ISSN: 1839-9940
J Genomics 2022; 10:49-56. doi:10.7150/jgen.74788 This volume Cite
Research Paper
Draft Genomes of Nitrogen-fixing Frankia Strains Ag45/Mut15 and AgPM24 Isolated from Root Nodules of Alnus Glutinosa
1. Université Claude-Bernard Lyon 1, Université de Lyon, UMR 5557 CNRS Ecologie Microbienne, Villeurbanne, France.
2. Texas State University, Department of Biology, 601 University Drive, San Marcos, TX 78666, USA.
Received 2022-5-6; Accepted 2022-5-26; Published 2022-6-6
Abstract
The genomes of two nitrogen-fixing Frankia strains, Ag45/Mut15 and AgPM24, isolated from root nodules of Alnus glutinosa are described as representatives of a novel candidate species. Phylogenomic and ANI analyses confirmed that both strains are related to cluster 1 frankiae, and that both strains belong to a novel species. At 6.4 - 6.7 Mb, their genomes were smaller than those of other cultivated Alnus-infective cluster 1 strains but larger than that of the non-cultivated Alnus-infective cluster 1 Sp+ strain AgTrS that was their closest neighbor as assessed by ANI. Comparative genomic analyses identified genes essential for nitrogen-fixation, gene composition as regards COGs, secondary metabolites clusters and transcriptional regulators typical of those from Alnus-infective cluster 1 cultivated strains in both genomes. There were 459 genes present in other cultivated Alnus-infective strains lost in the two genomes, spread over the whole of the genome, which indicates genome erosion is taking place in these two strains.
Keywords: Frankia, Actinorhizal symbiosis, genome, nitrogen-fixing frankiae, biosynthetic gene clusters
Introduction
The genus Frankia consists of nitrogen- and non-nitrogen-fixing actinobacteria that can occur in root nodules in symbiosis with a variety of woody plants [1, 2], and in soil [3]. Root nodule formation is host plant-specific, with host infection groups, i.e. the Alnus and Casuarina host infection group, the Rosaceae/Coriariaceae/Datiscaceae host infection group and the Elaeagnaceae/Rhamnaceae host infection group, respectively, largely represented by Frankia clusters 1, 2 and 3. These clusters were established by comparative analyses of ribosomal RNA gene sequences [4] and represent nitrogen-fixing frankiae, while cluster 4 frankiae are typically unable to fix N2, with one exception, and are often not able to form root nodules [4, 5]. Within clusters, assignment of strains to sub-clusters, OTUs, groups and genomospecies have been used to further describe diversity within the genus [4, 6-9].
Whole genome sequence analyses resulted in the description of several species within the genus Frankia. These analyses include isolates deposited as type strains in culture collections, as well as uncultured Frankia populations in root nodules of specific host plants described as candidate species [10]. As summarized by Normand and Fernandez [10], cluster 1 is currently the most extensively described cluster with four species and two candidate species described, while one species and three candidate species are identified in cluster 2. Four species belong to cluster 3, and three species to cluster 4, with genomes of two additional potential species published recently [11]. For cluster 1, comparative sequence analyses of amplicons of an actinobacteria-specific insertion in the 23S rRNA genes of frankiae identified several strains clustering together but distinct from type strains of cluster 1 [12]. These strains included strains Ag45/Mut15 and AgPM24 isolated from root nodules of Alnus glutinosa from two lake shores, one in Germany and one in The Netherlands about 500 km apart, i.e. Grossensee (53.631031, 10.359319) [13], and Hoogmade (52.162016, 4.591356) [7], respectively. The goal of this study was to use whole genome sequence analyses to assess the viability of our previous amplicon-based analysis, and thus affirm the potential of these strains for the description of a new species.
Materials and Methods
Sample preparation
Frankia strains Ag45/Mut15 and AgPM24 that were previously identified as members of cluster 1, representing a subcluster designated as subgroup II (14) or cluster 1b (12) were from a stock frozen at -20 °C in Defined Propionate Medium (DPM) containing propionate and NH4Cl as C and N source, respectively (15), at 30 °C for two weeks. Cells were harvested by centrifugation (15,000 × g, 5 min) and aggregates homogenized by brief sonication (10 s at 20% output in a S-450 sonifier, Branson Ultrasonics, Danbury, CT) (16). After an additional centrifugation, DNA was extracted from cell pellets using the SurePrepTM Soil DNA Isolation Kit (Fisher Scientific, Houston, TX) (17). DNA concentrations were measured with a Qubit® 2.0 Fluorometer (Life Technologies, Carlsbad, USA), and DNA sent to the Microbial Genomics Sequencing Center, Pittsburgh, PA, USA for library preparation and sequencing using the Illumina tagmentation protocol and the NextSeq Illumina platform (2 × 150 bp) using standard protocols.
Genome assembly
Sequence reads were filtered and trimmed using the default settings of fastp (18), and reads with average %GC<54 were removed using bbduk (https://jgi.doe.gov/data-and-tools/bbtools/bb-tools-user-guide/). Genomes were assembled using SPAdes 3.13.0 [19] and QUAST to check the quality of the assembled genomes [20]. Their completeness was estimated using the lineage workflow (lineage_set) CheckM v1.0.18 [21] with default values.
Comparative genomic analysis
Average Nucleotide Identity (ANI) comparisons (22) were performed for all Frankia genomes of type strains of described species and other selected genomes using the pyani platform with the b (Blast) setting ([23]; https://pyani.readthedocs.io). The genomes were compared to Frankia genomes of type strains of isolates on the Mage platform [24] to compute clusters of orthologous genes or COGs [25], to identify secondary metabolites clusters through antiSMASH [26], and identify genes specific to the new genomes or lost in the two genomes. A phylogenetic tree was reconstructed using a MASH distance matrix [27] and the tree computed dynamically directly in the Mage browser using a rapid neighbour joining algorithm [28].
Results
Sequence data
CheckM analyses showed that the assembled genomes for strains Ag45/Mut15 and AgPM24 were complete with scores of 98.09 for both strains. The number of contigs was 113 and 181 for Ag45/Mut15 and AgPM24, respectively. The largest contig was 550369 and 296366 for Ag45/Mut15 and AgPM24, respectively. The strain contamination index (CheckM) was 1.09 and 0.55 for Ag45/Mut15 and AgPM24, respectively.
Phylogenetic analysis of Frankia spp. Isolates
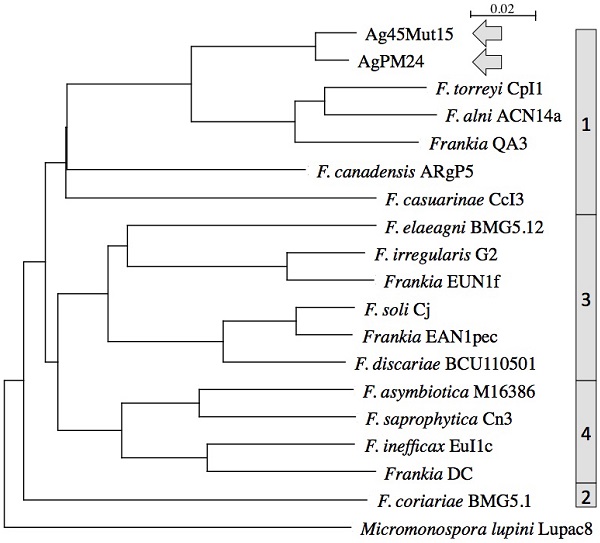
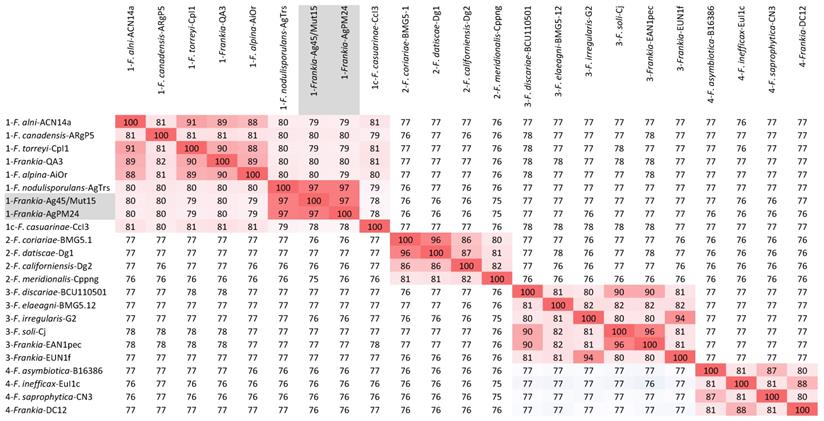
The two genomes were similar in size with about 6.4 Mb and 6.7 Mb, respectively, which is about 1 Mb smaller than those of other Alnus-infective cluster 1 cultivated strains (Table 1). They were also similar in DNA G+C% content at 71.35-71.37, which is 1% lower than values for other Alnus-infective cluster 1 cultivated strains. A phylogenetic tree generated from the MASH matrix with Frankia genomes of type strains revealed that the closest strains to Ag45/Mut15 and AgPM24 were members of cluster 1 (Figure 1). Average nucleotide identity (ANI) between strains Ag45/Mut15 and AgPM24 was 97%, indicating that they belong to a single genospecies (Figure 2). An ANI of 97% was also obtained with strain AgTrS, an uncultured Frankia population in root nodules representing Candidatus Frankia nodulisporulans. Since this strain is an obligate symbiont with a very different physiology, it will not be considered further in the present study. ANI values at or below 80% were obtained for both strains in comparison with Frankia genomes of type strains of all described species (Figure 2). The ANI values with other cluster 1 genomes ranged from 78% (CcI3) to 80% (ACN14a), while 75-76% values were obtained with cluster 2 genomes, and 76-77% with cluster 3 and 4 genomes (Figure 2).
Basic genome characteristics of Frankia strains Ag45/Mut15 and AgPM24 compared to those of type strains of Frankia species in clusters 1 to 4
| Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Strain | ACN14aT | ARgP5T | CpI1T | QA3 | Ag45/Mut15 | AgPM24 | CcI3T | BMG5.1T | BCU110501T | BMG5.12T | G2T | CjT | EAN1pec | EUN1f | M16386T | EuI1cT | Cn3T | DC12 |
| Collection | DSM 45986 | DSM 45898 | DSM 44263 | DSM 45818 | DSM 100624 | DSM 46785 | DSM 46783 | DSM 45899 | DSM 100623 | DSM 100626 | DSM 45817 | DSM 105290 | ||||||
| Frankia species | alni | canadensis | torreyi | casuarinae | coriariae | discariae | elaeagni | irregularis | soli | asymbiotica | inefficax | saprophytica | ||||||
| Genomic G+C content (mol%) | 72.8 | 72.4 | 72.4 | 72.6 | 71.37 | 71.35 | 70.1 | 71.0 | 72.3 | 71.7 | 70.9 | 71.1 | 70.94 | 70.82 | 71.93 | 72.3 | 71.8 | 71.93 |
| Genome length (nt) | 7497934 | 7730285 | 7624758 | 7590853 | 6443382 | 6672691 | 5433628 | 5795263 | 7891711 | 7589313 | 9537992 | 8061539 | 9035218 | 9322173 | 9435764 | 8815781 | 9978592 | 6884336 |
| # CDS | 6,714 | 7,500 | 7,201 | 7,307 | 6,088 | 6,370 | 5,593 | 6,487 | 7,567 | 6,977 | 8,663 | 8,108 | 9,063 | 9,428 | 8,884 | 8,099 | 9,262 | 6,630 |
| # secondary metabolite clusters* | 27 | 33 | 28 | 33 | 29 | 38 | 26 | 22 | 36 | 35 | 37 | 30 | 27 | 33 | 29 | 23 | 28 | 15 |
| nifH** | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 |
| shc | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| hupL | 2 | 2 | 2 | 2 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| sufD | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| celA1 | 2 | 2 | 2 | 0 | 2 | 2 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 1 |
| glxA | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 1 |
| bcsA | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| gvpJ | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| sodF | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 |
| geoA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 0 |
| argG | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 |
| accA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 |
| can | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 2 | 2 | 0 | 1 | 0 | 0 |
| rhbE | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 |
| lac | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 |
| phdA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 |
| dctA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 |
| tgsA | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 0 |
| ddnB | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 |
| mopB | 1 | 1 | 2 | 2 | 2 | 2 | 1 | 1 | 2 | 2 | 2 | 1 | 2 | 2 | 0 | 0 | 0 | 0 |
| qorB | 1 | 2 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 |
| glbN | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 0 | 1 |
| # contigs | 1 | 568 | 153 | 120 | 113 | 181 | 1 | 116 | 207 | 139 | 83 | 289 | 1 | 396 | 174 | 1 | 2 | 1 |
| Accession | NC_008278.1 | OESX01000001 | JYFN00000000 | WGS NZ_AJWA.1 | JALKFT000000000 | JALKFW000000000 | CP000249.1 | JWIO00000000 | ARDT00000000 | ARFH00000000 | FAOZ00000000 | MAXA00000000.1 | AAII00000000 | ADGX00000000 | MOMC00000000 | CP002299.1 | AGJN00000000 | LANG01000000 |
| Reference | (30) | (39) | (40) | (41) | this study | (30) | (38) | (42) | (43) | (44) | (45) | (30) | (46) | (47) | (46) | (30) | (46) | |
* indicates the number of clusters identified by AntiSMASH
** indicates the number of hits (>50%) following a BlastP. nif is nitrogenase, shc is squalene hopene cyclase, hup is hydrogenase uptake, suf is sulfur-iron cluster, cel is cellulase, glx is glucose oxidase, bcs is cellulose synthase,
gvp is gas vesicle cluster, sodF is superoxide dismutase iron, geoA is geosmine synthase, arG is arginine, acc is acetate carboxylase, can is carbonic anhydrase, rhb is rhizobactin, lac is laccase, phd is a phytoene desaturase,
dct is a dicarboxylate transporter, tgs is diacylglycerol O-acyltransferase. ddn is F420H(2)-dependent quinone nitroreductase, mop is molybdenum transport, qor is quinone oxydoreductase, glb is hemoglobin.
Phylogenetic tree of complete genomes using Micromonospora lupini (NZ_CAIE00000000.1) as outgroup. Clusters are indicated on the right.
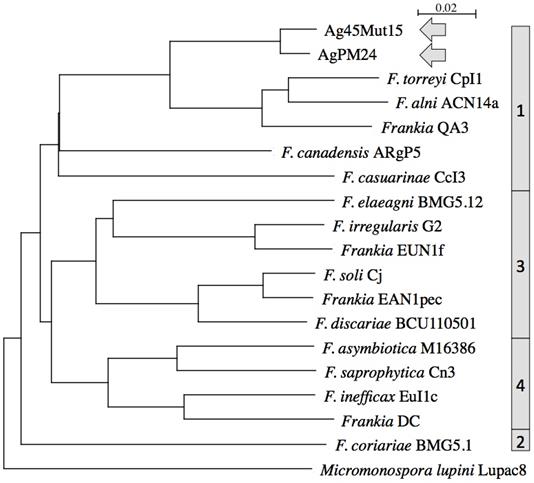
Heatmap matrix of Average Nucleotide Identity (ANI) comparisons (in percent) for the Frankia genomes of type strains of described species using the pyani platform with the b (Blast) setting (23); https://pyani.readthedocs.io). The two genomes described in the present study are highlighted in grey.
Analysis of functional genes in Frankia spp. isolates
Most genes associated with the Frankia-actinorhizal plant symbiosis were recovered in the two genomes, i.e. nif, hup, suf, shc, cel, glx, bcsA (Table 1). All genes that are more abundant in symbiotic lineages (clusters 1, 2 and 3) than in non-symbiotic lineages (cluster 4) (sodF, geoA, argF, accA, rhbE, dctA, phdA, tgsA, ddnB) were also recovered in Ag45/Mut15 and AgPM24 (Table 1). Conversely, gvp genes that code for gas vesicle proteins and one of the two hup clusters that are found in infective cluster 1 strains were not found in the two genomes.
The COG computation showed a set-up for Ag45/Mut15 and AgPM24 characteristic of other Alnus-infective cluster 1 strains with a low number of categories “N” (Cell motility) and “O” (Posttranslational modification, protein turnover, chaperones) (Table 2). These results are similar for the antiSMASH computation that showed Ag45/Mut15 and AgPM24 to have a set-up characteristic of other Alnus-infective cluster 1 strains with a high number of T1PKS and NRPS (Table 3) as were transcriptional regulators with a low number of GntR, IclR, LysR regulators (Table 4). A phyloprofile of genes present in Ag45/Mut15 and AgPM24 but without homologs at a threshold of 50% identity in AA and present in a synteny group in F. alni ACN14a, Frankia sp. QA3, F. torreyi CpI1 and F. canadensis ARgP5 yielded 1068 hits of which 621 were “proteins of unknown function”, 37 “HTH-transcriptional regulators”, 15 “acyl-CoA metabolism”, 5 “sigma factors”, 9 “amidohydrolase”, 14 “ABC transporter” and 7 “P450 cytochrome” (Table S1). Two were involved in the metabolism of xylose and xylulose.
COG characteristics of Frankia strains Ag45/Mut15 and AgPM24 compared to those of type strains of Frankia species in clusters 1 to 4
| Strain | Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ACN14aT | ARgP5T | CpI1T | QA3 | Ag45/Mut15 | AgPM24 | CcI3T | BMG5.1T | BCU110501T | BMG5.12T | G2T | CjT | EAN1pec | EUN1f | M16386T | EuI1cT | Cn3T | DC12 | |
| species | alni | canadensis | torreyi | casuarinae | coriariae | discariae | elaeagni | irregularis | soli | asymbiotica | inefficax | saprophytica | ||||||
| Class1 | ||||||||||||||||||
| D | 56 | 66 | 75 | 64 | 56 | 61 | 57 | 80 | 65 | 63 | 80 | 66 | 65 | 78 | 63 | 62 | 64 | 67 |
| M | 241 | 189 | 253 | 236 | 225 | 241 | 207 | 203 | 292 | 259 | 297 | 248 | 292 | 311 | 299 | 258 | 266 | 255 |
| N | 19 | 15 | 26 | 22 | 12 | 17 | 12 | 30 | 20 | 16 | 28 | 21 | 20 | 29 | 16 | 20 | 11 | 17 |
| O | 181 | 134 | 181 | 190 | 133 | 140 | 147 | 149 | 200 | 165 | 176 | 200 | 200 | 195 | 177 | 173 | 200 | 149 |
| T | 325 | 226 | 320 | 326 | 291 | 290 | 232 | 253 | 405 | 336 | 436 | 400 | 405 | 418 | 415 | 405 | 494 | 282 |
| U | 42 | 38 | 50 | 38 | 45 | 50 | 48 | 50 | 54 | 53 | 66 | 56 | 54 | 64 | 53 | 52 | 52 | 50 |
| V | 94 | 74 | 86 | 102 | 77 | 81 | 60 | 78 | 107 | 84 | 117 | 126 | 107 | 110 | 130 | 113 | 153 | 113 |
| J | 212 | 226 | 212 | 257 | 209 | 212 | 202 | 243 | 207 | 197 | 203 | 219 | 207 | 226 | 243 | 241 | 247 | 232 |
| K | 565 | 402 | 594 | 646 | 509 | 525 | 369 | 409 | 739 | 577 | 778 | 688 | 739 | 755 | 785 | 809 | 945 | 520 |
| L | 270 | 254 | 351 | 356 | 308 | 319 | 433 | 289 | 613 | 398 | 398 | 518 | 613 | 468 | 380 | 286 | 409 | 399 |
| C | 435 | 323 | 455 | 472 | 346 | 347 | 256 | 362 | 492 | 394 | 527 | 451 | 492 | 530 | 555 | 507 | 589 | 332 |
| E | 523 | 386 | 482 | 534 | 452 | 451 | 335 | 396 | 577 | 461 | 630 | 516 | 577 | 623 | 670 | 661 | 704 | 447 |
| F | 111 | 82 | 104 | 108 | 96 | 94 | 94 | 92 | 107 | 94 | 103 | 101 | 107 | 97 | 129 | 116 | 114 | 107 |
| G | 326 | 274 | 321 | 342 | 289 | 297 | 233 | 249 | 418 | 326 | 372 | 360 | 418 | 428 | 450 | 426 | 488 | 302 |
| H | 192 | 149 | 186 | 187 | 170 | 184 | 174 | 173 | 187 | 177 | 192 | 181 | 187 | 182 | 188 | 186 | 208 | 163 |
| I | 432 | 258 | 400 | 460 | 296 | 303 | 191 | 297 | 513 | 412 | 643 | 405 | 513 | 619 | 586 | 624 | 619 | 313 |
| P | 311 | 243 | 323 | 332 | 307 | 313 | 210 | 293 | 381 | 298 | 408 | 343 | 381 | 387 | 402 | 394 | 427 | 278 |
| Q | 376 | 226 | 368 | 371 | 304 | 339 | 197 | 320 | 488 | 369 | 565 | 417 | 488 | 550 | 531 | 534 | 569 | 256 |
| R | 1009 | 704 | 1005 | 1059 | 814 | 836 | 619 | 682 | 1216 | 969 | 1323 | 1064 | 1216 | 1280 | 1343 | 1332 | 1508 | 865 |
| S | 301 | 226 | 315 | 286 | 258 | 278 | 223 | 243 | 323 | 297 | 336 | 328 | 323 | 338 | 341 | 334 | 375 | 284 |
1class: D: Cell cycle control, cell division, chromosome partitioning; M: Cell wall/membrane/envelope biogenesis; N: Cell motility; O: Posttranslational modification, protein turnover, chaperones; T: Signal transduction mechanisms; U: Intracellular trafficking, secretion, and vesicular transport; V: Defense mechanisms; J: Translation, ribosomal structure and biogenesis; K: Transcription; L: Replication, recombination and repair; C: Energy production and conversion; E: Amino acid transport and metabolism; F: Nucleotide transport and metabolism; G: Carbohydrate transport and metabolism; H: Coenzyme transport and metabolism; I: Lipid transport and metabolism; P: Inorganic ion transport and metabolism; Q: Secondary metabolites biosynthesis, transport and catabolism; R: General function prediction only; S: Function unknown.
Number of secondary metabolites clusters (antiSMASH) of Frankia strains Ag45/Mut15 and AgPM24 compared to those of cultivated type strains of Frankia species in clusters 1 to 4
| Strain | Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ACN14aT | ARgP5T | CpI1T | QA3 | Ag45/Mut15 | AgPM24 | CcI3T | BMG5.1T | BCU110501T | BMG5.12T | G2T | CjT | EAN1pec | EUN1f | M16386T | EuI1cT | Cn3T | DC12 | |
| species | alni | canadensis | torreyi | casuarinae | coriariae | discariae | elaeagni | irregularis | soli | asymbiotica | inefficax | saprophytica | ||||||
| t1PKS1 | 6 | 9 | 8 | 8 | 9 | 11 | 1 | 6 | 16 | 13 | 6 | 9 | 5 | 9 | 6 | 5 | 2 | 1 |
| t2PKS | 1 | 3 | 1 | 3 | 1 | 1 | 2 | 2 | 1 | 1 | 2 | 1 | 1 | 2 | 1 | 2 | 1 | 1 |
| t3PKS | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 2 | 0 | 1 | 1 | 1 | 1 | 1 | 3 | 1 | 1 | 2 |
| otherKS | 4 | 4 | 3 | 3 | 3 | 5 | 4 | 1 | 4 | 3 | 6 | 4 | 4 | 6 | 2 | 1 | 2 | 1 |
| t1pks-NRPS | 1 | 0 | 1 | 0 | 1 | 2 | 1 | 0 | 1 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 |
| NRPS | 3 | 6 | 2 | 2 | 6 | 6 | 0 | 1 | 1 | 2 | 9 | 5 | 3 | 5 | 4 | 2 | 7 | 1 |
| terpene | 5 | 3 | 5 | 5 | 4 | 4 | 4 | 3 | 4 | 4 | 3 | 5 | 4 | 4 | 5 | 4 | 4 | 3 |
| lanthipeptide | 1 | 1 | 1 | 3 | 0 | 3 | 6 | 2 | 4 | 3 | 2 | 1 | 4 | 3 | 1 | 2 | 1 | 2 |
| bacteriocin | 2 | 1 | 2 | 2 | 1 | 2 | 1 | 1 | 2 | 2 | 2 | 2 | 3 | 0 | 3 | 1 | 2 | 0 |
| siderophore | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 |
| lassopeptide | 1 | 1 | 1 | 2 | 1 | 1 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 2 | 1 | 2 |
| betalactone | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 1 | 2 | 0 | 1 |
| thiopeptide | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| butyrolactone | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| phosphonate | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| arylpolyene | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| nucleoside | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| ladderane | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| oligosaccharide | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
| resorcinol | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
| LAP | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| other | 0 | 2 | 2 | 3 | 1 | 0 | 4 | 1 | 1 | 5 | 1 | 1 | 0 | 1 | 2 | 1 | 5 | 1 |
| Total/strain | 27 | 33 | 28 | 33 | 29 | 38 | 26 | 22 | 36 | 35 | 37 | 30 | 27 | 33 | 29 | 23 | 28 | 15 |
1-tnPKS is type “n” PolyKetide Synthase, NRPS is Non Ribosomal Peptide Synthase, LAP is Linear Azole/azoline-containing Peptide.
Number of transcriptional regulators of Frankia strains Ag45/Mut15 and AgPM24 compared to those of type strains of Frankia species in clusters 1 to 4
| Strain | Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ACN14aT | ARgP5T | CpI1T | QA3 | Ag45/Mut15 | AgPM24 | CcI3T | BMG5.1T | BCU110501T | BMG5.12T | G2T | CjT | EAN1pec | EUN1f | M16386T | EuI1cT | Cn3T | DC12 | |
| species | alni | canadensis | torreyi | casuarinae | coriariae | discariae | elaeagni | irregularis | soli | asymbiotica | inefficax | saprophytica | ||||||
| Class1 | ||||||||||||||||||
| AraC | 9 | 9 | 10 | 16 | 6 | 6 | 2 | 5 | 15 | 13 | 17 | 16 | 28 | 17 | 20 | 22 | 21 | 6 |
| ArsR | 9 | 6 | 5 | 1 | 7 | 6 | 6 | 5 | 4 | 4 | 11 | 6 | 11 | 9 | 9 | 16 | 8 | 8 |
| AsnC | 3 | 2 | 2 | 4 | 4 | 3 | 3 | 2 | 3 | 3 | 3 | 3 | 5 | 4 | 5 | 5 | 5 | 3 |
| CRP | 4 | 2 | 1 | 1 | 4 | 4 | 2 | 3 | 3 | 3 | 5 | 2 | 3 | 5 | 3 | 5 | 2 | 3 |
| DeoR | 4 | 1 | 0 | 0 | 1 | 2 | 0 | 0 | 2 | 1 | 0 | 2 | 4 | 0 | 2 | 2 | 2 | 1 |
| DtxR | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| FurC | 2 | 3 | 3 | 4 | 3 | 3 | 2 | 2 | 5 | 4 | 5 | 5 | 4 | 4 | 5 | 4 | 4 | 5 |
| GntR | 25 | 19 | 10 | 20 | 7 | 5 | 6 | 8 | 21 | 12 | 19 | 19 | 20 | 24 | 27 | 35 | 30 | 11 |
| IclR | 3 | 6 | 4 | 9 | 4 | 3 | 2 | 1 | 4 | 7 | 12 | 6 | 12 | 12 | 6 | 13 | 11 | 7 |
| LuxR | 10 | 19 | 19 | 36 | 10 | 14 | 20 | 15 | 22 | 9 | 18 | 15 | 58 | 40 | 18 | 64 | 29 | 14 |
| LysR | 18 | 16 | 12 | 22 | 11 | 10 | 5 | 5 | 14 | 10 | 20 | 13 | 13 | 17 | 24 | 20 | 22 | 13 |
| MarR | 21 | 19 | 13 | 33 | 16 | 15 | 15 | 23 | 18 | 20 | 31 | 25 | 27 | 30 | 32 | 33 | 35 | 19 |
| MerR | 8 | 17 | 9 | 22 | 10 | 10 | 12 | 4 | 13 | 12 | 15 | 13 | 15 | 17 | 16 | 19 | 18 | 7 |
| TetR | 92 | 78 | 117 | 127 | 61 | 65 | 30 | 47 | 113 | 77 | 39 | 126 | 147 | 156 | 156 | 191 | 18 | 59 |
| WhiB | 7 | 7 | 8 | 7 | 5 | 6 | 6 | 6 | 10 | 5 | 6 | 9 | 6 | 5 | 8 | 10 | 7 | 8 |
1class: AraC: arabinose regulator; ArsR: arsenic resistance; AsnC: asparagine synthase regulator; CRP: cyclic AMP receptor protein (catabolite repression); DeoR: deoxyribonucleoside synthesis operon regulator; DtxR: diphtheria toxin repressor; FurC: ferric uptake regulator; GntR: gluconate regulator; IclR: isocitrate lyase regulator; LuxR: quorum-sensing luminescence regulator; LysR: lysine regulator; MarR: Multiple antibiotic resistance regulator; MerR: mercury resistance regulator; TetR: Tetracycline repressor; WhiB: regulation of morphological differentiation.
A reverse study for genes present in F. alni ACN14a, Frankia sp. QA3, F. torreyi CpI1 and F. canadensis ARgP5 but absent in Ag45/Mut15 and AgPM24 yielded 459 hits among which an xanthine dehydrogenase locus, a CRISPR-locus, a acetyl/propionyl CoA carboxylase locus, an uptake hydrogenase locus, a dicarboxylate transporter, a hup locus, the GVP locus, several transporters (Table S1).
The genes lost in Ag45/Mut15 and AgPM24 have been mapped on the genome of ACN14a and found to be evenly spread over the whole genome (Supplementary Fig. 1).
Discussion
Phylogenomic and ANI analyses confirmed that strains Ag45/Mut15 and AgPM24 are related to cluster 1 frankiae, and indicate that both strains isolated from Alnus glutinosa belong to a novel species. Genome sizes of both strains were about 6.4 Mb and 6.7 Mb, respectively, and thus smaller than genomes of most cluster 1 and some cluster 3 frankiae (7.5 Mb to 7.9 Mb), though genomes of Frankia casuarinae (4.9 to 5.6 Mb) and F. nodulisporulans (4.9 Mb), as well as F. coriariae as cluster 2 representative (5.8 Mb) were even smaller (see [5] for review). Some cluster 3 frankiae were much larger in size (9.0 to 10.4 Mb) [29-31], similar to many cluster 4 frankiae (8.8 to 10.7 Mb) [11]. Frankiae with larger genomes that often result from duplications of genes involved in substrate transfers into central metabolic pathways (e.g. cluster 1, 3 and 4 frankiae), might have an enhanced potential to exploit a large variety of environments [30, 32], compared to frankiae with smaller genomes due to genome reductions that are associated to reduced saprotrophic potential, while maintaining their symbiotic potential (e.g. F. casuarinae, F. nodulisporulans and F. coriariae).
Both Ag45/Mut15 and AgPM24 belong to a group of strains within cluster 1 that are able to use leaf litter compounds as carbon resource in addition to root exudates commonly used by other strains of clusters 1, 3 and 4 [16, 33]. Together with cluster 3 frankiae, members of this group have been identified as major populations in soils, with absolute numbers depending on the sampling depth, physicochemical conditions and the vegetation [12, 14, 34, 35]. Young stands of both host trees (e.g. natural stands of A. glutinosa) [35], and non-host trees (e.g. young plantations of Betula nigra) [14] seem to promote members of this group, as do leaf litter amendments to soils, both for introduced and indigenous populations [36]. Thus, this group with strains Ag45/Mut15 and AgPM24 as representatives could be adapted to carbon resources provided by the decomposition of plant material and represent a group of frankiae characteristic of soils in early stages of plant-mediated organic matter accumulation.
Functional genes typically found in nitrogen-fixing frankiae (i.e. clusters 1, 2 and 3) within the Frankia-actinorhizal plant symbiosis were recovered in the genomes of strains Ag45/Mut15 and AgPM24. These strains appear to have lost a large number of genes dispensable for saprotrophic life as is the case in Sp+ lineages [37] or in cl2 lineages [32] where one megabase or more relative to the closest neighbor has been lost. This process of gene erosion is slow with seemingly a conserved possibility of growth under physiologically demanding conditions [38]. The full extent of diversity with cluster 1 is slowly emerging with the description of genomes from new lineages such as the present one. It appears some lineages such as F. torreyi and even more so F. casuarinae have been isolated repeatedly while others such as QA3, F. canadensis and the two lineages Ag45/Mut15 and AgPM24 described in the present study have been more rarely isolated. Two related factors have probably caused this distortion, one is the evolutionary success over long eras resulting in a higher abundance in nature and the other is the physiological ability to grow relatively fast on a large range of substrates resulting in a higher isolation success rate. Their genome composition should now be analyzed in that light of adaptation to contrasted biotopes.
Data Summary
Genomes of the strains sequenced in this study from Dr. Dittmar Hahn culture collection and were deposited in the National Center for Biotechnology Information (NCBI), under BioProject Number PRJNA680372. Whole Genome Sequencing (WGS) accession numbers are JALKFT000000000 for strain Ag45/Mut15, and JALKFW000000000 for strain AgPM24.
A list of other Frankia genomes utilized in this study can be found in Table 1. All sequences were downloaded from the NCBI Assembly database.
Supplementary Material
Supplementary figure and table.
Acknowledgements
The authors are indebted to the Graduate College (Doctoral Research Support Fellowship to S. Vemulapally), and the Department of Biology at Texas State University for financial support.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Benson DR, Dawson JO. Recent advances in the biogeography and genecology of symbiotic Frankia and its host plants. Physiol Plant. 2007;130:318-330
2. Dawson JO. Actinorhizal plants: their use in forestry and agriculture. Outlook Agricult. 1986;15:202-208
3. Chaia EE, Wall LG, Huss-Danell K. Life in soil by the actinorhizal root nodule endophyte Frankia. A review. Symbiosis. 2010;51:201-226
4. Normand P, Orso S, Cournoyer B, Jeannin P, Chapelon C. et al. Molecular phylogeny of the genus Frankia and related genera and emendation of the family Frankiaceae. Int J Syst Bacteriol. 1996;46:1-9
5. Gtari M, Nouioui I, Sarkar I, Ghodhbane-Gtari F, Tisa LS. et al. An update on the taxonomy of the genus Frankia Brunchorst, 1886, 174AL. Anton Leeuw. 2019;112:5-21
6. Hahn D. Polyphasic taxonomy of the genus Frankia. In: Pawlowski K, Newton WE. (Eds.) Nitrogen-fixing actinorhizal symbioses. Springer Verlag, Berlin, Germany. 2007 pp. 25-48
7. Hahn D, Mirza B, Benagli C, Vogel G, Tonolla M. Typing of nitrogen-fixing Frankia strains by matrix-assisted laser desorption ionization-time-of-flight (MALDI-TOF) mass spectrometry. Syst Appl Microbiol. 2011;34:63-68
8. Pozzi AC, Bautista-Guerrero HH, Abby SS, Herrera-Belaroussi A, Abrouk D. et al. Robust Frankia phylogeny, species delineation and intra species diversity based on Multi-Locus Sequence Analysis (MLSA) and Single-Locus Strain Typing (SLST) adapted to a large sample size. Syst Appl Microbiol. 2018;41:311-323
9. Fernandez MP, Meugnier H, Grimont PAD, Bardin R. Deoxyribonucleic acid relatedness among members of the genus Frankia. Int J Syst Bacteriol. 1989;39:424-429
10. Normand P, Fernandez MP. Frankia Brunchorst 1886, 174AL. In: Whitman WB, Rainey FA, Kämpfer P, Trujillo ME, DeVos P, Hedlund B, Dedysh S. (Eds.) Bergey's Manual of Systematics of Archaea and Bacteria. 2019 10.1002/9781118960608.gbm00042.pub2
11. Carlos-Shanley C, Guerra T, Hahn D. Draft genomes of non-nitrogen-fixing Frankia strains. J Genomics. 2021;9:68-75
12. Ben Tekaya S, Ganesan AS, Guerra T, Dawson JO, Forstner MRJ, Hahn D. SybrGreen- and TaqMan-based quantitative PCR approaches allow assessment of the abundance and relative distribution of Frankia clusters in soils. Appl Environ Microb. 2017 83(5), e02833-16
13. Hahn D, Starrenburg MJC, Akkermans ADL. Variable compatibility of cloned Alnus glutinosa ecotypes against ineffective Frankia strains. Plant Soil. 1988;107:233-243
14. Samant S, Huo T, Dawson JO, Hahn D. Abundance and relative distribution of Frankia host infection groups under actinorhizal Alnus glutinosa and non-actinorhizal Betula nigra trees. Microb Ecol. 2016;71:473-481
15. Meesters TM, van Genesen ST, Akkermans ADL. Growth, acetylene reduction activity and localization of nitrogenase in relation to vesicle formation in Frankia strains Cc1.17 and Cp1.2. Arch Microbiol. 1985;143:137-142
16. Mirza BS, Welsh A, Hahn D. Saprophytic growth of inoculated Frankia sp. in soil microcosms. FEMS Microbiol Ecol. 2007;62:280-289
17. Samant S, Sha Q, Iyer A, Dhabekar P, Hahn D. Quantification of Frankia in soils using SYBR Green based qPCR. Syst Appl Microbiol. 2012;35:191-197
18. Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884-i890
19. Prjibelski A, Antipov D, Meleshko D, Lapidus A, Korobeynikov A. Using SPAdes De Novo Assembler. Curr Protoc Bioinformatics. 2020;70:e102
20. Gurevich A, Saveliev V, Vyahhi N, Tesler G. QUAST: quality assessment tool for genome assemblies. Bioinformatics. 2013;29:1072-1075
21. Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043-1055
22. Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P. et al. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol. 2007;57:81-91
23. Pritchard L, Glover RH, Humphris S, Elphinstone JG, Toth IK. Genomics and taxonomy in diagnostics for food security: soft-rotting enterobacterial plant pathogens. Anal Methods-Uk. 2016;8:12-24
24. Vallenet D, Calteau A, Cruveiller S, Gachet M, Lajus A. et al. MicroScope in 2017: an expanding and evolving integrated resource for community expertise of microbial genomes. Nucleic Acids Res. 2017;45:D517-D528
25. Tatusov RL, Natale DA, Garkavtsev IV, Tatusova TA, Shankavaram UT. et al. The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res. 2001;29:22-28
26. Medema MH, Blin K, Cimermancic P, de Jager V, Zakrzewski P. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 2011;39:W339-346
27. Ondov BD, Treangen TJ, Melsted P, Mallonee AB, Bergman NH. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 2016;17:132
28. Simonsen M, Mailund T, Pedersen CNS. Rapid Neighbour-Joining. In: Crandall KA, Lagergren J. (Eds.) WABI 2008: Algorithms in Bioinformatics, Springer Verlag, Heidelberg. 2008 pp. 113-122
29. Pujic P, Bolotin A, Fournier P, Sorokin A, Lapidus A. et al. Genome sequence of the atypical symbiotic Frankia R43 strain, a nitrogen-fixing and hydrogen-producing actinobacterium. Genome Announc. 2015;3(6):e01387-15
30. Normand P, Lapierre P, Tisa LS, Gogarten JP, Alloisio N. et al. Genome characteristics of facultatively symbiotic Frankia sp strains reflect host range and host plant biogeography. Genome Res. 2007;17:7-15
31. Nouioui I, Ghodhbane-Gtari F, Rhode M, Sangal V, Klenk HP. et al. Frankia irregularis sp nov, an actinobacterium unable to nodulate its original host, Casuarina equisetifolia, but effectively nodulates members of the actinorhizal Rhamnales. Int J Syst Evol Microbiol. 2018;68:2883-2890
32. Persson T, Battenberg K, Demina IV, Vigil-Stenman T, Heuvel BV. et al. Candidatus Frankia datiscae Dg1, the actinobacterial microsymbiont of Datisca glomerata, expresses the canonical nod genes nodABC in symbiosis with its host plant. PLoS One. 2015;10(5):e0127630
33. Mirza BS, Welsh AK, Hahn D. Growth of Frankia strains in leaf litter-amended soil and the rhizosphere of a non-actinorhizal plant. FEMS Microbiol Ecol. 2009;70:132-141
34. Samant SS, Dawson JO, Hahn D. Growth responses of indigenous Frankia populations to edaphic factors in actinorhizal rhizospheres. Syst Appl Microbiol. 2015;38:501-505
35. Samant S, Amann RI, Hahn D. Evaluation of the 23S rRNA gene as target for qPCR based quantification of Frankia in soils. Syst Appl Microbiol. 2014;37:229-234
36. Nickel A, Pelz O, Hahn D, Saurer M, Siegwolf R, Zeyer J. Effect of inoculation and leaf litter amendment on establishment of nodule-forming Frankia populations in soil. Appl Environ Microbiol. 2001;67:2603-2609
37. Bethencourt L, Vautrin F, Taib N, Dubost A, Castro-Garcia L. et al. Draft genome sequences for three unisolated Alnus-infective Frankia Sp+ strains, AgTrS, AiOr and AvVan, the first sequenced Frankia strains able to sporulate in-planta. J Genomics. 2019;7:50-55
38. Gtari M, Ghodhbane-Gtari F, Nouioui I, Ktari A, Hezbri K. et al. Cultivating the uncultured: growing the recalcitrant cluster-2 Frankia strains. Sci Rep-Uk. 2015;5:13112 doi: 13110.11038/srep13112
39. Normand P, Nouioui I, Pujic P, Fournier P, Dubost A. et al. Frankia canadensis sp nov, isolated from root nodules of Alnus incana subspecies rugosa. Int J Syst Evol Microbiol. 2018;68:3001-3011
40. Oshone R, Hurst SGt, Abebe-Akele F, Simpson S, Morris K. et al. Permanent draft genome sequences for two variants of Frankia sp. strain CpI1, the first Frankia strain isolated from root nodules of Comptonia peregrina. Genome Announc. 2016;4(1):e01588-15
41. Sen A, Beauchemin N, Bruce D, Chain P, Chen A. et al. Draft genome sequence of Frankia sp. strain QA3, a nitrogen-fixing actinobacterium isolated from the root nodule of Alnus nitida. Genome Announc. 2013;1:e0010313
42. Wall LG, Beauchemin N, Cantor MN, Chaia E, Chen A. et al. Draft genome sequence of Frankia sp. strain BCU110501, a nitrogen-fixing actinobacterium isolated from nodules of Discaria trinevis. Genome Announc. 2013;1(4):e00503-13
43. Nouioui I, Beauchemin N, Cantor MN, Chen A, Detter JC. et al. Draft genome sequence of Frankia sp. strain BMG5.12, a nitrogen-fixing actinobacterium isolated from Tunisian soils. Genome Announc. 2013;1(4):e00468-13
44. Nouioui I, Gtari M, Goker M, Ghodhbane-Gtari F, Tisa LS. et al. Draft genome sequence of Frankia strain G2, a nitrogen-fixing actinobacterium isolated from Casuarina equisetifolia and able to nodulate actinorhizal plants of the order Rhamnales. Genome Announc. 2016;4(3):e00437-16
45. Gtari M, Ghodhbane-Gtari F, Nouioui I. Frankia soli sp. nov, an actinobacterium isolated from soil beneath Ceanothus jepsonii. Int J Syst Evol Microbiol. 2020;70:1203-1209
46. Tisa LS, Oshone R, Sarkar I, Ktari A, Sen A. et al. Genomic approaches toward understanding the actinorhizal symbiosis: an update on the status of the Frankia genomes. Symbiosis. 2016;70:5-16
47. Nouioui I, Gueddou A, Ghodhbane-Gtari F, Rhode M, Gtari M. et al. Frankia asymbiotica sp nov, a non-infective actinobacterium isolated from Morella californica root nodule. Int J Syst Evol Microbiol. 2017;67:4897-4901
Author contact
Corresponding author: Philippe Normand, Université Claude-Bernard Lyon 1, Université de Lyon, UMR 5557 CNRS Ecologie Microbienne, Villeurbanne, France. E-mail: philippe.normandfr.

 Global reach, higher impact
Global reach, higher impact