ISSN: 1839-9940
J Genomics 2018; 6:113-116. doi:10.7150/jgen.27637 This volume Cite
Short Research Paper
Whole-Genome Sequence of the Novel Antarctobacter heliothermus Strain SMS3, Found in Association with the Marine Diatom Skeletonema marinoi
1. Department of Marine Sciences, University of Gothenburg, Göteborg, Sweden
2. Gothenburg Global Biodiversity Centre, Göteborg, Sweden
3. Department of Biological and Environmental Sciences, University of Gothenburg, Göteborg, Sweden
Received 2018-6-4; Accepted 2018-7-26; Published 2018-9-4
Abstract
As part of an ongoing investigation into the microbiome of the marine diatom Skeletonema marinoi, the bacterial strain SMS3 was isolated from a culture of S. marinoi strain ST54, which had been propagated from a sample of top layer marine sediments taken from the Swedish west coast. We present here the sequenced genome of this bacterium, which we place in the taxon Antarctobacter heliothermus, based on a phylotaxonomic analysis and its high 16S rRNA sequence similarity to the A. heliothermus type strain DSM 11445T. Its 5,331,190 bp genome consists of a circular chromosome and three circular plasmids, and contains 5,019 CDSs. Strain SMS3 contains a phosphatidylcholine synthase gene, as well as genes involved in DMSP degradation, both of which imply a potential symbiotic relationship with its host.
Keywords: Whole Genome Sequencing, Antarctobacter, Diatom, Skeletonema, Microbiome
Introduction
Diatoms are an ecologically important group of phytoplankton, performing around 20% of the Earth's photosynthesis [1] while also being responsible for some varieties of harmful algal blooms [2]. The study of diatom-microbiome interactions has provided important insight in this area [3], with future work on the topic standing to benefit from the use of new sequencing technologies, allowing the genomes of diatom-associated bacteria to be examined in depth. The microbiome of the chain-forming diatom Skeletonema marinoi strain ST54 has not previously been examined, however, and so efforts are being made to identify the bacteria in this species' microbiome, and determine how they interact with their host. One such strain identified is Antarctobacter heliothermus strain SMS3, whose genome sequence is presented here.
Strain SMS3 was isolated from a culture of S. marinoi strain ST54, which was originally established from a germinated resting cell embedded in top layer sediment at 102 m depth [4]. Sediment was collected with a box corer in Kosterfjord, Sweden (58°51.0 N, 10°45.7 E) in May 2009, and the strain has been kept in culture at the Gothenburg University Marine Culture Collection (GUMACC) algal bank (https://marine.gu.se/english/research/marine-biology/algal-bank).
Isolation was performed in April 2016 by dilution streaking on marine agar plates incubated in darkness at 16°C. The bacterial culture has since been maintained on marine agar plates and sub-cultured monthly. Strain SMS3 displays an ovoid, rod-like cell shape, and colonies appear white with a red tint when grown on marine agar, and always display firm edges. No apparent pigment could be isolated by methanolic extraction and UV-vis/HPLC analysis. Cells grow well in a wide range of conditions, including temperatures from 10°C to 30°C (optimum), salt concentrations over 8%, and pH 6-8.5.
Sequencing of the bacterial genome was performed on one SMRT cell using PacBio RSII technology (Pacific Biosciences, Menlo Park, CA, USA), which produced 107,814 uncorrected reads totalling 1.26 Gbp. Canu version 1.3 [5] was used for genome assembly (using a genomeSize parameter of 5.4m), and BLASTn [6] was used to identify where the ends of circular sequences overlapped, after which these areas were trimmed manually. Contig circularization was confirmed by joining the corresponding ends and realigning the reads using the RS_Resequencing.1 protocol on SMRT Portal version 2.3.0 (Pacific Biosciences), which also included a correction step using the Quiver algorithm [7]. The final assembly contained four circular contigs totalling 5,331,190 bp, with average read coverage of 184.56x, where the chromosome is 4,723,013 bp long with a G+C content of 61.6%, plasmid pSMS3-1 is 372,263 bp (G+C 60.3%), plasmid pSMS3-2 is 154,467 bp (G+C 62.8%) and plasmid pSMS3-3 is 81,447 bp (G+C 60.4%). Annotation using Prokka version 1.12beta [8] inferred 5,019 CDSs (4,314 proteins with a functional prediction and 705 labelled as hypothetical), 26 pseudogenes, 47 tRNAs, 6 rRNAs and 11 ncRNAs (per-replicon figures are given in Table 1).
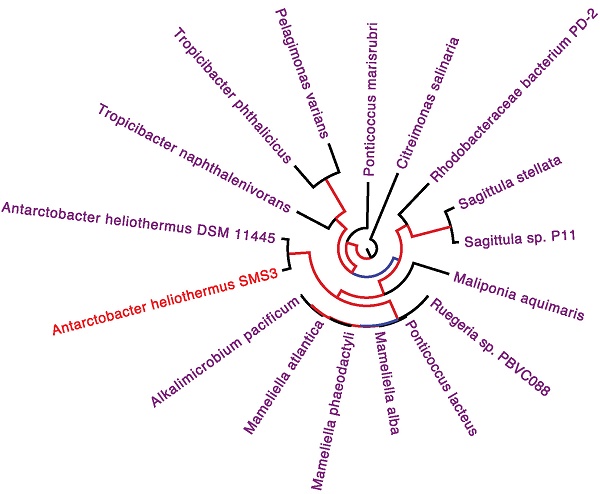
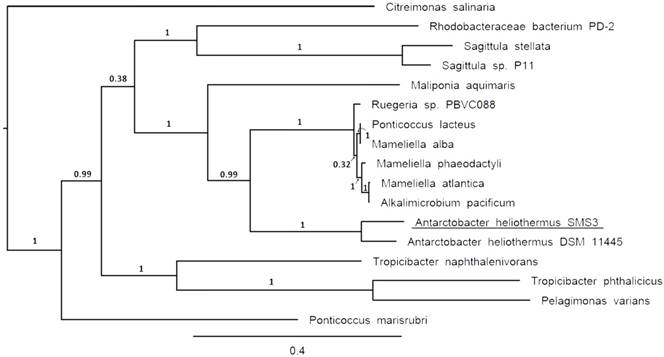
The two identical 16S rRNA sequences of strain SMS3 (both located on the chromosome) have 99.9% identity with that of Antarctobacter heliothermus strain DSM 11445T (accession no. NR_115889). A phylotaxonomic analysis was also performed using PhyloPhlAn version 0.99 [9], which compared strain SMS3 to all whole-genome sequenced species in the family Rhodobacteraceae available at the NCBI RefSeq ftp site (as of 16 January 2017) (ftp://ftp.ncbi.nlm.nih.gov/genomes/refseq/bacteria/). This showed strain SMS3 to be sister to A. heliothermus strain DSM 11445T, forming a clade with 100% bootstrap support (Figure 1). Based on this analysis and the high level of 16S rRNA similarity, we place this strain in the taxon Antarctobacter heliothermus. This placement is further supported by similar G+C content (61.53% for strain SMS3, 62.70% for strain DSM 11445T) and a similar number of protein-coding genes (when run through NCBI's Prokaryotic Genome Annotation Pipeline [10], 4,833 are reported for strain SMS3, and 4,862 are reported for strain DSM 11445T).
Assembly and annotation statistics of Antarctobacter heliothermus strain SMS3. Overlapping bases trimmed from start of contig refers to the number of overlapping bases at contig ends that were removed in order to form circular sequences.
| Total assembly | Chromosome | pSMS3-1 | pSMS3-2 | pSMS3-3 | |
|---|---|---|---|---|---|
| Assembly | |||||
| Number of reads | 107,814 | ||||
| Number of bases | 1,255,903,408 bp | ||||
| Overlapping bases trimmed from start of contig | 15,834 bp | 15,633 bp | 18,046 bp | 17,200 bp | |
| Final assembly size | 5,331,190 bp | 4,723,013 bp | 372,263 bp | 154,467 bp | 81,447 bp |
| G+C content | 61.5% | 61.6% | 60.3% | 62.8% | 60.4% |
| Average read coverage | 184.56x | ||||
| Annotation | |||||
| CDS | 5,019 | 4,440 | 378 | 126 | 75 |
| Pseudogenes | 26 | 18 | 7 | 0 | 1 |
| tRNA | 47 | 47 | 0 | 0 | 0 |
| rRNA | 6 | 6 | 0 | 0 | 0 |
| ncRNA | 11 | 11 | 0 | 0 | 0 |
Clade of a phylogenetic tree of the family Rhodobacteraceae, showing the placement of strain SMS3 (underlined) within the family. Adapted from tree generated using PhyloPhlAn version 0.99 [9], and visualised using FigTree version 1.4.3 [16]. Branch labels represent bootstrap values and the scale bar indicates the mean number of nucleotide substitutions per site.
Metabolic reactions involved in the breakdown of DMSP into either dimethyl sulfide and acrylate, or methanethiol and acetaldehyde, and the genes encoding the enzymes which perform these reactions. Green arrows indicate reactions unique to this pathway according to the MetaCyc database [17]. Bold red text denotes reaction substrates and products and yellow text denotes the enzyme involved in this reaction in strain SMS3. The purple text denotes the locus tag of the gene encoding the enzyme, and blue text denotes the EC number of the enzyme(s) required to perform each reaction. Figure adapted from output generated by Pathway Tools version 21.0 [13].
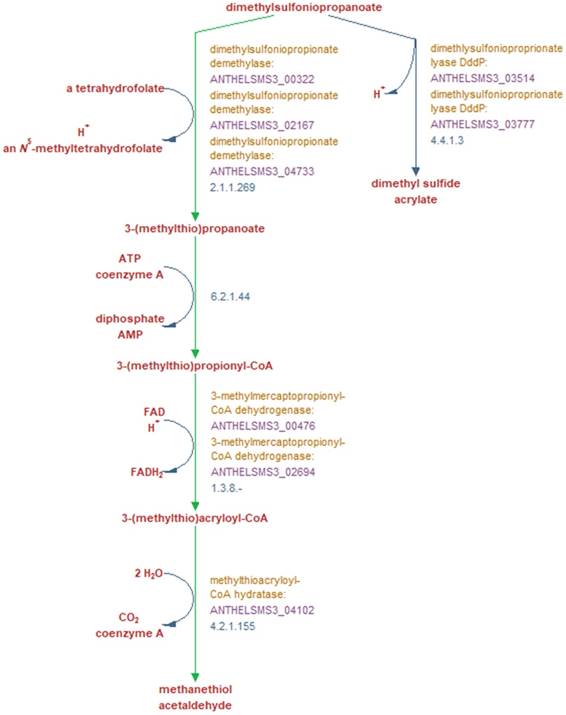
In terms of its potential interactions with S. marinoi, A. heliothermus strain SMS3 contains a phosphatidylcholine synthase gene (pcs; ANTHELSMS3_01509) which is suggestive of a symbiotic or pathogenic relationship with a eukaryote host (reviewed in [11]). Strain SMS3 is also inferred to digest the organosulfur compound DMSP, produced by many phytoplankton and used by some bacteria as a carbon and sulfur source [12]. When strain SMS3 was examined in Pathway Tools version 21.0 [13], predictions for four of the five enzymes in the superpathway of DMSP degradation were made (Figure 2). Of these, three are noted as being unique to this pathway (dimethylsulfoniopropionate demethylase, 3-methylmercaptopropionyl-CoA dehydrogenase, and methylthioacryloyl-CoA hydratase), which reinforces the notion that this strain is capable of DMSP degradation.
Antarctobacter heliothermus is one of only two described species in its genus (the other being A. jejuensis [14]), and the only Antarctobacter species for which a genome sequence has been published. Along with the strain SMS3 genome presented here, the genome of the type strain DSM 11445T has also been sequenced (accession no. NZ_FZON00000000.1). The type strain assembly is however fragmented and comprises 159 scaffolds, making a direct comparison between replicons difficult. Still, it appears that strain SMS3's three plasmids are absent from the type strain reference sequence. Comparison of the two A. heliothermus genomes using Mauve version 20150226 build 10 [15] shows that only relatively short regions are shared between the strain SMS3 plasmids and the type strain assembly. Whereas 80% of the strain SMS3 chromosome aligns to the type strain assembly, only 6% and 9% of plasmids pSMS3-1 and pSMS3-2 align, respectively. In the case of pSMS3-3, Mauve does not note any alignment at all, suggesting this plasmid is entirely novel when compared to the type strain.
Nucleotide sequence accession numbers. This whole-genome project has been deposited in GenBank under the accession numbers CP022540-CP022543, as part of BioProject No. PRJNA380207.
Abbreviations
CDS: coding sequence; DMSP: dimethylsulfoniopropionate; ncRNA: noncoding RNA; SMRT: single-molecule real-time.
Acknowledgements
This work was supported by the Gordon and Betty Moore Foundation (A.K.C., M.T., A.G.; Grant 4967), the Swedish Research Council VR (A.K.C., grant 2015-04286), and the Swedish Research Council Formas (A.G., grant 219-2012-2070).
We thank the Linnéus Center for Marine Evolutionary Biology (CeMEB, http://cemeb.science.gu.se/) for support. All bioinformatics analyses were run on the Albiorix computer cluster (http://albiorix.bioenv.gu.se/) at the Department of Marine Sciences, University of Gothenburg.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Armbrust EV. The life of diatoms in the world's oceans. Nature. 2009;459(7244):185-192 doi:10.1038/nature08057
2. Bates SS. Domoic acid-producing diatoms: Another genus added!. J Phycol. 2000;36(6):978-983 doi:10.1046/j.1529-8817.2000.03661.x
3. Amin SA, Parker MS, Armbrust EV. Interactions between Diatoms and Bacteria. Microbiol Mol Biol Rev. 2012;76(3):667-684 doi:10.1128/MMBR.00007-12
4. Godhe A, Egardt J, Kleinhans D, Sundqvist L, Hordoir R, Jonsson PR. Seascape analysis reveals regional gene flow patterns among populations of a marine planktonic diatom. Proceedings of the Royal Society B. 2013;280(1773):20131599. doi:10.1098/rspb.2013.1599
5. Koren S, Walenz BP, Berlin K, Miller JR, Bergman NH, Phillippy AM. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017;27(5):722-736 doi:10.1101/gr.215087.116
6. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403-410 doi:10.1016/S0022-2836(05)80360-2
7. Chin C-S, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods. 2013;10(6):563-569 doi:10.1038/nmeth.2474
8. Seemann T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068-2069 doi:10.1093/bioinformatics/btu153
9. Segata N, Börnigen D, Morgan XC, Huttenhower C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat Commun. 2013;4:2304. doi:10.1038/ncomms3304
10. Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44(14):6614-6624 doi:10.1093/nar/gkw569
11. Sohlenkamp C, López-Lara IM, Geiger O. Biosynthesis of phosphatidylcholine in bacteria. Prog Lipid Res. 2003;42(2):115-162 doi:10.1016/S0163-7827(02)00050-4
12. Reisch CR, Moran MA, Whitman WB. Bacterial Catabolism of Dimethylsulfoniopropionate (DMSP). Front Microbiol. 2011;2:172. doi:10.3389/fmicb.2011.00172
13. Karp PD, Paley S, Romero P. The Pathway Tools Software. Bioinformatics. 2002;18(suppl_1):S225-S232 doi:10.1093/bioinformatics/18.suppl_1.S225
14. Kim H, Lee JB, Oh HW, Lee H, Shin KS, Bae KS, Park DS. Antarctobacter jejuensis sp. nov, isolated from seawater in Jeju, Korea. Antonie van Leeuwenhoek. 2014;106(6):1081-1087 doi:10.1007/s10482-014-0276-3
15. Darling ACE, Mau B, Blattner FR, Perna NT. Mauve: Multiple Alignment of Conserved Genomic Sequence With Rearrangements. Genome Res. 2004;14(7):1394-1403 doi:10.1101/gr.2289704
16. Rambaut A. http://tree.bio.ed.ac.uk/software/figtree/
17. Caspi R, Billington R, Ferrer L, Foerster H, Fulcher CA, Keseler IM, Kothari A, Krummenacker M, Latendresse M, Mueller LA, Ong Q, Paley S, Subhraveti P, Weaver DS, Karp PD. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 2015;44(D1):D471-D480 doi:10.1093/nar/gkv1164
Author contact
![]() Corresponding author: Mats Töpel, mats.topelgu.se
Corresponding author: Mats Töpel, mats.topelgu.se

 Global reach, higher impact
Global reach, higher impact