ISSN: 1839-9940
J Genomics 2018; 6:74-87. doi:10.7150/jgen.25648 This volume Cite
Research Paper
Genome Wide Analysis Reveals the Extrinsic Cellulolytic and Biohydrogen Generating Abilities of Neocallimastigomycota Fungi
Department of Biology, Lakehead University, 955 Oliver Road, Thunder Bay, Ontario, P7B 5E1, Canada.
Abstract
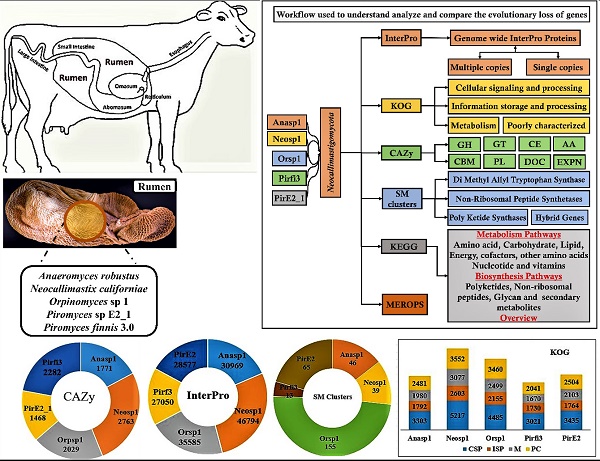
Ruminating animals, especially cattle lack the carbohydrate active enzyme encoding genes which are required for the degradation of the glycosidic linkages of plant cell wall carbohydrates (such as cellulose, hemicellulose, lignin and pectin). Thus, ruminating animals are completely dependent on the microorganisms (anaerobic bacteria and fungi, methanogenic archaea and protozoa) residing in their rumen (hindgut). In this study, we have retrieved and analyzed the complete genome wide annotations of the Neocallimastigomycota division fungi such as Anaeromyces robustus, Neocallismatix californiae, Orpinomyces sp, Piromyces finnis, Piromyces sp E2. We have retrieved the InterPro, CAZy, KOG, KEGG, SM Clusters and MEROPS genome level data of these anaerobic fungi from JGI-MycoCosm database. Results obtained in our study reveals that, the genomes of anaerobic fungi completely lack genes encoding for lignin degrading auxiliary activity enzymes. Contrastingly, these fungi outnumbered other fungi by having highest number of CAZyme encoding genes. The genes encoding for dockerins and carbohydrate binding modules exaggerated other CAZymes which are involved in the structure and functioning of cellulosomes. Presence of cellulosomes and higher number of carbohydrate transport and metabolism genes also endorses the plant cell wall carbohydrate degrading abilities of these fungi. We also reported the tentative total cellulolytic, hemicellulolytic and pectinolytic abilities. And we have explicitly reported the genes, enzymes and the mechanisms involved in structure and functioning of the cellulosomes and hydrogenosomes. Our present work reveals the genomic machinery underlying the extrinsic plant cell wall degrading abilities of the anaerobic fungi. Results obtained in our study can be significantly applied in improving the gut health of cattle and especially in the fields of biofuel, biorefining and bioremediation-based industries.
Keywords: Ruminating animals (cattle), Neocallimastigomycota (Anaerobic fungi), Cellulose, Plant biomass, Cellulosomes, Hydrogenosomes

 Global reach, higher impact
Global reach, higher impact