ISSN: 1839-9940
J Genomics 2019; 7:7-10. doi:10.7150/jgen.30559 This volume Cite
Short Research Communication
Complete Genome Sequence of Novel Sulfitobacter pseudonitzschiae Strain SMR1, Isolated from a Culture of the Marine Diatom Skeletonema marinoi
1. Department of Marine Sciences, University of Gothenburg, Göteborg, Sweden;
2. Gothenburg Global Biodiversity Centre, Göteborg, Sweden;
3. Department of Biological and Environmental Sciences, University of Gothenburg, Göteborg, Sweden
*M.T. and M.I.M.P. contributed equally to this work.
Received 2018-10-10; Accepted 2018-11-13; Published 2019-1-1
Abstract
When studying diatoms, an important consideration is the role of associated bacteria in the diatom-microbiome holobiont. To that end, bacteria isolated from a culture of Skeletonema marinoi strain R05AC were sequenced, one of which being bacterial strain SMR1, presented here. The genome consists of a circular chromosome and seven circular plasmids, totalling 5,121,602 bp. After phylotaxonomic analysis and 16S rRNA sequence comparison, we place this strain in the taxon Sulfitobacter pseudonitzschiae on account of similarity to the type strain. The annotated genome suggests similar interactions between strain SMR1 and its host diatom as have been shown previously in diatom-associated Sulfitobacter, for example bacterial production of growth hormone for its host, and breakdown of diatom-derived DMSP by Sulfitobacter for use as a sulfur source.
Keywords: Whole Genome Sequencing, Sulfitobacter, Diatom, Skeletonema, Microbiome, Marine sediment
Introduction
The diatom groups of phytoplankton are responsible for around a fifth of the world's photosynthesis, as well as being an important component of marine food webs [1]. As with other organisms, diatoms are found associated with a microbiome with which they can have beneficial interactions including exchange of nutrients, but also be exposed to adverse effects such as parasitism and algicidal activity [2]. Growth of the diatom Skeletonema marinoi is impaired when attempts are made to culture it axenically (Johansson et al., unpublished data), highlighting the importance of the microbiome, and so efforts are being made to identify these associated bacteria, in order to better understand the roles they may play in aiding, or indeed hindering, S. marinoi's growth.
The culture of the host diatom, S. marinoi strain R05AC, was established from a resting cell revived from sediments collected from 14 m depth. This sediment was collected in May 2010 from Öresund, Sweden (55°59.16′ N, 12°44.02′ E), using a box corer. The culture is available from the Gothenburg University Marine Culture Collection (GUMACC) algal bank (https://marine.gu.se/english/research/marine-biology/algal-bank).
Isolation of the bacterial strain SMR1 was performed in April 2016, by dilution streaking on marine agar plates incubated in darkness at 16°C. The culture is currently maintained through monthly subculturing on marine agar plates. Strain SMR1 produces white colonies with firm edges under standard growth conditions, and grows well down to a media pH of 5 and salt ranges between 1-8%.
Assembly and annotation statistics for Sulfitobacter pseudonitzschiae strain SMR1.
| Total assembly | Chromosome | pSMR1-1 | pSMR1-2 | pSMR1-3 | pSMR1-4 | pSMR1-5 | pSMR1-6 | pSMR1-7 | |
|---|---|---|---|---|---|---|---|---|---|
| Assembly | |||||||||
| Number of reads | 94,470 | ||||||||
| Number of bases | 1,055,734,106 bp | ||||||||
| Final assembly size | 5,121,602 bp | 3,572,445 bp | 428,095 bp | 292,917 bp | 284,777 bp | 209,222 bp | 142,107 bp | 99,245 bp | 92,794 bp |
| G+C content | 59.5% | 59.9% | 58.4% | 58.8% | 58.4% | 56.8% | 60.3% | 60.7% | 58.9% |
| Average read coverage | 166.75x | ||||||||
| Annotation | |||||||||
| CDS | 4,938 | 3,482 | 404 | 253 | 273 | 193 | 152 | 93 | 88 |
| Pseudogenes | 11 | 4 | 4 | 1 | 2 | 0 | 0 | 0 | 0 |
| tRNA | 43 | 40 | 0 | 3 | 0 | 0 | 0 | 0 | 0 |
| rRNA | 6 | 3 | 0 | 3 | 0 | 0 | 0 | 0 | 0 |
| ncRNA | 17 | 13 | 2 | 0 | 2 | 0 | 0 | 0 | 0 |
| tmRNA | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
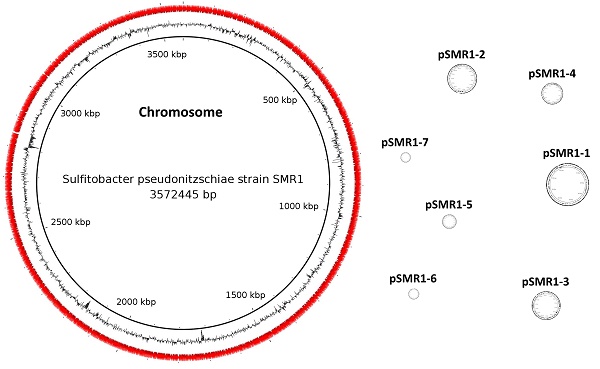
The PacBio RSII platform (Pacific Biosciences, Menlo Park, CA, USA) was used for sequencing strain SMR1. A single SMRT cell was used to sequence a 10kb fragment library, yielding 94,470 uncorrected reads of total size 1.1 Gbp. The de novo PacBio assembler Falcon version 1.7.5 (https://github.com/PacificBiosciences/FALCON/, [3]) was used to assemble these reads (seed read length parameter - 10,600 bp), and contig circularization was confirmed by joining the opposite ends of each contig and realigning the sequencing reads using SMRT Portal version 2.3.0's RS_Resequencing.1 protocol (Pacific Biosciences, [4]) to ensure consistent supporting read coverage. This circularization confirmation also included a correction step using the Quiver algorithm [4]. The final assembly consists of eight circular contigs - one chromosome and seven plasmids - totalling 5,121,602 bp (per-contig figures are given in Table 1), with average read coverage of 166.75x.
Annotation was performed using Prokka version 1.12beta [5]. This pipeline inferred 4,938 CDSs (of which 4,299 have a functional prediction, and 639 are labelled 'hypothetical protein'), 11 pseudogenes, 43 tRNAs, 6 rRNAs, 17 ncRNAs and one tmRNA (per-contig figures are given in Table 1).
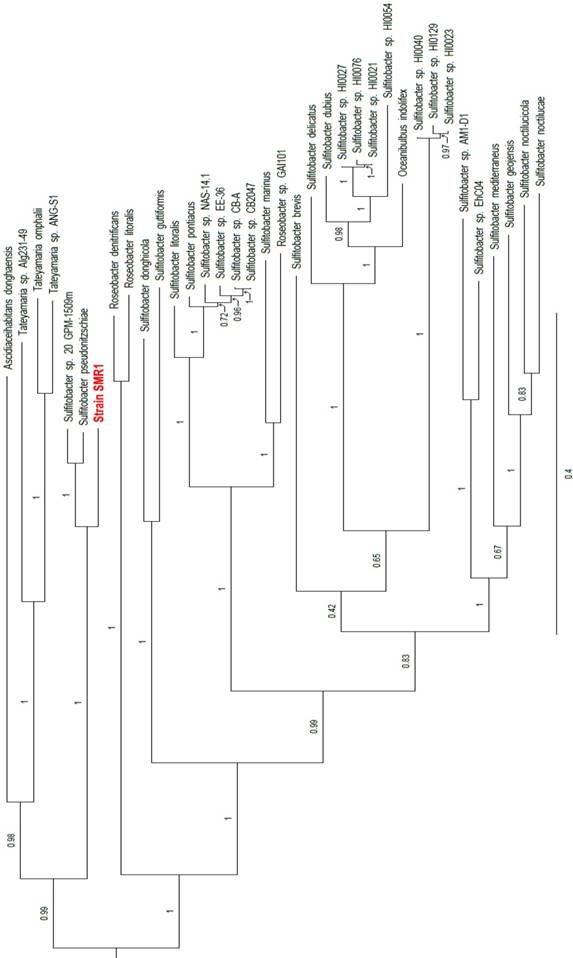
Two rRNA operons were identified in the annotation - one on the chromosome and one on plasmid pSMR1-2 - in which the 16S rRNA sequences are identical to one another. These 16S rRNA genes share 99.2% identity with those of the Sulfitobacter pseudonitzschiae type strain H3T (= DSM 26824T) (accession nos. NZ_JAMD00000000 and NZ_FQVP00000000, respectively), S. guttiformis strain R16 (accession no. AB607871) and S. sp. 20_GPM-1509m (accession no. NZ_JIBC00000000), as well as 99.9% identity with Sulfitobacter sp. SAG13 (accession no. KX268604). To complement the 16S comparison, a phylotaxonomic analysis was performed using PhyloPhlAn version 0.99 [6], which compared strain SMR1 to all whole-genome sequenced Rhodobacteraceae species on NCBI's RefSeq ftp site (ftp://ftp.ncbi.nlm.nih.gov/genomes/refseq/bacteria/; accessed 11 June 2018). This showed strain SMR1 to be sister to the clade of the S. pseudonitzschiae type strain and S. sp. 20_GPM-1509m; the three-strain clade has 100% bootstrap support (Figure 1). Despite the 16S rRNA similarity to strain SMR1, S. guttiformis appears elsewhere in the Sulfitobacter clade. Based on these analyses, we place this strain in the taxon Sulfitobacter pseudonitzschiae.
A point of interest regarding this genome assembly is the large number of plasmids present, five of which are larger than 100 kbp. Multiple techniques were used to try and eliminate the possibility that this was the result of a misassembly, including the use of different assemblers (Canu [7] and HGAP [4], in addition to different parameters in Falcon [3]). Manually joining contigs was also attempted, however nothing yielded fewer circular contigs than the assembly presented here, so we believe this result to be accurate. In addition, there is a precedent for a Sulfitobacter species to have a high number of plasmids - Sulfitobacter sp. AM1-D1 (accession nos. CP018076-CP018081) contains five plasmids, of which three are larger than 100 kbp. A strain of another Rhodobacteraceae species from the same clade, Tateyamaria omphalii (accession nos. CP019312-CP019319), also contains 7 plasmids, although these are smaller (46.3 - 84.5 kbp, compared to 92.8 - 428.1 kbp in strain SMR1).
A previous study by Amin et al. into the interaction between a Sulfitobacter species and a diatom (Sulfitobacter sp. SA11 and Pseudo-nitzschia multiseries) revealed a wide array of interactions between the two organisms, including production of the growth hormone indole-3-acetic acid (auxin) by Sulfitobacter and bacterial breakdown of diatom-produced DMSP for use as a sulfur source [8]. Based on our genomics data, these two examples are also potential interactions in the S. marinoi-SMR1 system. Pathway Tools version 21.0 [9] was used to predict potential pathways in strain SMR1, and this analysis found components of an indole-3-acetate biosynthesis pathway (nitrile hydratase and amidase) in the genome, as well as the dddL gene for DMSP lyase. This implies that strain SMR1 may have a similar relationship with S. marinoi as Sulfitobacter sp. SA11 has with P. multiseries. In addition, associations between Sulfitobacter and Skeletonema have previously been noted [10,11].
Clade of a phylogenetic tree indicating the placement of strain SMR1 (highlighted in red) within the family Rhodobacteraceae. Adapted from tree generated using PhyloPhlAn version 0.99 [6], and visualised using FigTree version 1.4.3 [12]. Branch labels represent bootstrap values; scale bar indicates the mean number of substitutions per site.

Nucleotide sequence accession numbers: This whole-genome project has been deposited in GenBank under the accession numbers CP022415-CP022422, as part of BioProject No. PRJNA380207.
Abbreviations
DMSP: dimethylsulfoniopropionate; SMRT: single-molecule real-time; CDS: coding sequence; tmRNA: transfer-messenger RNA; ncRNA: noncoding RNA; HGAP: hierarchical genome assembly process.
Acknowledgements
This work was supported by the Gordon and Betty Moore Foundation (A.K.C., M.T., A.G.; Grant 4967), the Swedish Research Council VR (A.K.C., grant 2015-04286), Swedish Research Council Formas (M.T., A.G., grant 2017-00466; A.G., grant 219-2012-2070).
We thank the Linnéus Center for Marine Evolutionary Biology (CeMEB, http://cemeb.science.gu.se/) for support. All bioinformatics analyses were run on the Albiorix computer cluster (http://albiorix.bioenv.gu.se/) at the Department of Marine Sciences, University of Gothenburg.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Armbrust EV. The life of diatoms in the world's oceans. Nature. 2009;459(7244):185-192 doi:10.1038/nature08057
2. Amin SA, Parker MS, Armbrust EV. Interactions between Diatoms and Bacteria. Microbiol Mol Biol Rev. 2012;76(3):667-684 doi:10.1128/MMBR.00007-12
3. Gordon D, Huddleston J, Chaisson MJ, Hill CM, Kronenberg ZN, Munson KM, Malig M, Raja A, Fiddes I, Hillier LW, Dunn C, Baker C, Armstrong J, Diekhans M, Paten B, Shendure J, Wilson RK, Haussler D, Chin C-S, Eichler EE. Long-read sequence assembly of the gorilla genome. Science. 2016;352(6281):aae0344. doi:10.1126/science.aae0344
4. Chin C-S, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods. 2013;10(6):563-569 doi:10.1038/nmeth.2474
5. Seemann T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068-2069 doi:10.1093/bioinformatics/btu153
6. Segata N, Börnigen D, Morgan XC, Huttenhower C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat Commun. 2013;4:2304. doi:10.1038/ncomms3304
7. Koren S, Walenz BP, Berlin K, Miller JR, Bergman NH, Phillippy AM. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017;27(5):722-736 doi:10.1101/gr.215087.116
8. Amin SA, Hmelo LR, van Tol HM, Durham BP, Carlson LT, Heal KR, Morales RL, Berthiaume CT, Parker MS, Djunaedi B, Ingalls AE, Parsek MR, Moran MA, Armbrust EV. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature. 2015;522(7554):98-101 doi:10.1038/nature14488
9. Karp PD, Paley S, Romero P. The Pathway Tools Software. Bioinformatics. 2002;18(suppl_1):S225-S232 doi:10.1093/bioinformatics/18.suppl_1.S225
10. Sapp M, Schwaderer AS, Wiltshire KH, Hoppe H-G, Gerdts G, Wichels A. Species-Specific Bacterial Communities in the Phycosphere of Microalgae? Microb Ecol. 2007;53(4):683-699 doi:10.1007/s00248-006-9162-5
11. Grossart H-P, Levold F, Allgaier M, Simon M, Brinkhoff T. Marine diatom species harbour distinct bacterial communities. Environ Microbiol. 2005;7(6):860-873 doi:10.1111/j.1462-2920.2005.00759.x
12. Rambaut A. http://tree.bio.ed.ac.uk/software/figtree/
Author contact
![]() Corresponding author: Mats Töpel, mats.topelgu.se
Corresponding author: Mats Töpel, mats.topelgu.se

 Global reach, higher impact
Global reach, higher impact