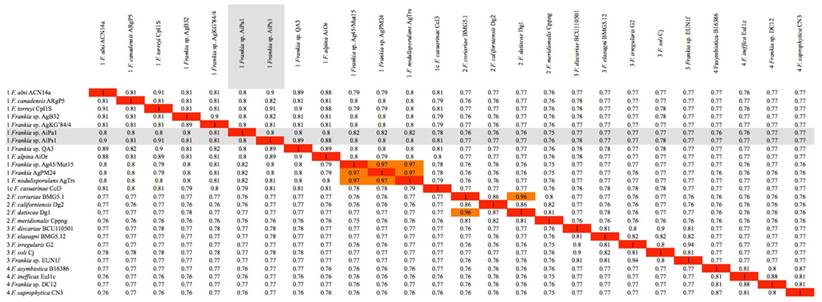
Heatmap matrix of Average Nucleotide Identity (ANI) comparisons for the Frankia genomes of strains and type strains of described species using the pyani platform with the b (Blast) setting [38]; https://pyani.readthedocs.io). The two genomes described in the present study are highlighted in grey.
Analysis of functional genes in Frankia spp. isolates
Genes such as nif, hup, suf, shc, cel, glx, bcsA meant to be characteristic for the symbiosis were present in the genomes of symbiotic lineages (clusters 1, 2 and 3) compared to non-symbiotic lineages (cluster 4) (Table 1; Table S1). Both genomes lacked gvp genes encoding gas vesicle proteins, and one of the two hup clusters commonly found in cluster 1 frankiae was absent in AiPs1 but present in AiPa1.
Both the COG and the antiSMASH computation retrieved values for AiPa1 and AiPs1 characteristic of other Alnus-infective cluster 1 strains (Table 2 and 3). These data include high number of T1PKS and NRPS (Table 3) as well as transcriptional regulators such as ArsR, and LuxR (Table 4). Additional comparisons with cluster 2, 3 and 4 frankiae are provided in Supplemental tables S2, S3, and S4. A search for genes present in F. alni ACN14a, Frankia sp. QA3, F. torreyi CpI1 and F. canadensis ARgP5 but absent in AiPa1 and AiPs1 yielded 475 or 112 hits, respectively among which a hup cluster and a dipeptide transporter in AiPa1, and the gvp cluster in AiPs1 (Table S5).
COG characteristics of Frankia strains AiPa1 and AiPs1 compared to those of type strains and strains of Frankia species in cluster 1. Additional comparisons with strains and type strains of clusters 2, 3 and 4 are provided in Supplemental Table S2.
| Cluster 1 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Strain | ACN14aT | ARgP5T | CpI1T | QA3 | AgB32 | AgKG'84/4 | Ag45/Mut15 | AgPM24 | AiPa1 | AiPs1 | CcI3T |
| COG1 | alni | canadensis | torreyi | casuarinae | |||||||
| D | 56 | 66 | 75 | 64 | 64 | 51 | 56 | 61 | 63 | 63 | 57 |
| M | 241 | 189 | 253 | 236 | 216 | 212 | 225 | 241 | 221 | 289 | 207 |
| N | 19 | 15 | 26 | 22 | 17 | 13 | 12 | 17 | 21 | 16 | 12 |
| O | 181 | 134 | 181 | 190 | 143 | 150 | 133 | 140 | 158 | 184 | 147 |
| T | 325 | 226 | 320 | 326 | 318 | 281 | 291 | 290 | 292 | 327 | 232 |
| U | 42 | 38 | 50 | 38 | 45 | 54 | 45 | 50 | 46 | 47 | 48 |
| V | 94 | 74 | 86 | 102 | 83 | 76 | 77 | 81 | 83 | 80 | 60 |
| J | 212 | 226 | 212 | 257 | 222 | 219 | 209 | 212 | 218 | 253 | 202 |
| K | 565 | 402 | 594 | 646 | 537 | 507 | 509 | 525 | 534 | 610 | 369 |
| L | 270 | 254 | 351 | 356 | 332 | 266 | 308 | 319 | 289 | 306 | 433 |
| C | 435 | 323 | 455 | 472 | 348 | 355 | 346 | 347 | 376 | 458 | 256 |
| E | 523 | 386 | 482 | 534 | 440 | 463 | 452 | 451 | 458 | 534 | 335 |
| F | 111 | 82 | 104 | 108 | 91 | 100 | 96 | 94 | 98 | 105 | 94 |
| G | 326 | 274 | 321 | 342 | 307 | 298 | 289 | 297 | 286 | 352 | 233 |
| H | 192 | 149 | 186 | 187 | 181 | 186 | 170 | 184 | 166 | 186 | 174 |
| I | 432 | 258 | 400 | 460 | 299 | 331 | 296 | 303 | 328 | 489 | 191 |
| P | 311 | 243 | 323 | 332 | 274 | 293 | 307 | 313 | 274 | 299 | 210 |
| Q | 376 | 226 | 368 | 371 | 323 | 331 | 304 | 339 | 330 | 449 | 197 |
| R | 1009 | 704 | 1005 | 1059 | 863 | 869 | 814 | 836 | 893 | 1055 | 619 |
| S | 301 | 226 | 315 | 286 | 281 | 268 | 258 | 278 | 266 | 288 | 223 |
1class: D: Cell cycle control, cell division, chromosome partitioning; M: Cell wall/membrane/envelope biogenesis; N: Cell motility; O: Posttranslational modification, protein turnover, chaperones; T: Signal transduction mechanisms; U: Intracellular trafficking, secretion, and vesicular transport; V: Defense mechanisms; J: Translation, ribosomal structure and biogenesis; K: Transcription; L: Replication, recombination and repair; C: Energy production and conversion; E: Amino acid transport and metabolism; F: Nucleotide transport and metabolism; G: Carbohydrate transport and metabolism; H: Coenzyme transport and metabolism; I: Lipid transport and metabolism; P: Inorganic ion transport and metabolism; Q: Secondary metabolites biosynthesis, transport and catabolism; R: General function prediction only; S: Function unknown.
Number of secondary metabolites clusters (antiSMASH) of Frankia strains AiPa1 and AiPs1 compared to those of cultivated strains and type strains of Frankia species in cluster 1. Additional comparisons with strains and type strains of clusters 2, 3 and 4 are provided in Supplemental Table S3.
| Cluster 1 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Strain | ACN14aT | ARgP5T | CpI1T | QA3 | AgB32 | AgKG'84/4 | Ag45/Mut15 | AgPM24 | AiPa1 | AiPs1 | CcI3T |
| alni | canadensis | torreyi | casuarinae | ||||||||
| t1PKS1 | 6 | 9 | 8 | 8 | 10 | 13 | 9 | 11 | 9 | 10 | 1 |
| t2PKS | 1 | 3 | 1 | 3 | 2 | 1 | 1 | 1 | 1 | 2 | 2 |
| t3PKS | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 |
| otherKS | 4 | 4 | 3 | 3 | 5 | 4 | 3 | 5 | 4 | 3 | 4 |
| t1pks-NRPS | 1 | 0 | 1 | 0 | 1 | 0 | 1 | 2 | 0 | 0 | 1 |
| NRPS | 3 | 6 | 2 | 2 | 6 | 8 | 6 | 6 | 4 | 8 | 0 |
| terpene | 5 | 3 | 5 | 5 | 3 | 6 | 4 | 4 | 5 | 6 | 4 |
| lanthipeptide | 1 | 1 | 1 | 3 | 2 | 1 | 0 | 3 | 3 | 0 | 6 |
| bacteriocin | 2 | 1 | 2 | 2 | 2 | 3 | 1 | 2 | 1 | 1 | 1 |
| siderophore | 1 | 1 | 1 | 1 | 1 | 2 | 1 | 1 | 2 | 1 | 1 |
| lassopeptide | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 0 |
| betalactone | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 3 | 0 | 0 |
| thiopeptide | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
| butyrolactone | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| phosphonate | 1 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 |
| arylpolyene | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| nucleoside | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
| ladderane | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| oligosaccharide | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| resorcinol | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| LAP | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 |
| other | 0 | 2 | 2 | 3 | 2 | 3 | 1 | 0 | 2 | 0 | 4 |
| Total/strain | 27 | 33 | 28 | 33 | 38 | 43 | 29 | 38 | 36 | 33 | 26 |
1tnPKS is type "n" Polyketide Synthase, NRPS is Non Ribosomal Peptide Synthase, LAP is Linear Azole/azoline-containing Peptide.
Number of transcriptional regulators of Frankia strains AiPa1 and AiPs1 compared to those of strains and type strains of Frankia species in cluster 1. Additional comparisons with strains and type strains of clusters 2, 3 and 4 are provided in Supplemental Table S4.
| Cluster 1 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Strain | ACN14aT | ARgP5T | CpI1T | QA3 | AgB32 | AgKG'84/4 | Ag45/Mut15 | AgPM24 | AiPa1 | AiPs1 | CcI3T |
| Class1 | alni | canadensis | torreyi | casuarinae | |||||||
| AraC | 9 | 9 | 10 | 16 | 9 | 13 | 6 | 6 | 12 | 16 | 2 |
| ArsR | 9 | 6 | 5 | 1 | 14 | 13 | 7 | 6 | 22 | 15 | 6 |
| AsnC | 3 | 2 | 2 | 4 | 3 | 4 | 4 | 3 | 4 | 4 | 3 |
| CRP | 4 | 2 | 1 | 1 | 4 | 3 | 4 | 4 | 5 | 6 | 2 |
| DeoR | 4 | 1 | 0 | 0 | 4 | 4 | 1 | 2 | 6 | 7 | 0 |
| DtxR | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| FurC | 2 | 3 | 3 | 4 | 3 | 3 | 3 | 3 | 3 | 4 | 2 |
| GntR | 25 | 19 | 10 | 20 | 12 | 15 | 7 | 5 | 22 | 20 | 6 |
| IclR | 3 | 6 | 4 | 9 | 7 | 8 | 4 | 3 | 12 | 5 | 2 |
| LuxR | 10 | 19 | 19 | 36 | 40 | 36 | 10 | 14 | 37 | 29 | 20 |
| LysR | 18 | 16 | 12 | 22 | 18 | 18 | 11 | 10 | 28 | 25 | 5 |
| MarR | 21 | 19 | 13 | 33 | 23 | 20 | 16 | 15 | 34 | 27 | 15 |
| MerR | 8 | 17 | 9 | 22 | 19 | 18 | 10 | 10 | 25 | 16 | 12 |
1class: AraC: arabinose regulator; ArsR: arsenic resistance; AsnC: asparagine synthase regulator; CRP: cyclic AMP receptor protein (catabolite repression); DeoR: deoxyribonucleoside synthesis operon regulator; DtxR: diphtheria toxin repressor; FurC: ferric uptake regulator; GntR: gluconate regulator; IclR: isocitrate lyase regulator; LuxR: quorum-sensing luminescence regulator; LysR: lysine regulator; MarR: Multiple antibiotic resistance regulator; MerR: mercury resistance regulator; TetR: Tetracycline repressor; WhiB: regulation of morphological differentiation.
Discussion
Genome sizes of Frankia strains have been found to be quite variable, with size ranging from 4.9 Mb to 10.7 Mb. Initial studies determined relatively consistent sizes of 7.5 Mb to 7.7 Mb for genomes of Alnus-infective cluster 1 and some cluster 3 frankiae, however, other genomes of cluster 3 frankiae were much larger (9.0 to 10.4 Mb) [45-47], similar to many cluster 4 frankiae (8.8 to 10.7 Mb) [21, 48, 49]. In contrast, genomes of cluster 1 species Frankia casuarinae (4.9 to 5.6 Mb) and Candidate species F. nodulisporulans (4.9 Mb) were smaller, similar to F. coriariae as cluster 2 representative (5.8 Mb) (see [50] for review).
Larger genomes have commonly been associated with duplications of genes involved in substrate transfers into central metabolic pathways [46]. Strains with larger genomes might therefore be considered to have a higher potential to exploit a large variety of environments [20, 46]. Smaller genomes like those of F. casuarinae, F. nodulisporulans and F. coriariae have been linked to genome reductions resulting in reduced saprotrophic potential, though their symbiotic potential is maintained [46]. The genome sizes of 6.98 and 7.35 of strains AiPa1 and AiPs1 fit into recent discoveries of smaller genome sizes for many cluster 1 strains, with a size range from 6.4 Mb to 6.7 MB (Table 1). Smaller genome sizes coincide with the loss of some duplicated genes such as, for example, the shc gene coding for the synthesis of hopanoid lipids [46], or the hup genes coding for hydrogen uptake for the recycling of hydrogen derived from nitrogenase [51]. Genes such as hup are lost by many strains (e.g. Ag45/Mut15, AgPM24, AgB32, AgKG'84/4) [22] including AiPa1, but not AiPs1 in this study, while others such as shc are lost less frequently (e.g. only in AgB32, AgKG'84/4) [23]. The numbers of genes lost by individual strains differ significantly, e.g. 475 or 112 genes for AiPa1 and AiPs1, respectively, 380 or 409 genes for AgB32 and AgKG'84/4, or 459 genes for Ag45/Mut15 and AgPM24 [22]. Thus, genome reductions could be caused by genome erosion, which could be more pronounced in some strains such as Ag45/Mut15, AgPM24, AgB32 and AgKG'84/4 compared to others, including strains AiPa1 and AiPs1.
While smaller genomes have been found so far in frankiae with high symbiotic specificity and reduced saprophytic capabilities (e.g. F. casuarinae, F. nodulisporulans and F. coriariae), potential genome reduction is not indicative of reduced saprophytic growth in strains Ag45/Mut15, AgPM24 and AiPa1. Similar to species described for clusters 1 and 3, as well as for strains AgB32, AgKG'84/4 and AiPs1, all three strains grow in the rhizosphere of host and non-host plants, but in contrast to these species also proliferate in the presence of leaf litter as sole C- and N-sources [32, 52]. Thus, these strains are able to grow on complex organic material such as leaf litter, and do not require easily available C-resources such as root exudates [53-56]. Thus, genome erosions could have affected other traits of these strains, as indicated by the low competitive ability for nodule formation of strain Ag45/Mut15 compared to F. torreyi [57, 58].
Relatives of Frankia strain AiPa1 were exclusively or most prominently found in recently vegetated soils [59, 60], while relatives of AiPs1 were only found in the older part of the plantation (65 yrs), together with relatives of AiPa1, suggesting that environmental changes in time might have promoted Frankia population changes [59]. Nodulation capacities of soils under spruce and pine monocultures were similar [61], with root nodule populations representing relatives of AiPa1 under spruce and AiPs1 under pine [29]. However, both soils harbored other Frankia populations related to strain Ag45/Mut15 that is closely related to AiPa1 [29, 61]. Thus, it remains highly speculative to draw any conclusions about potential effects of plant species on the development of infective Frankia populations in soils under 25 year-old monocultures of spruce and pine.
The availability of genome information of strains AiPa1 and AiPs1 as representatives of two new species within cluster 1 of the genus Frankia provides the possibility to assess the importance of gene erosion, but also the presence of unique genes with respect to different physiological and potentially ecological adaptations. Thus, additional studies need to include analyses of unique and additional genes to highlight differences between these two strains and other representatives of cluster 1, and relate those differences to environmental characteristics.
Supplementary Material
Supplementary tables.
Acknowledgements
The authors are indebted to the Graduate College (Doctoral Research Support Fellowship to S. Vemulapally), and the Department of Biology at Texas State University for financial support.
Data availability
The genome sequences of strains AiPa1 and AiPs1 have been deposited to Genbank under accession numbers JALKFZ000000000 and JAMQQF000000000, respectively.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Vollmers J, Wiegand S, Kaster AK. Comparing and evaluating metagenome assembly tools from a microbiologist's perspective - not only size matters!. PLoS One. 2017;12(1):e0169662
2. Tambong JT, Xu RL, Cuppels D, Chapados J, Gerdis S, Eyres J. et al. Whole-genome resources and species-level taxonomic validation of 89 plant-pathogenic Xanthomonas strains isolated from various host plants. Plant Dis. 2022;106(6):1558-65
3. Zareba-Marchewka K, Szymanska-Czerwinska M, Livingstone M, Longbottom D, Niemczuk K. Whole genome sequencing and comparative genome analyses of Chlamydia abortus strains of avian origin suggests that Chlamydia abortus species should be expanded to include avian and mammalian subgroups. Pathogens. 2021;10(11):1405
4. Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol. 2007;57:81-91
5. Gilroy R, Leng J, Ravi A, Adriaenssens EM, Oren A, Baker D. et al. Metagenomic investigation of the equine faecal microbiome reveals extensive taxonomic diversity. Peerj. 2022;10:e13084
6. Nguyen TV, Wibberg D, Vigil-Stenman T, Berckx F, Battenberg K, Demchenko KN. et al. Frankia-enriched metagenomes from the earliest diverging symbiotic Frankia cluster: they come in teams. Genome Biol Evol. 2019;11:2273-91
7. Nguyen HDT, Dodge A, Dadej K, Rintoul TL, Ponomareva E, Martin FN. et al. Whole genome sequencing and phylogenomic analysis show support for the splitting of g genus Pythium. Mycologia. 2022;114(3):501-15
8. Callaham D, Del Tredici P, Torrey JG. Isolation and cultivation in vitro of the actinomycete causing root nodulation in Comptonia. Science. 1978;199:899-902
9. Benson DR. Isolation of Frankia strains from alder actinorhizal root nodules. Appl Environ Microbiol. 1982;44:461-5
10. Caru M, Becerra A, Sepulveda D, Cabello A. Isolation of infective and effective Frankia strains from root nodules of Alnus acuminata (Betulaceae). World J Microb Biot. 2000;16:647-51
11. Gtari M, Brusetti L, Skander G, Mora D, Boudabous A, Daffonchio D. Isolation of Elaeagnus-compatible Frankia from soils collected in Tunisia. FEMS Microbiol Lett. 2004;234:349-55
12. Gtari M, Ghodhbane-Gtari F, Nouioui I, Ktari A, Hezbri K, Mimouni W. et al. Cultivating the uncultured: growing the recalcitrant cluster-2 Frankia strains. Sci Rep-Uk. 2015;5:13112 doi: 13110.11038/srep13112
13. Fernandez MP, Meugnier H, Grimont PAD, Bardin R. Deoxyribonucleic acid relatedness among members of the genus Frankia. Int J Syst Bacteriol. 1989;39:424-9
14. Lechevalier MP. Taxonomy of the genus Frankia (Actinomycetales). Int J Syst Bacteriol. 1994;44:1-8
15. Lechevalier MP, Lechevalier HA. Systematics, isolation, and culture of Frankia. In: Schwintzer C.R, Tjepkema J.D. (Eds.) The biology of Frankia and actinorhizal plants, Academic Press, New York. 1990 pp. 35-60
16. Normand P, Queiroux C, Tisa LS, Benson DR, Rouy Z, Cruveiller S. et al. Exploring the genomes of Frankia. Physiol Plantarum. 2007;130:331-43
17. Tisa LS, Oshone R, Sarkar I, Ktari A, Sen A, Gtari M. Genomic approaches toward understanding the actinorhizal symbiosis: an update on the status of the Frankia genomes. Symbiosis. 2016;70:5-16
18. Normand P, Fernandez MP. Frankia Brunchorst 1886, 174AL. In: Whitman W.B, Rainey F.A, Kämpfer P, Trujillo M.E, DeVos P, Hedlund B, Dedysh S. (Eds. ) Bergey's Manual of Systematics of Archaea and Bacteria. 2019 doi:10.1002/9781118960608.gbm00042.pub2
19. Bethencourt L, Vautrin F, Taib N, Dubost A, Castro-Garcia L, Imbaud O. et al. Draft genome sequences for three unisolated Alnus-infective Frankia Sp+ strains, AgTrS, AiOr and AvVan, the first sequenced Frankia strains able to sporulate in-planta. J. Genomics. 2019;7:50-5
20. Persson T, Battenberg K, Demina IV, Vigil-Stenman T, Heuvel BV, Pujic P. et al. Candidatus Frankia datiscae Dg1, the actinobacterial microsymbiont of Datisca glomerata, expresses the canonical nod genes nodABC in symbiosis with its host plant. PLoS One. 2015;10(5):e0127630
21. Carlos-Shanley C, Guerra T, Hahn D. Draft genomes of non-nitrogen-fixing Frankia strains. J Genomics. 2021;9:68-75
22. Normand P, Pujic P, Abrouk D, Vemulapally S, Guerra T, Carlos-Shanley C. et al. Draft genomes of nitrogen-fixing Frankia strains Ag45/Mut15 and AgPM24 isolated from root nodules of Alnus glutinosa. J Genomics. 2022;10:49-56
23. Normand P, Pujic P, Abrouk D, Vemulapally S, Guerra T, Carlos-Shanley C. et al. Draft genomes of symbiotic Frankia strains AgB32 and AgKG'84/4 from root nodules of Alnus glutinosa growing under contrasted environmental conditions. J Genomics. 2022;10:61-8
24. Hahn D. P Polyphasic taxonomy of the genus Frankia In: Pawlowski K, Newton W.E. (Eds.) Nitrogen-fixing actinorhizal symbioses. Springer Verlag, Berlin, Germany, 2007, pp. 25-48
25. Hahn D, Mirza B, Benagli C, Vogel G, Tonolla M. Typing of nitrogen-fixing Frankia strains by matrix-assisted laser desorption ionization-time-of-flight (MALDI-TOF) mass spectrometry. Syst Appl Microbiol. 2011;34:63-8
26. Pozzi AC, Bautista-Guerrero HH, Abby SS, Herrera-Belaroussi A, Abrouk D, Normand P. et al. Robust Frankia phylogeny, species delineation and intraspeciesdiversity based on Multi-Locus Sequence Analysis (MLSA) and Single-Locus Strain Typing (SLST) adapted to a large sample size. Syst Appl Microbiol. 2018;41(4):311-23
27. Normand P, Orso S, Cournoyer B, Jeannin P, Chapelon C, Dawson J. et al. Molecular phylogeny of the genus Frankia and related genera and emendation of the family Frankiaceae. Int J Syst Bacteriol. 1996;46:1-9
28. Ben Tekaya S, Ganesan AS, Guerra T, Dawson JO, Forstner MRJ, Hahn D. SybrGreen- and TaqMan-based quantitative PCR approaches allow assessment of the abundance and relative distribution of Frankia clusters in soils. Appl Environ Microb. 2017;83(5):e02833-16
29. Maunuksela L, Zepp K, Koivula T, Zeyer J, Haahtela K, Hahn D. Analysis of Frankia populations in three soils devoid of actinorhizal plants. FEMS Microbiol Ecol. 1999;28:11-21
30. Priha O, Smolander A. Microbial biomass and activity in soil and litter under Pinus sylvestris, Picea abies and Betula pendula at originally similar field afforestation sites. Biol Fertil Soils. 1997;24:45-51
31. Meesters TM, van Genesen ST, Akkermans ADL. Growth, acetylene reduction activity and localization of nitrogenase in relation to vesicle formation in Frankia strains Cc1.17 and Cp1.2. Arch Microbiol. 1985;143:137-42
32. Mirza BS, Welsh A, Hahn D. Saprophytic growth of inoculated Frankia sp. in soil microcosms. FEMS Microbiol Ecol. 2007;62:280-9
33. Samant S, Sha Q, Iyer A, Dhabekar P, Hahn D. Quantification of Frankia in soils using SYBR Green based qPCR. Syst Appl Microbiol. 2012;35:191-7
34. Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884-i890
35. Prjibelski A, Antipov D, Meleshko D, Lapidus A, Korobeynikov A. Using SPAdes de novo assembler. Curr Protoc Bioinformatics. 2020;70:e102
36. Gurevich A, Saveliev V, Vyahhi N, Tesler G. QUAST: quality assessment tool for genome assemblies. Bioinformatics. 2013;29:1072-5
37. Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043-55
38. Pritchard L, Glover RH, Humphris S, Elphinstone JG, Toth IK. Genomics and taxonomy in diagnostics for food security: soft-rotting enterobacterial plant pathogens. Anal Methods-Uk. 2016;8:12-24
39. Tatusov RL, Natale DA, Garkavtsev IV, Tatusova TA, Shankavaram UT, Rao BS. et al. The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res. 2001;29:22-28
40. Medema MH, Blin K, Cimermancic P, de Jager V, Zakrzewski P, Fischbach MA. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 2011;39:W339-346
41. Vallenet D, Calteau A, Cruveiller S, Gachet M, Lajus A, Josso A. et al. MicroScope in 2017: an expanding and evolving integrated resource for community expertise of microbial genomes. Nucleic Acids Res. 2017;45:D517-D528
42. Ondov BD, Treangen TJ, Melsted P, Mallonee AB, Bergman NH, Koren S. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 2016;17:132
43. Simonsen M, Mailund T, Pedersen CNS. Rapid Neighbour-Joining. In: Crandall K.A, Lagergren J. (Eds.) WABI 2008: Algorithms in Bioinformatics, Springer Verlag, Heidelberg. 2008 pp. 113-22
44. Felsenstein J. Confidence limits of phylogenies: an approach using the bootstrap. Evolution. 1985;39:783-91
45. Pujic P, Bolotin A, Fournier P, Sorokin A, Lapidus A, Richau KH. et al. Genome sequence of the atypical symbiotic Frankia R43 strain, a nitrogen-fixing and hydrogen-producing actinobacterium. Genome Announc. 2015;3(6):e01387-15
46. Normand P, Lapierre P, Tisa LS, Gogarten JP, Alloisio N, Bagnarol E. et al. Genome characteristics of facultatively symbiotic Frankia sp strains reflect host range and host plant biogeography. Genome Res. 2007;17:7-15
47. Nouioui I, Ghodhbane-Gtari F, Rhode M, Sangal V, Klenk HP, Gtari M. Frankia irregularis sp nov, an actinobacterium unable to nodulate its original host, Casuarina equisetifolia, but effectively nodulates members of the actinorhizal Rhamnales. Int J System Evol Microbiol. 2018;68:2883-90
48. Ghodhbane-Gtari F, Beauchemin N, Bruce D, Chain P, Chen A, Walston Davenport K. et al. Draft genome sequence of Frankia sp. strain CN3, an atypical, noninfective (Nod-) ineffective (Fix-) isolate from Coriaria nepalensis. Genome Announc. 2013;1:e0008513
49. Nouioui I, Gueddou A, Ghodhbane-Gtari F, Rhode M, Gtari M, Klenk HP. Frankia asymbiotica sp nov, a non-infective actinobacterium isolated from Morella californica root nodule. Int J Syst Evol Micr. 2017;67:4897-901
50. Gtari M, Nouioui I, Sarkar I, Ghodhbane-Gtari F, Tisa LS, Sen A. et al. An update on the taxonomy of the genus Frankia Brunchorst, 1886, 174AL. Anton Leeuw. 2019;112:5-21
51. Richau KH, Kudahettige RL, Pujic P, Kudahettige NP, Sellstedt A. Structural and gene expression analyses of uptake hydrogenases and other proteins involved in nitrogenase protection in Frankia. J Biosciences. 2013;38:703-12
52. Mirza BS, Welsh AK, Hahn D. Growth of Frankia strains in leaf litter-amended soil and the rhizosphere of a non-actinorhizal plant. FEMS Microbiol. Ecol. 2009;70:132-41
53. Smolander A. Frankia populations in soils under different tree species - with special emphasis on soils under Betula pendula. Plant Soil. 1990;121:1-10
54. Rönkkö R, Smolander A, Nurmiaho-Lassila EL, Haahtela K. Frankia in the rhizosphere of nonhost plants: A comparison with root-associated nitrogen-fixing Enterobacter, Klebsiella and Pseudomonas. Plant Soil. 1993;153:85-95
55. Nickel A, Pelz O, Hahn D, Saurer M, Siegwolf R, Zeyer J. Effect of inoculation and leaf litter amendment on establishment of nodule-forming Frankia populations in soil. Appl Environ Microbiol. 2001;67:2603-9
56. Samant S, Dawson JO, Hahn D. Growth responses of introduced Frankia strains to edaphic factors. Plant Soil. 2016;400:123-32
57. Vemulapally S, Guerra T, Hahn D. Effect of different Alnus taxa on abundance and diversity of introduced and indigenous Frankia in soils and root nodules. FEMS Microbiol Ecol. 2022;9898(3):fiac020
58. Vemulapally S, Guerra T, Weckerly FW, Hahn D. Competition of two inoculated Frankia strains in root nodulation of Alnus glutinosa seedlings and associated Frankia-strain growth in rhizospheric and non-rhizospheric soils. Plant Soil. 2022;474:115-24
59. Samant S, Amann RI, Hahn D. Evaluation of the 23S rRNA gene as target for qPCR based quantification of Frankia in soils. Syst Appl Microbiol. 2014;37:229-34
60. Samant S, Sha Q, Iyer A, Dhabekar P, Hahn D. Quantification of Frankia in soils using SYBR Green based qPCR. Syst Appl Microbiol. 2012;35:191-7
61. Maunuksela L, Hahn D, Haahtela K. Effect of freezing of soils on nodulation capacities of total and specific Frankia populations. Symbiosis. 2000;29:107-19
62. Normand P, Nouioui I, Pujic P, Fournier P, Dubost A, Schwob G. et al. Frankia canadensis sp nov, isolated from root nodules of Alnus incana subspecies rugosa. Int J Syst Evol Microbiol. 2018;68:3001-11
63. Oshone R, Hurst SGt, Abebe-Akele F, Simpson S, Morris K, Thomas WK. et al. Permanent draft genome sequences for two variants of Frankia sp. strain CpI1, the first Frankia strain isolated from root nodules of Comptonia peregrina. Genome Announc. 2016;4(1):e01588-15
64. Sen A, Beauchemin N, Bruce D, Chain P, Chen A, Walston Davenport K. et al. Draft genome sequence of Frankia sp. strain QA3, a nitrogen-fixing actinobacterium isolated from the root nodule of Alnus nitida. Genome Announc. 2013;1:e0010313
65. Wall LG, Beauchemin N, Cantor MN, Chaia E, Chen A, Detter JC. et al. Draft genome sequence of Frankia sp. strain BCU110501, a nitrogen-fixing actinobacterium isolated from nodules of Discaria trinevis. Genome Announc. 2013 1(4) e00503-13
66. Nouioui I, Beauchemin N, Cantor MN, Chen A, Detter JC, Furnholm T. et al. Draft genome sequence of Frankia sp. strain BMG5.12, a nitrogen-fixing actinobacterium isolated from Tunisian soils. Genome Announc. 2013;1(4):e00468-13
67. Nouioui I, Gtari M, Goker M, Ghodhbane-Gtari F, Tisa LS, Fernandez MP. et al. Draft genome sequence of Frankia strain G2, a nitrogen-fixing actinobacterium isolated from Casuarina equisetifolia and able to nodulate actinorhizal plants of the order Rhamnales. Genome Announc. 2016;4(3):e00437-16
68. Gtari M, Ghodhbane-Gtari F, Nouioui I. Frankia soli sp. nov, an actinobacterium isolated from soil beneath Ceanothus jepsonii. Int J Syst Evol Microbiol. 2020;70:1203-9
69. Tisa LS, Beauchemin N, Cantor MN, Furnholm T, Ghodhbane-Gtari F, Goodwin L. et al. Draft genome sequence of Frankia sp. strain DC12, an atypical, noninfective, ineffective isolate from Datisca cannabina. Genome Announc. 2015;3(4):e00889-15
Author contact
Corresponding author: Philippe Normand, Université Claude-Bernard Lyon 1, Université de Lyon, UMR 5557 CNRS Ecologie Microbienne, Villeurbanne, France. Email: philippe.normandfr

 Global reach, higher impact
Global reach, higher impact