ISSN: 1839-9940
J Genomics 2022; 10:1-7. doi:10.7150/jgen.67277 This volume Cite
Research Paper
Comparative analysis of mRNA transcripts of HT-29 cell line expressed in identical quantities for pathogenic E. coli strains UM146 and UM147 with control Escherichia coli Nissle 1917
Department of Molecular Biotechnology and Microbiology, Gdansk University of Technology, Faculty of Chemistry, Narutowicza 11/12, 80-233 Gdansk, Poland.
Abstract
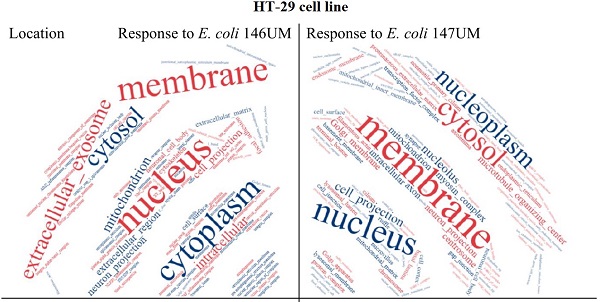
Aim of study was comparative analysis of mRNA transcripts of HT-29 cell line, expressed in identical quantities for the combination of pathogenic and non-pathogenic Escherichia coli strains. HT-29 confluent monolayers infection with two pathogenic E. coli strains UM146 and UM147 resulted in two sets of mRNA transcripts that were identical with RNA transcripts obtained for non-pathogenic one strain E. coli Nissle 1917. In this study genome-wide experiments were conducted using expression microarray-system. Only one common mRNA transcript coding for CCDC65 gene was equally expressed by HT-29 cells after incubation challenge with three different E. coli strains used. This gene and its bacterial analogue are important in the ciliary or flagellar motility, respectively. Altogether, 78 and 81 HT-29 mRNA transcripts for E. coli UM146 and E. coli UM147 had identical RNA quantity in comparison to the response obtained for non-pathogenic E. coli Nissle 1917 interactions with HT-29 monolayers. Specific analysis using REACTOME and agriGO terms enrichment data-mining tools as well as word-cloud analysis allowed for identification the most important processes characteristic during HT-29 cell line infections for each pathogenic E. coli strain used. The importance of results may contribute to recognition of those processes during bacterial infections that are identical with processes arising from human interaction with non-pathogenic strains that belong to the same bacterial species.
Keywords: HT-29 cell line, Escherichia coli, PAI I, trypsin-like activity, infection

 Global reach, higher impact
Global reach, higher impact