ISSN: 1839-9940
J Genomics 2019; 7:60-63. doi:10.7150/jgen.39039 This volume Cite
Short Research Paper
Whole Genome Sequence of Marinobacter salarius Strain SMR5, Shown to Promote Growth in its Diatom Host
1. Department of Marine Sciences, University of Gothenburg, Göteborg, Sweden;
2. Gothenburg Global Biodiversity Centre, Göteborg, Sweden;
3. Department of Biological and Environmental Sciences, University of Gothenburg, Göteborg, Sweden.
*M.T and M.I.M.P. contributed equally to this work.
Received 2019-8-5; Accepted 2019-9-8; Published 2019-10-1
Abstract
Attempts to obtain axenic cultures of the marine diatom Skeletonema marinoi often result in poor growth, indicating the importance of the microbiome to the growth of its host. In order to identify the precise roles played by these associated bacteria, individual strains were isolated, cultured and sequenced. We report the genome of one such strain - SMR5, isolated from a culture of S. marinoi strain R05AC sampled from top layer sediments of the Swedish west coast. Its genome of 4,630,160 bp consists of a circular chromosome and one circular plasmid, and 4,263 CDSs were inferred in the annotation. Comparison of 16S rRNA sequences and other markers, along with phylotaxonomic analysis, leads us to place strain SMR5 in the taxon Marinobacter salarius. Pathway analysis and previous experimental work suggest that this strain may produce a growth factor, as well as improve iron availability for its host via siderophores.
Keywords: Whole Genome Sequencing, Marinobacter, Diatom, Skeletonema, Microbiome, Marine sediment
Introduction
We have identified several bacterial strains living in association with strain R05AC of the chain-forming centric diatom Skeletonema marinoi [1-4]. Bacteria and diatoms are known to interact in a number of ways, ranging from mutually beneficial nutrient exchange to parasitism [5], but the precise roles of the bacteria within the strain R05AC holobiont are as yet unknown. Therefore, we undertook a systematic approach to investigating these potential interactions by isolating individual bacterial strains associated with cultures of the diatom, then sequencing and annotating their genomes. One bacterial strain isolated this way was SMR5, whose genome is presented here.
The host culture, S. marinoi strain R05AC, was originally established from a germinated resting cell, collected in May 2010 from top layer sediment at 14m depth in Öresund, Sweden (55°59.16′ N, 12°44.02′ E). Strain SMR5 was isolated by dilution streaking from strain R05AC, and thenceforth maintained on marine agar plates (BD Difco - Fisher Scientific, USA). A distinguishing feature of strain SMR5, compared to the other bacteria identified so far in the S. marinoi microbiome, is the formation of white colonies with diffuse edges when grown on marine agar plates, compared to the firm colonies produced by all other bacterial species identified. Under microscopic observation, strain SMR5 can be seen to have a rod-shaped cell morphology and flagella.
Sequencing of strain SMR5 was performed using one SMRT cell on the PacBio RSII platform (Pacific Biosciences, Menlo Park, CA, USA), producing 163,482 unfiltered reads totalling 2.3 Gbp. The reads were filtered using the default parameters of SMRT Portal version 2.3.0's P_Filter module (minSubReadLength 500, readScore 0.80, minLength 100) [6], giving 101,723 filtered reads of 1.3 Gbp total. These reads were then assembled using the de novo assembler Canu version 1.3 [7] (genomeSize parameter 4.5m). As Canu generates overlapping regions at the ends of contigs when assembling circular sequences, BLASTn [8] was used to identify the extent of these overlaps, in order to allow the contigs to be circularized. In this way, regions of 17,082 bp and 21,124 bp, highly similar between the beginning and end of their respective contigs, were manually trimmed from the start of the assembled chromosome and plasmid, respectively. Contig circularization was then confirmed by joining each contig's corresponding ends and realigning the reads using the RS_Resequencing.1 protocol on SMRT Portal version 2.3.0 (Pacific Biosciences, [6]), which showed consistent read coverage across the joins. Correction of the assembled contigs was also performed during this step using the Quiver algorithm [6]. The final assembly contained one circular chromosome and one circular plasmid, totalling 4,630,160 bp, with an average read coverage of 213.58x. The strain SMR5 chromosome is 4,386,892 bp long with a G+C content of 57.2%, and plasmid pSMR5 is 243,268 bp (G+C 53.7%) (assembly details summarized in Table 1). Annotation of the genome was performed using Prokka version 1.12beta [9], which inferred 4,263 CDSs (3,508 with functional predictions), 7 pseudogenes, 51 tRNAs, 9 rRNAs, 11 ncRNAs, and one tmRNA (annotation details summarized in Table 1).
Statistics for the assembly and annotation of Marinobacter salarius strain SMR5.
| Total assembly | Chromosome | pSMR5 | |
|---|---|---|---|
| Assembly | |||
| Number of reads (filtered) | 101,723 | ||
| Number of bases (filtered) | 1,318,128,196 bp | ||
| Overlapping bases trimmed from start of contig | 17,082 bp | 21,124 bp | |
| Final assembly size | 4,630,160 bp | 4,386,892 bp | 243,268 bp |
| G+C content | 57.0% | 57.2% | 53.7% |
| Average read coverage | 213.58x | ||
| Annotation | |||
| CDS | 4,263 | 3,999 | 264 |
| Pseudogenes | 7 | 6 | 1 |
| tRNA | 51 | 51 | 0 |
| rRNA | 9 | 9 | 0 |
| ncRNA | 11 | 10 | 1 |
| tmRNA | 1 | 1 | 0 |
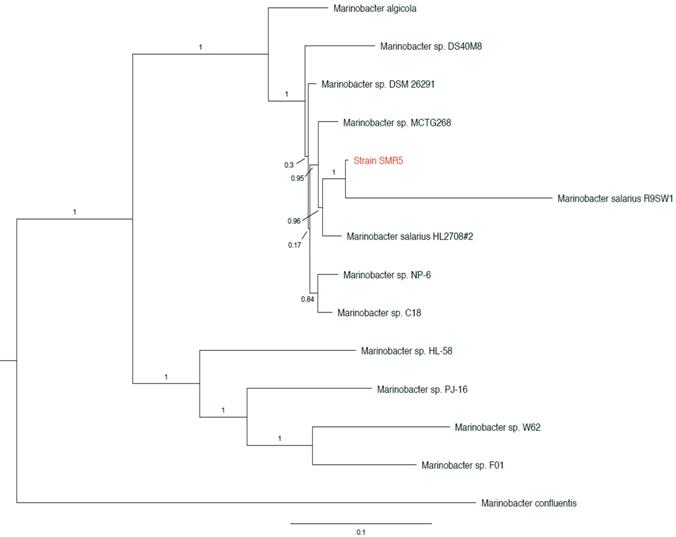
The three 16S rRNA sequences of strain SMR5 (one differing from the other two by a single base substitution) share 99.8%/99.9% identity with the 16S sequence of Marinobacter algicola strain DG893T (accession no. NZ_ABCP00000000), and 99.4%/99.5% identity with that of Marinobacter salarius strain R9SW1T (accession no. NZ_CP007152). While not a type strain, the three identical 16S sequences of M. salarius strain HL2708#2 (accession no. NZ_CP021333) were found to have 99.9%/100% identity to strain SMR5. In addition, we found multiple sequences from unnamed Marinobacter sp. strains with ≥99.8% 16S identity to strain SMR5 in NCBI's RefSeq database [10]. We then used PhyloPhlAn version 0.99 [11] to perform a phylotaxonomic comparison of strain SMR5 against representatives of all whole-genome sequenced Alteromonadaceae species available on NCBI's RefSeq ftp site (ftp://ftp.ncbi.nlm.nih.gov/genomes/refseq/bacteria/; accessed 27 May 2019). This analysis showed that strain SMR5 is sister to M. salarius strain R9SW1T (100% bootstrap support), with M. algicola appearing more distantly related (Figure 1).
Considering this apparent discrepancy between the 16S and phylotaxonomic analyses with regard to type strains, we also examined several conserved genes which have previously been used in Marinobacter classification - gyrB, rpoB and rpoD [12,13] - in order to more precisely classify strain SMR5. Whereas M. algicola strain DG893T had <96% sequence identity to the homologs in strain SMR5, M. salarius strain R9SW1T showed sequence identity of >99%. On this basis, we place strain SMR5 in the taxon Marinobacter salarius.
In terms of strain SMR5's relationship with its host, enhanced growth of S. marinoi strain R05AC in the presence of excess strain SMR5 has been observed in culture [14]. Based on examination of the annotation using Pathway Tools version 21.0 [15], our current hypothesis is that this may be caused by the production of indole-3-acetic acid (auxin) by strain SMR5. This hormone is known to promote growth in diatoms as well as plants [16], and the Pathway Tools analysis highlighted a nitrilase gene in the SMR5 genome (MARSALSMR5_03867) as being potentially involved in an auxin biosynthesis pathway (indole-3-acetate biosynthesis V in MetaCyc (https://metacyc.org; [17]). Of additional note is the presence of a membrane transport protein (MARSALSMR5_00349) that, when compared to other Marinobacter proteins using BLASTp [8], appears to be an auxin efflux carrier, a protein present in many Marinobacter species. If this growth enhancement is indeed caused by auxin, it would be consistent with observations of other diatom-bacteria interactions, such as those between Pseudo-nitzschia multiseries and Sulfitobacter [16].
Clade of a phylogenetic tree showing placement of strain SMR5 (highlighted in red) within the family Alteromonadaceae. Adapted from tree generated with PhyloPhlAn version 0.99 [11]. Branch labels represent bootstrap values; scale bar indicates the mean number of nucleotide substitutions per site. Tree visualised with FigTree version 1.4.3 [20].
In addition, some phytoplankton-associated Marinobacter species are known to produce siderophores, which make iron available to both the bacteria and their phytoplankton host [18]. Secondary metabolite gene cluster prediction using antiSMASH version 5.0.0 [19] highlighted a region of the chromosome potentially involved in siderophore biosynthesis (corresponding to loci MARSALSMR5_03200 - MARSALSMR5_03239), indicating that strain SMR5 may also be capable of this.
Nucleotide sequence accession numbers
This whole-genome project has been deposited in GenBank under the accession numbers CP020931 and CP020932 as part of BioProject no. PRJNA380207.
Abbreviations
CDS: coding sequence; SMRT: single-molecule real-time; ncRNA: non-coding RNA; tmRNA: transfer-messenger RNA.
Acknowledgements
This work was supported by the Gordon and Betty Moore Foundation (A.K.C., M.T., A.G.; Grant No. 4967), the Swedish Research Council VR (A.K.C., Grant No. 2015-04286) and the Swedish Research Council Formas (M.T., Grant No. 2017-00466; A.G., Grant No. 217-2012-692).
We thank the Linnéus Centre for Marine Evolutionary Biology (CeMEB, http://cemeb.science.gu.se/) for support. All bioinformatics analyses were run on the Albiorix computer cluster (http://albiorix.bioenv.gu.se/) at the Department of Marine Sciences, University of Gothenburg.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Töpel M, Pinder MIM, Johansson ON, Kourtchenko O, Godhe A, Clarke AK. Genome Sequence of Roseovarius mucosus Strain SMR3, Isolated from a Culture of the Diatom Skeletonema marinoi. Genome Announc. 2017;5(22):e00394-17 doi:10.1128/genomeA.00394-17
2. Töpel M, Pinder MIM, Johansson ON, Kourtchenko O, Godhe A, Clarke AK. Complete Genome Sequence of Loktanella vestfoldensis Strain SMR4r, a Novel Strain Isolated from a Culture of the Chain-Forming Diatom Skeletonema marinoi. Genome Announc. 2018;6(12):e01558-17 doi:10.1128/genomeA.01558-17
3. Töpel M, Pinder MIM, Johansson ON, Kourtchenko O, Clarke AK, Godhe A. Complete Genome Sequence of Novel Sulfitobacter pseudonitzschiae Strain SMR1, Isolated from a Culture of the Marine Diatom Skeletonema marinoi. J Genomics. 2019;7:7-10 doi:10.7150/jgen.30559
4. Töpel M, Pinder MIM, Johansson ON, Kourtchenko O, Godhe A, Clarke AK. Complete Genome Sequence of the Diatom-Associated Bacterium Sphingorhabdus sp. Strain SMR4y. Microbiol Resour Announc. 2019;8(29):e00482-19 doi: 10.1128/MRA.00482-19
5. Amin SA, Parker MS, Armbrust EV. Interactions between Diatoms and Bacteria. Microbiol Mol Biol Rev. 2012;76(3):667-684 doi:10.1128/MMBR.00007-12
6. Chin C-S, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods. 2013;10(6):563-569 doi:10.1038/nmeth.2474
7. Koren S, Walenz BP, Berlin K, Miller JR, Bergman NH, Phillippy AM. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017;27(5):722-736 doi:10.1101/gr.215087.116
8. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403-410 doi:10.1016/S0022-2836(05)80360-2
9. Seemann T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068-2069 doi:10.1093/bioinformatics/btu153
10. O'Leary NA, Wright MW, Brister JR, Ciufo S, Haddad D, McVeigh R, Rajput B, Robbertse B, Smith-White B, Ako-Adjei D, Astashyn A, Badretdin A, Bao Y, Blinkova O, Brover V, Chetvernin V, Choi J, Cox E, Ermolaeva O, Farrell CM, Goldfarb T, Gupta T, Haft D, Hatcher E, Hlavina W, Joardar VS, Kodali VK, Li W, Maglott D, Masterson P, McGarvey KM, Murphy MR, O'Neill K, Pujar S, Rangwala SH, Rausch D, Riddick LD, Schoch C, Shkeda A, Storz SS, Sun H, Thibaud-Nissen F, Tolstoy I, Tully RE, Vatsan AR, Wallin C, Webb D, Wu W, Landrum MJ, Kimchi A, Tatusova T, DiCuccio M, Kitts P, Murphy TD, Pruitt KD. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 2016;44(D1):D733-D745 doi:10.1093/nar/gkv1189
11. Segata N, Börnigen D, Morgan XC, Huttenhower C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat Commun. 2013;4:2304. doi:10.1038/ncomms3304
12. Ng HJ, López-Pérez M, Webb HK, Gomez D, Sawabe T, Ryan J, Vyssotski M, Bizet C, Malherbe F, Mikhailov VV, Crawford RJ, Ivanova EP. Marinobacter salarius sp. nov. and Marinobacter similis sp. nov, Isolated from Sea Water. PLOS ONE. 2014;9(9):e106514. doi:10.1371/journal.pone.0106514
13. Gao B, Mohan R, Gupta RS. Phylogenomics and protein signatures elucidating the evolutionary relationships among the Gammaproteobacteria. Int J Syst Evol Microbiol. 2009;59(2):234-247 doi:10.1099/ijs.0.002741-0
14. Johansson ON, Pinder MIM, Ohlsson F, Egardt J, Töpel M, Clarke AK. Friends With Benefits: Exploring the Phycosphere of the Marine Diatom Skeletonema marinoi. Front Microbiol. 2019;10:1828. doi:10.3389/fmicb.2019.01828
15. Karp PD, Paley S, Romero P. The Pathway Tools Software. Bioinformatics. 2002;18(suppl_1):S225-S232 doi:10.1093/bioinformatics/18.suppl_1.S225
16. Amin SA, Hmelo LR, van Tol HM, Durham BP, Carlson LT, Heal KR, Morales RL, Berthiaume CT, Parker MS, Djunaedi B, Ingalls AE, Parsek MR, Moran MA, Armbrust EV. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature. 2015;522:98-101 doi:10.1038/nature14488
17. Caspi R, Altman T, Billington R, Dreher K, Foerster H, Fulcher CA, Holland TA, Keseler IM, Kothari A, Kubo A, Krummenacker M, Latendresse M, Mueller LA, Ong Q, Paley S, Subhraveti P, Weaver DS, Weerasinghe D, Zhang P, Karp PD. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases. Nucleic Acids Res. 2014;42(D1):D459-D471 doi:10.1093/nar/gkt1103
18. Amin SA, Green DH, Hart MC, Küpper FC, Sunda WG, Carrano CJ. Photolysis of iron-siderophore chelates promotes bacterial-algal mutualism. PNAS. 2009;106(40):17071-17076 doi:10.1073/pnas.0905512106
19. Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T. antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019;47(W1):W81-W87 doi:10.1093/nar/gkz310
20. Rambaut A. http://tree.bio.ed.ac.uk/software/figtree/
Author contact
![]() Corresponding author: Mats Töpel, mats.topelgu.se
Corresponding author: Mats Töpel, mats.topelgu.se

 Global reach, higher impact
Global reach, higher impact