ISSN: 1839-9940
J Genomics 2019; 7:46-49. doi:10.7150/jgen.35061 This volume Cite
Short Research Paper
Genome Sequence of Kordia sp. Strain SMS9 Identified in a Non-Axenic Culture of the Diatom Skeletonema marinoi
1. Department of Marine Sciences, University of Gothenburg, Göteborg, Sweden;
2. Department of Biological and Environmental Sciences, University of Gothenburg, Göteborg, Sweden;
3. Clinical Genomics Göteborg, SciLifeLab, Göteborg, Sweden;
4. Gothenburg Global Biodiversity Centre, Göteborg, Sweden.
Received 2019-3-20; Accepted 2019-4-18; Published 2019-5-9
Abstract
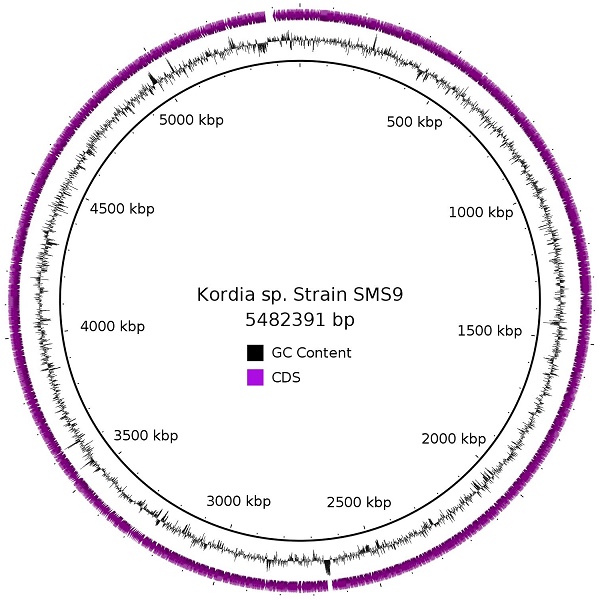
Initial efforts to sequence the genome of the marine diatom Skeletonema marinoi were hampered by the presence of genetic material from bacteria, and there was sufficient material from some of these bacteria to enable the assembly of full chromosomes. Here, we report the genome of strain SMS9, one such bacterial species identified in a non-axenic culture of S. marinoi strain ST54. Its 5,482,391 bp circular chromosome contains 4,641 CDSs, and has a G+C content of 35.6%. Based on 16S rRNA comparison, phylotaxonomic analysis, and the genome similarity metrics dDDH and OrthoANI, we place this strain in the genus Kordia, and to the best of our knowledge, this is the first Kordia species to be initially described from European waters. As attempts to culture this strain have failed, however, the specifics of its relationship with S. marinoi are still uncertain.
Keywords: Whole Genome Sequencing, Kordia, Diatom, Skeletonema, Microbiome, Marine sediment
Introduction
A variety of different bacteria are known to be associated with diatoms, and their interactions can take a variety of forms, ranging from mutually beneficial to parasitic [1]. One side-effect of these associations is the difficulty of obtaining an axenic diatom culture for the purposes of genome sequencing. Such problems were encountered during attempts to sequence the genome of the chain-forming marine diatom Skeletonema marinoi strain ST54, resulting in the assembly of both diatom and bacterial contigs. However, as complete bacterial chromosomes were discovered in the assembly, the opportunity was taken to gain insight into bacteria which may form part of this diatom strain's microbiome. One of the bacterial chromosomes assembled in this way was that of strain SMS9, which we present here.
The culture of S. marinoi strain ST54 was established from a revived resting-stage cell taken from top layer sediment in Kosterfjord, Sweden (58°51.0' N, 10°45.7' E; 102m depth) in May 2009. Sediment collection was performed using a box corer. Sequencing of DNA from this non-axenic culture was performed on 11 SMRT cells using PacBio RSII technology (Pacific Biosciences, Menlo Park, CA, USA), with an output of 183,658 uncorrected reads (1.1 Gbp total). Assembly was performed using the de novo assembler HGAP.2, as part of SMRT Portal version 2.3.0 (Pacific Biosciences, [2]), with a seed read length parameter of 7.5 kbp and an approximate genome size set to 55 Mb. Metaxa2 version 2.1.2 was used to search the resultant assembly for 16S ribosomal RNA sequences in order to find bacterial contigs [3], which identified a 5,489,227 bp contig containing a Kordia 16S sequence.
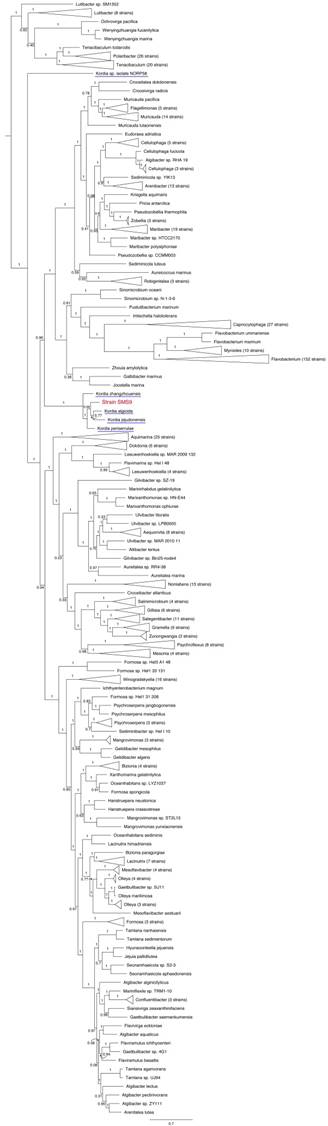
Subtree taken from an outgroup-rooted phylogenetic tree of 734 species, highlighting the position of strain SMS9 (in red) within the Flavobacteriaceae; other Kordia species are underlined in blue (note the distant location of K. sp. isolate NORP58 relative to the other Kordia species). Crocinitomix algicola (accession no. LYPF00000000) and C. catalasitica (accession no. JHXV00000000) were used as the outgroup, belonging to Crocinitomicaceae, a sister family to Flavobacteriaceae within the order Flavobacteriales. Adapted from tree generated using PhyloPhlAn version 0.99 [6], and visualised using FigTree version 1.4.3 [16]. Branch labels represent bootstrap values; scale bar indicates the mean number of substitutions per site.
Due to the nature of the assembly process, overlapping sequences were present at each end of the contig. Two 4,227 bp regions at the start and end of the contig, respectively, were identified by BLASTn as being almost identical to one another [4], and the region at the start of the contig was manually removed. The corresponding ends were then joined, and the PacBio reads were realigned to this sequence using the RS_Resequencing.1 protocol on SMRT Portal [2], to ensure that read coverage was consistent across the join, supporting a true circular molecule. This protocol includes a correction step using the Quiver algorithm [2], and was run an additional two times for further polishing. The final result is a circular chromosome of 5,482,391 bp, with G+C content of 35.6% and average read coverage of 67.46x. As this chromosome was assembled from a metagenome, we do not know whether any plasmids are also present in this strain; however, to the best of our knowledge, no other Kordia species described so far has been reported to contain a plasmid. Annotation of strain SMS9 was performed using the prokaryotic annotation software Prokka version 1.12beta [5], which inferred 4,641 CDSs (2,747 protein-coding genes with a functional prediction and 1,894 labelled as hypothetical), 6 pseudogenes, 63 tRNAs, 9 rRNAs, 6 ncRNAs, and one tmRNA.
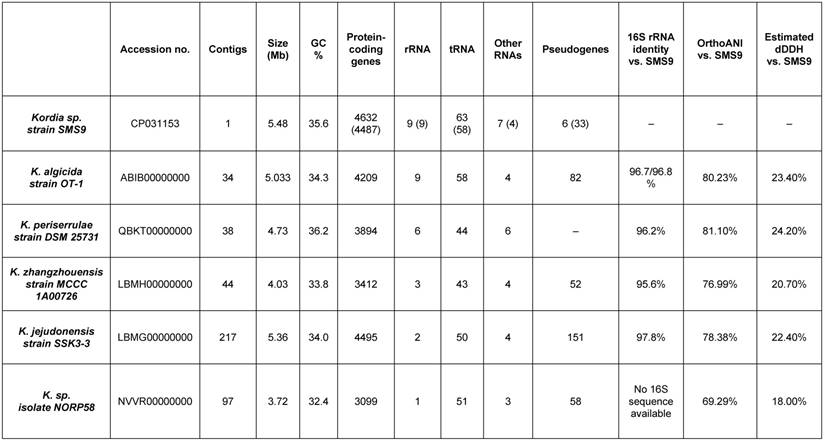
A phylotaxonomic analysis was performed using PhyloPhlAn version 0.99 [6], comparing strain SMS9 to all whole-genome sequenced Flavobacteriaceae species available on NCBI's RefSeq ftp site (ftp://ftp.ncbi.nlm.nih.gov/genomes/refseq/bacteria/; accessed 16 October 2018), in addition to Kordia sp. isolate NORP58 (accession no. NVVR00000000; [7]), which was obtained from GenBank (ftp://ftp.ncbi.nlm.nih.gov/genomes/genbank/bacteria/; accessed 16 October 2018) (Figure 1). With the exception of isolate NORP58, which appears more distantly in the tree, the remaining Kordia species form a well-supported clade which includes strain SMS9 (bootstrap value 1). While the relationships within the clade are less well-supported, this result supports the placement of strain SMS9 within the genus Kordia. Further evidence for this placement was found when comparing the 16S rRNA sequences of strain SMS9 to those of other Kordia strains: 16S similarity to K. jejudonensis strain SSK3-3T (accession no. LBMG00000000; [8]), K. algicida strain OT-1T (accession no. ABIB00000000; [9],[10]), K. periserrulae strain DSM 25731T (accession no. QBKT00000000; [11]) and K. zhangzhouensis strain MCCC 1A00726T (accession no. LBMH00000000; [12]) fell between the proposed species and genus thresholds of 98.7% and 95%, respectively [13], implying that strain SMS9 may belong to an as-yet undescribed Kordia species. Lastly, this placement is supported by the use of the genome similarity metrics dDDH ([14]; calculated using Formula 2 of the Genome-to-Genome Distance Calculator version 2.1 [https://ggdc.dsmz.de/]) and OrthoANI (calculated using OAT version 0.93.1 [15]). Comparison of strain SMS9 to the sequenced Kordia genomes available at NCBI (as mentioned above) gave OrthoANI values of between 69.29% and 81.10% (below the species delineation threshold of 95-96%), and dDDH values of between 18.00% and 24.20% (below the species delineation threshold of 70%, and a figure comparable to other Kordia-Kordia comparisons) (data not shown). Taken together, we propose that strain SMS9 represents a new species within the genus Kordia. As such, we believe this to be the first Kordia species to be initially described from European waters. A comparison of assembly and annotation statistics of strain SMS9 and other Kordia strains is presented in Table 1.
To try and identify genes potentially denoting novel Kordia functionality in strain SMS9, protein sequences obtained from the annotation were subjected to tBLASTn against a database of the other sequenced Kordia genomes, as noted above [4]. Sequences which returned no hits were then subjected to BLASTp to ensure that no further Kordia hits were found [4]. In this way, 23 proteins were found which do not appear to be present in other known Kordia strains. Of these, 21 were labelled as 'hypothetical protein', one was labelled as 'DinB family protein' (although other 'DinB family proteins' are noted for Kordia strains in NCBI), and one was labelled as 'DUF3164 family protein'.
The related species (and type species for the genus) Kordia algicida was first isolated from an algal bloom of the diatom Skeletonema costatum [9], itself related to Skeletonema marinoi, the diatom from whose culture strain SMS9 was obtained. As the name suggests, K. algicida has been shown to be capable of lysing and killing various algae, a process regulated in a quorum sensing-dependent manner [9],[18]. However, as attempts to isolate strain SMS9 from S. marinoi cultures have proven unsuccessful, and the genes behind the aforementioned lysis and quorum sensing remain unknown, further work is required before we can determine whether strain SMS9 and S. marinoi share a similar relationship to that found between K. algicida and S. costatum.
Assembly and annotation features of strain SMS9 compared to other sequenced Kordia species. Bracketed SMS9 annotation figures denote figures obtained from the PGAP annotation [17] (cf. the Prokka annotation).
Nucleotide sequence accession numbers
This whole-genome project has been deposited in GenBank under the accession number CP031153, as part of BioProject No. PRJNA380207.
Abbreviations
CDS: coding sequence; dDDH: digital DNA-DNA hybridisation; OrthoANI: average nucleotide identity by orthology; SMRT: single-molecule real-time; HGAP: Hierarchical Genome Assembly Process; ncRNA: non-coding RNA; tmRNA: transfer-messenger RNA; PGAP: Prokaryotic Genome Annotation Pipeline.
Acknowledgements
This work was supported by the Swedish Research Council Formas (M.T., A.G., grant 2017-00466; A.G., grant 219-2012-2070), the Gordon and Betty Moore Foundation (A.K.C., M.T., A.G.; Grant 4967), and the Swedish Research Council VR (A.K.C., grant 2015-04286).
We thank the Linnéus Centre for Marine Evolutionary Biology (CeMEB, http://cemeb.science.gu.se/) for support. All bioinformatics analyses were run on the Albiorix computer cluster (http://albiorix.bioenv.gu.se/) at the Department of Marine Sciences, University of Gothenburg.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Amin SA, Parker MS, Armbrust EV. Interactions between Diatoms and Bacteria. Microbiol Mol Biol Rev. 2012;76(3):667-684 doi:10.1128/MMBR.00007-12
2. Chin C-S, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods. 2013;10(6):563-569 doi:10.1038/nmeth.2474
3. Bengtsson-Palme J, Hartmann M, Eriksson KM, Pal C, Thorell K, Larsson DGJ, Nilsson RH. metaxa2: improved identification and taxonomic classification of small and large subunit rRNA in metagenomic data. Mol Ecol Resour. 2015;15(6):1403-1414 doi:10.1111/1755-0998.12399
4. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403-410 doi:10.1016/S0022-2836(05)80360-2
5. Seemann T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068-2069 doi:10.1093/bioinformatics/btu153
6. Segata N, Börnigen D, Morgan XC, Huttenhower C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat Commun. 2013;4:2304. doi:10.1038/ncomms3304
7. Tully BJ, Wheat CG, Glazer BT, Huber JA. A dynamic microbial community with high functional redundancy inhabits the cold, oxic subseafloor aquifer. ISME J. 2018;12(1):1-16 doi:10.1038/ismej.2017.187
8. Park S, Jung Y, Yoon J. Kordia jejudonensis sp. nov, isolated from the junction between the ocean and a freshwater spring, and emended description of the genus Kordia. Int J Syst Evol Microbiol. 2014;64(2):657-662 doi:10.1099/ijs.0.058776-0
9. Sohn JH, Lee JH, Yi H, Chun J, Bae KS, Ahn TY, Kim SJ. Kordia algicida gen. nov, sp. nov, an algicidal bacterium isolated from red tide. Int J Syst Evol Microbiol. 2004;54(3):675-680 doi:10.1099/ijs.0.02689-0
10. Lee HS, Kang SG, Kwon KK, Lee JH, Kim SJ. Genome sequence of the algicidal bacterium Kordia algicida OT-1. J Bacteriol. 2011;193(15):4031-4032 doi:10.1128/JB.05241-11
11. Choi A, Oh HM, Yang SJ, Cho JC. Kordia periserrulae sp. nov, isolated from a marine polychaete Periserrula leucophyrna, and emended description of the genus Kordia. Int J Syst Evol Microbiol. 2011;61(4):864-869 doi:10.1099/ijs.0.022764-0
12. Du J, Liu Y, Lai Q, Dong C, Xie Y, Shao Z. Kordia zhangzhouensis sp. nov, isolated from surface freshwater. Int J Syst Evol Microbiol. 2015;65(10):3379-3383 doi:10.1099/ijsem.0.000424
13. Rossi-Tamisier M, Benamar S, Raoult D, Fournier P-E. Cautionary tale of using 16S rRNA gene sequence similarity values in identification of human-associated bacterial species. Int J Syst Evol Microbiol. 2015;65(6):1929-1934 doi:10.1099/ijs.0.000161
14. Meier-Kolthoff JP, Auch AF, Klenk H-P, Göker M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 2013;14:60. doi:10.1186/1471-2105-14-60
15. Lee I, Ouk Kim Y, Park S-C, Chun J. OrthoANI: An improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol. 2016;66(2):1100-1103 doi:10.1099/ijsem.0.000760
16. Rambaut A. http://tree.bio.ed.ac.uk/software/figtree/
17. Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44(14):6614-6624 doi:10.1093/nar/gkw569
18. Paul C, Pohnert G. Interactions of the algicidal bacterium Kordia algicida with diatoms: regulated protease excretion for specific algal lysis. PLoS ONE. 2011;6(6):e21032. doi:10.1371/journal.pone.0021032
Author contact
![]() Corresponding author: Mats Töpel, mats.topelgu.se
Corresponding author: Mats Töpel, mats.topelgu.se

 Global reach, higher impact
Global reach, higher impact