ISSN: 1839-9940
J Genomics 2019; 7:26-30. doi:10.7150/jgen.31911 This volume Cite
Research Paper
SELfies and CELLfies: Whole Genome Sequencing and Annotation of Five Antibiotic Resistant Bacteria Isolated from the Surfaces of Smartphones, An Inquiry Based Laboratory Exercise in a Genomics Undergraduate Course at the Rochester Institute of Technology
The Thomas H. Gosnell School of Life Sciences, Rochester Institute of Technology, Rochester NY, USA
Received 2018-11-30; Accepted 2019-2-5; Published 2019-2-19
Abstract
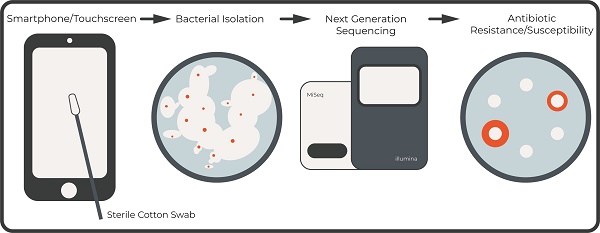
Are touchscreen devices a public health risk for the transmission of pathogenic bacteria, especially those that are resistant to antibiotics? To investigate this, we embarked on a project aimed at isolating and identifying bacteria that are resistant to antibiotics from the screens of smartphones. Touchscreen devices have become ubiquitous in society, and it is important to evaluate the potential risks they pose towards public health, especially as it pertains to the harboring and transmission of pathogenic bacteria that are resistant to antibiotics. Sixteen bacteria were initially isolated of which five were unique (four Staphylococcus species and one Micrococcus species). The genomes of the five unique isolates were subsequently sequenced and annotated. The genomes were analyzed using in silico tools to predict the synthesis of antibiotics and secondary metabolites using the antibiotics and Secondary Metabolite Analysis SHell (antiSMASH) tool in addition to the presence of gene clusters that denote resistance to antibiotics using the Resistance Gene Identifier (RGI) tool. In vivo analysis was also done to assess resistance/susceptibility to four antibiotics that are commonly used in a research laboratory setting. The data presented in this manuscript is the result of a semester-long inquiry based laboratory exercise in the genomics course (BIOL340) in the Thomas H. Gosnell School of Life Sciences/College of Science at the Rochester Institute of Technology.
Keywords: smartphones, touchscreen, public health, antibiotic resistance, Staphylococcus, Micrococcus, secondary metabolites
Introduction
Antibiotic resistance is one of the major public health challenges of the 21st century. The recent death of a United States citizen who became ill with an infection caused by Klebsiella pneumoniae that was resistant to twenty-six antibiotics highlights this important issue (1). Pathogenic bacteria can colonize a number of everyday objects such as dishwashers, personal computers and smartphones, which have become ubiquitous in daily life (2-4). They act as potential vectors for the transmission of bacteria, many of which could be pathogenic and/or resistant to clinically relevant antibiotics. Mobile phones have already been shown to be frequently contaminated with potential pathogens in universities and hospitals (5, 6). Studies have even found that the harboring of bacteria that are resistant to antibiotics is more prevalent on smartphones with touchscreens compared to phones that do not have touchscreens (7). The surfaces of electronic devices in hospitals have already been identified as potential infection threats to patients (3). Alongside personal mobile phones, devices with touchscreens are increasingly being used in public settings. The prevalence of shared public touchscreen devices meant for usage as payment kiosks and entertainment presents a particular risk for transmission of potentially pathogenic bacteria amongst vulnerable segments of the population. For example, touchscreens in chain restaurants are becoming very popular and are often used by young children for entertainment. Many children prefer “finger foods” and may not practice sufficient hand hygiene before food consumption. Therefore, such devices have the potential to become vectors for bacteria and other organisms. Studies conducted so far regarding the presence of bacteria on touchscreen devices have yielded mixed results. There is not a significant difference between the rates of incidence of microbial contamination on touchscreen cell phones versus cell phones that are of the keypad variety (8). A separate study concluded that touchscreen phones have even a lower risk of contamination than keypad phones (9). This difference in the literature pertaining to which type of device is more prevalent regarding the harboring bacteria is not settled. However, because of how widespread touchscreens have become, including and extending beyond mobile phones, the specific types of bacterial contamination are a potential source for concern. Many bacteria isolated from personal smartphones and other touchscreen devices are deemed a normal constituent of the human skin microbiome. For instance, a study conducted at a German university found that Gram-positive cocci associated with then human skin were among the most common microbes isolated from touchscreen smartphones (10). In addition, a 2014 study using Illumina-based next generation sequencing showed an 82% overlap of the dominant operational taxonomic unit (OTU) being shared between mobile phone surfaces and the index finger skin microflora of the respective owners (11). Resistance to both commonly used and last-resort antibiotics has been found in various environmental settings. A study assessing the bacterial diversity from the phones of students and health care workers in Gaza found that over a quarter of the Staphylococcus aureus isolates were resistant to methicillin, and almost 4% were resistant to vancomycin (12). Another study found hospital touchscreens that were contaminated with vancomycin resistant Enterococcus bacteria (13). The presence of antibiotic resistance in hospital settings is especially disconcerting since some patients are particularly susceptible to opportunistic infections.
Here we report the isolation, identification, whole genome sequencing and annotation of five antibiotic resistant bacteria from the surfaces of student owned smartphones. This study was facilitated by an inquiry based laboratory exercise in a genomics course (BIOL 340) in the Thomas H. Gosnell School of Life Sciences on the campus of the Rochester Institute of Technology. The course is primarily designed to introduce undergraduate students to next generation sequencing employing in vivo and in silco aspects of bacterial genomics using questions related to antibiotic resistance as the overarching theme.
Methods
Isolation of bacteria from the screens of smartphones
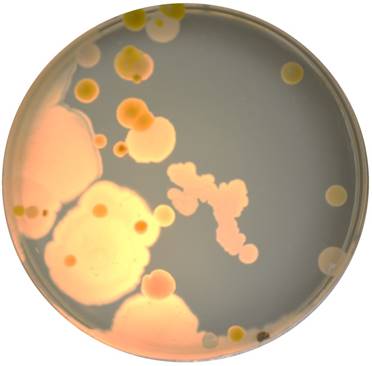
Sixteen student owned smartphones were used in this study. Bacteria were initially isolated on agar plates of tryptic soy (TS), potato dextrose (PD), Reasoner's 2A (R2A), and Luria broth (LB) using sterile cotton swab moistened with R2A broth. The plates were incubated for 30℃ for 48-96 hours. Figure 1 shows the diversity of organisms cultured on a TS agar plate from the screen of one of the smartphone devices.
Example of bacteria cultured from one of the smartphones used in this study. The bacteria were cultured on a tryptic soy (TS) agar plate incubated at 30oC for 48 hours.
Genomic DNA isolation and PCR amplification of the 16S V3/V4 rDNA regions
Genomic DNA was extracted from 1 mL of individual bacteria grown in TSA broth using MolBio DNA extraction kit according to the manufacturer's instructions. The genera of the bacteria were initially identified through nucleotide sequence analysis of the variable 3 and 4 (V3/V4) regions of the 16S rRNA gene, using the 314f/805r primer set (14). The V3/V4 regions were amplified using 15 picomoles of forward and reverse primer (5'-CCTACGGGNGGCWGCAG-3' and 5'-GACTACHVGGGTATCTAATCC-3') using the GoTaq™ Green PCR master mix (Promega). The following PCR conditions was used: 1 cycle at 95oC for 5 minutes, followed by 30 cycles at 95oC for 30 seconds, 55oC for 30 seconds and 72oC for 1 minute. The ~500 bp V3/V4 amplicons were resolved by electrophoresis on a 0.8% (w/v) agarose gel followed by gel extraction using the QIAquick Gel Extraction Kit (Qiagen) followed by Sanger nucleotide sequencing in both directions using the primers that were used for amplification. The individual genera were initially identified using the Basic Local Alignment Search Tool (BLAST) querying the 16S ribosomal RNA sequence database (15).
Bacterial genome sequencing, assembly and annotation
For whole genome sequencing, the genomic DNA was processed using the Nextera XT (Illumina), quantified using a Qubit 2.0 fluorometer and sequenced using a V3 600 cycle cartridge on the MiSeq Illumina platform at the Rochester Institute of Technology Genomics Facility. Sequence quality was assessed using FASTQC version 0.11.6. Adapter trimming was performed on the raw paired-end reads using Trimmomatic version 0.36.5 (16). The trimmed reads were subsequently assembled de novo with Unicycler version 0.3.0b with a minimum contig length of 200 bp (17). Genome assembly metrics were evaluated using the Quality Assessment Tool for Genome Assemblies (QUAST) version 4.1 (18).
Completed FASTA assemblies were uploaded to the NCBI Whole Genome Shotgun project for annotation via the Prokaryotic Genome Annotation Pipeline (PGAP). Secondary metabolite biosynthesis pathways were predicted by uploading completed genome assemblies to antiSMASH using the most recent version 4.6 as of January 7, 2019, using default parameters (20). Each assembled genome was also uploaded to the Comprehensive Antibiotic Resistance Database (CARD), and antibiotic resistance gene prediction was performed using the Resistance Gene Identifier (RGI) version 4.2.2 (21).
Antibiotic resistance/susceptibility assay
Bacteria were streaked on individual LB agar plates supplemented with the antibiotics ampicillin, kanamycin, chloramphenicol, or gentamicin to a final concentration of 50 µg/mL. To ensure consistency with respect to the number of bacterial colonies on each plate, one colony was chosen, suspended and mixed by vortexing in 100 µL of LB broth. Five microliters of the bacteria suspension was pipetted onto each plate, and was then streaked. The inoculated plates were incubated at room temperature for four days to assess resistance and/or susceptibility.
Results and Discussion
Growth was observed on tryptic soy, potato dextrose, and Luria broth after 48 hours when compared to Reasoner's 2A medium that showed growth after 96 hours. Based on the initial amplification and sequencing of the 16S rRNA gene, 5 unique bacteria from the screen were chosen for whole-genome sequencing and annotation. The annotation features for each of the bacterial genomes are summarized in Table 1. The antibiotic resistance/ susceptibility assay revealed that the five isolated strains were resistant to ampicillin, but sensitive to gentamicin and chloramphenicol, while two of the Staphylococcus strains were also able to resist kanamycin (Table 2). Analysis employing antiSMASH showed the production of various secondary metabolites, including antibiotics (Table 3). The results show that all five isolates possess gene clusters for the synthesis of siderophores. Since the host imposes iron limitation, the ability to produce siderophores offers a selective advantage for pathogens. The production of siderophores has been correlated with resistance to carbanepam antibiotics, particularly in P. aeruginosa clinical isolates (22). However, the occurrence of resistance to individual antibiotics did not follow any particular pattern (23). Moreover, siderophores have a wide variety of functions including the regulation of gene expression and quorum sensing (24). Interestingly, Micrococcus sp RIT 608 was the only isolate that possessed a gene cluster for ectoine, a compound that is involved in protecting bacteria from osmotic stress. Given the fact that the bacteria were resistant to antibiotics as per the results in Table 2, we wanted to assess if the isolates possess gene clusters that confer resistance to other antibiotics. The results in Table 4 show a number of predicted gene clusters with homology to antimicrobial resistance genes. Predicted gene clusters were not identified in RIT 608 even though the bacterium is resistant to both ampicillin and kanamycin. Please note that the bacteria in this study could be resistant to additional antibiotics that were not included in this study. Even if the isolates reported in this study are not virulent pathogens, the occurrence of terpene and NRPS biosynthesis gene clusters could indicate antibiotic production by skin commensal Staphylococcus strains. Lugdunin is a recently discovered antibiotic produced by Staphylococcus lugdunensis, which commonly colonizes the human nasal cavity and offers protection against Staphylococcus aureus (25).
Genome annotation information for the isolated strains.
| Organism | Accession no. | Genome size (bp) | % GC Content | Genome Coverage (X) | No. of contigs | No. of ORFs | No. of tRNAs | No. of rRNAs |
|---|---|---|---|---|---|---|---|---|
| Staphylococcus pasteuri sp RIT 605 | RJLY01000000 | 2,546,404 | 31.47 | 202 | 28 | 2,502 | 60 | 4 |
| Micrococcus sp RIT 608 | RJLX00000000 | 2,482,291 | 72.97 | 243 | 140 | 2,332 | 48 | 3 |
| Staphylococcus epidermis sp RIT 611 | RJMA01000000 | 2,483,518 | 31.98 | 251 | 49 | 2,470 | 56 | 4 |
| Staphylococcus hominis sp RIT 612 | RDSL00000000 | 2,146,799 | 31.44 | 239 | 46 | 2,179 | 53 | 4 |
| Staphylococcus cohnii sp RIT 614 | RJLZ01000000 | 2,728,720 | 32.43 | 209 | 44 | 2,653 | 53 | 4 |
The genome size and the GC content were obtained from the Quality Assessment Tool for Genome Assemblies (QUAST) and the other annotation features were obtained from the Prokaryotic Genome Annotation Pipeline (PGAP).
Summary of antibiotic resistance/susceptibility.
| Strain | Ampicillin (50 µg/mL) | Kanamycin (50 µg/mL) | Gentamicin (50 µg/mL) | Chloramphenicol (50 µg/mL) |
|---|---|---|---|---|
| RIT 605 | + | - | - | - |
| RIT 608 | + | + | - | - |
| RIT 611 | + | + | - | - |
| RIT 612 | + | - | - | - |
| RIT 614 | + | - | - | - |
+ indicates resistance and - indicates susceptibility.
Summary of the antibiotics and secondary metabolite analysis shell (antiSMASH) analysis showing the presence/absence of gene clusters involved in the synthesis of antibiotics and secondary metabolites.
| Strain | Terpene | Ectoine | Siderophore | Non-ribosomal peptide synthetase |
|---|---|---|---|---|
| RIT 605 | - | - | + | - |
| RIT 608 | + | + | + | - |
| RIT 611 | - | - | + | + |
| RIT 612 | + | - | + | - |
| RIT 614 | + | - | + | - |
+ indicates the presence of a gene cluster and - indicates that a cluster was not identified.
Summary of antimicrobial resistance gene prediction using the Comprehensive Antibiotic Resistance Database (CARD) database v3.0.0 and the Resistance Gene Identifier (RGI) tool v4.2.2.
| Strain | Drug Class | % ID of matching region | % Length of Reference Sequence |
|---|---|---|---|
| RIT605 | Fusidic acid | 100 | 100 |
| RIT608 | - | - | - |
| RIT611 | Streptogramin antibiotic, Macrolide antibiotic | 100 | 100 |
| Tetracycline antibiotic, Fluoroquinolone antibiotic, Penam, Peptide antibiotic, Cephalosporin, Acridine dye | 95.24 | 100 | |
| Penam | 94.66 | 100 | |
| Diaminopyrimidine antibiotic | 98.14 | 100 | |
| Fluoroquinolone antibiotic, Acridine dye | 99.48 | 100 | |
| RIT612 | Streptogramin antibiotic, Macrolide antibiotic | 99.98 | 100 |
| Fluoroquinolone antibiotic | 99.03 | 100 | |
| Macrolide antibiotic | 90.3 | 100 | |
| RIT614 | Macrolide antibiotic | 90.3 | 100 |
| Streptogramin antibiotic, Macrolide antibiotic | 98.98 | 100 |
The rise in the number of multidrug-resistant bacteria such as methicillin-resistant Staphylococcus aureus (MRSA), Vancomycin intermediate and resistant Staphylococcus aureus, Pseudomonas aeruginosa, Mycobacterium tuberculosis, among others, has led to a significant increase in the morbidity and mortality of humans infected with pathogenic bacteria (26). In the UK, at least 10% of phones used by patients in a hospital were contaminated by hospital-associated pathogens (27). Several studies documented the risk of pathogenic contamination of mobile phones in hospitals with or without antibiotic resistance, and some suggested these as the source of hospital-acquired infections (28-31). One study reported a higher prevalence of MRSA on mobile phones vs the hands of hospital workers in the intensive care and surgical units, suggesting the enrichment of resistant strains rather than merely indicating the presence of skin commensals (32). Another study reported the prevalence of antibiotic resistant bacteria on the surface of mobile phones used by patients in a hospital (33). From a public health perspective, our study further supports the need for awareness of the threat that touchscreen devices may pose with respect to the harboring and/or transmission of pathogenic bacteria, especially those that are resistant to clinically relevant antibiotics. The regular disinfection of patient and hospital worker cell phones and clear usage guidelines for both patients and staff could minimize the risk of nosocomial infections. The current study lends further support to the observation that antibiotic resistant bacteria colonize mobile phone touchscreens that are potential vectors for the transmission of bacteria in a number of settings, other than just the well-studied case of hospitals. It should be noted that in addition to smartphones, other kiosk-like devices at airports, automatic teller machines, restaurants, hotels among others are potential vectors for the transmission of antibiotic resistant pathogens especially since these devices are used by more than one individual and the potential for vectoring pathogens may depend on how often these devices are sanitized and or sterilized.
Nucleotide sequence accession numbers
The genome sequences of the strains described in this study have been deposited in the GenBank database with the accession numbers and annotation features described in Table 1. The version described in this paper is the first version.
Acknowledgements
The authors acknowledge the Thomas H. Gosnell School of Life Sciences (GSoLS) and the College of Science (COS) at the Rochester Institute of Technology (RIT) for ongoing support in addition to the GSoLS laboratory support staff (Allison Healy, Jennifer Liedkie and Gabrielle Crandall) for their hard work and excellent leadership. This work is dedicated to the memory of Dr. David A. Lawlor, our late colleague, teacher, mentor and friend.
Competing Interests
The authors have declared that no competing interest exists.
References
1. STAT: 2017. Woman Killed by a Superbug Resistant to Every Available Antibiotic. https://www.scientificamerican.com/article/woman-killed-by-a-superbug-resistant-to-every-available-antibiotic/
2. Raghupathi PK, Zupančič J, Brejnrod AD. et al. Microbiomes in Dishwashers: Analysis of the microbial diversity and putative opportunistic pathogens in dishwasher biofilm communities. Appl Environ Microbiol. 2018;84(5):e02755-17 doi: 10.1128/AEM.02755-17
3. Neely AN, Sittig DF. Basic microbiologic and infection control information to reduce the potential transmission of pathogens to patients via computer hardware. J Am Med Inform Assoc. 2002;9:500-8
4. Neely AN, Maley MP. Dealing with contaminated computer keyboards and microbial survival. Am J Infect Control. 2001;29:131-2
5. Annand JW, Bajaj N, Sheth A. et al. Potential pathogens and effective disinfectants on public telephones at a large urban United States university. J Environ Health. 2009;71:24-8
6. Tunç K, Olgun U. Microbiology of public telephones. J Infect. 2006;53:140-3
7. Lee YJ, Yoo CG, Lee CT. et al. Contamination rates between smart cell phones and non-smart cell phones of healthcare workers. J Hosp Med. 2013;8:144-7
8. Koroglu M, Gunal S, Yildiz F. et al. Comparison of keypads and touch-screen mobile phones/devices as potential risk for microbial contamination. J Infect Dev Ctries. 2015;9:1308-14
9. Pal P, Roy A, Moore G. et al. Keypad mobile phones are associated with a significant increased risk of microbial contamination compared to touch screen phones. J Infect Prev. 2013;14:65-68
10. Egert M, Späth K, Weik K. et al. Bacteria on smartphone touchscreens in a German university setting and evaluation of two popular cleaning methods using commercially available cleaning products. Folia Microbiol (Praha). 2015;60:159-64
11. Meadow JF, Altrichter AE, Green JL. Mobile phones carry the personal microbiome of their owners. PeerJ. 2014;2:e447
12. Elmanama A, Hassona I, Marouf A. et al. Microbial Load of Touch Screen Mobile Phones Used by University Students and Healthcare Staff. Journal of Arab American University. 2015;1:1-21
13. Gerba CP, Wuollet AL, Raisanen P. et al. Bacterial contamination of computer touch screens. Am J Infect Control. 206; 44: 358-60.
14. Klindworth A, Pruesse E, Schweer T. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013;41:e1
15. Altschul SF, Gish W, Miller W. et al. Basic local alignment search tool. J Mol Biol. 1990;215:403-10
16. Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114-20
17. Wick RR, Judd LM, Gorrie CL. et al. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol. 2017;13:e1005595
18. Gurevich A, Saveliev V, Vyahhi N. et al. QUAST: quality assessment tool for genome assemblies. Bioinformatics. 2013;29:1072-5
19. Weber T, Blin K, Duddela S. et al. antiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 2015;43:W237-43
20. Blin K, Wolf T, Chevrette MG. et al. antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 2017;45:W36-W41
21. Jia B, Raphenya AR, Alcock B. et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017;45:D566-D573
22. Aoki S, Hirakata Y, Kondoh A. et al. Virulence of metallo-beta-lactamase-producing Pseudomonas aeruginosa in vitro and in vivo. Antimicrob Agents Chemother. 2004;48:1876-8
23. Sadeqi Nezhad M, Pordeli H, Ghasemi N. et al. Evaluation of multidrug resistance patterns in siderophore-producing. New Microbes New Infect. 2018;24:38-41
24. Johnstone TC, Nolan EM. Beyond iron: non-classical biological functions of bacterial siderophores. Dalton Trans. 2015;44:6320-39
25. Zipperer A, Konnerth MC, Laux C. et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature. 2016;535:511-6
26. Spellberg B, Guidos R, Gilbert D. et al. The epidemic of antibiotic-resistant infections: a call to action for the medical community from the Infectious Diseases Society of America. Clin Infect Dis. 2008;46:155-64
27. Brady RR, Hunt AC, Visvanathan A. et al. Mobile phone technology and hospitalized patients: a cross-sectional surveillance study of bacterial colonization, and patient opinions and behaviours. Clin Microbiol Infect. 2011;17:830-5
28. Singh S, Acharya S, Bhat M. et al. Mobile phone hygiene: potential risks posed by use in the clinics of an Indian dental school. J Dent Educ. 2010;74:1153-8
29. Ustun C, Cihangiroglu M. Health care workers' mobile phones: a potential cause of microbial cross-contamination between hospitals and community. J Occup Environ Hyg. 2012;9:538-42
30. Akinyemi KO, Atapu AD, Adetona OO. et al. The potential role of mobile phones in the spread of bacterial infections. J Infect Dev Ctries. 2009;3:628-32
31. Jullian-Desayes I, Landelle C, Mallaret MR. et al. Clostridium difficile contamination of health care workers' hands and its potential contribution to the spread of infection: Review of the literature. Am J Infect Control. 2017;45:51-58
32. Ulger F, Esen S, Dilek A. et al. Are we aware how contaminated our mobile phones with nosocomial pathogens? Ann Clin Microbiol Antimicrob. 2009;8:7
33. Vinod Kumar B, Hobani YH, Abdulhaq A. et al. Prevalence of antibacterial resistant bacterial contaminants from mobile phones of hospital inpatients. Libyan J Med. 2014;9:25451
Author contact
Corresponding author: André O. Hudson, Ph.D., Thomas H. Gosnell School of Life Sciences, Rochester Institute of Technology, 85 Lomb Memorial Dr., Rochester, NY 14623 U.S.A. Telephone: 585-475-4259; FAX: 585-475-5766; E-mail: aohsbiedu

 Global reach, higher impact
Global reach, higher impact