ISSN: 1839-9940
J Genomics 2016; 4:19-22. doi:10.7150/jgen.16082 This volume Cite
Short Research Communication
External RNA Controls Consortium Beta Version Update
1. Section of Developmental Genomics, Laboratory of Cellular and Developmental Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA
2. Joint Initiative for Metrology in Biology, National Institute of Standards and Technology, Gaithersburg, MD 20899, USA
*These authors contributed equally to this work.
Published 2016-7-26
Abstract
Spike-in RNAs are valuable controls for a variety of gene expression measurements. The External RNA Controls Consortium developed test sets that were used in a number of published reports. Here we provide an authoritative table that summarizes, updates, and corrects errors in the test version that ultimately resulted in the certified Standard Reference Material 2374. We have noted existence of anti-sense RNA controls in the material, corrected sub-pool memberships, and commented on control RNAs that displayed inconsistent behavior.
Keywords: ERCC, spike-in controls, external RNA controls, NIST standard reference materials.
Advances in gene expression profiling technologies not only make it possible for individual groups to ask genome-wide questions, but properly controlled experiments with well-described metadata can be used over and over to make discoveries not envisaged by the data producers. Making these data robust and durable is greatly augmented by standard reference materials. The National Institute of Standards and Technology (NIST) as a part of the External RNA Controls Consortium (ERCC) developed 176 DNA plasmids that can be used as templates for RNA controls [1-3]. NIST Standard Reference Material (SRM) 2374 is a library composed of a subset of 96 plasmids. These same materials were used for commercially available ERCC RNA spike-in mixtures (Ambion/Thermo Fisher Scientific, Waltham, MA), which are formulations of 92 RNA molecules derived from the plasmids. The Commercial collection does not include ERCC-00007, -00018, -00023, and, -00128. One of the test versions that led to the SRM contained 96 RNA sequences transcribed from the plasmids, quantified, and mixed to form defined pools to be added to unknowns in transcription profiling experiments by array, sequencing, PCRs, or other assays. These test pools were widely distributed and were used by the human and model organisms Encyclopedia of DNA Elements projects [4, 5].
When “spiked” into an individual RNA sample, the readout from a single pool of ERCC controls can be used as a ruler. Each pool is designed to have dynamic range of 220. It is noteworthy that the actual linear range of their measurement depends on experimental platforms. Distribution of spike-in measurement fits to straight linear line in RNA-Seq and a monotonic sigmoidal pattern against actual abundance in microarrays or bead-arrays [6], consistent with data compression in hybridization-based techniques [7].
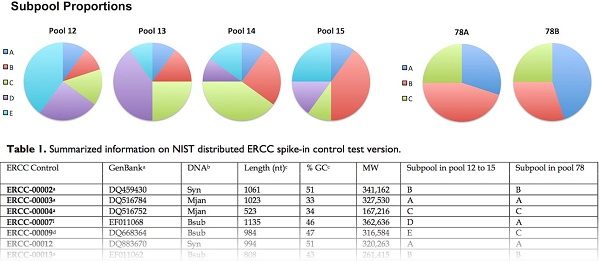
Addition of a single pool of ERCC controls generates useful information, but their use can be enhanced when different pools of spike-in controls from different samples are directly compared. The “pools” of ERCC controls were mixed from multiple “subpools”, such that comparisons between “subpools” that belong to different “pools” generate abundance ratios that can be used as differential expression standards. There were two distinct sets of pools in the test version. Pools 12-15 follow a modified Latin-square design, using 5 different subpools (A-E). The numbers of RNA molecules in subpool A are equal in pools 12-15, and thus subpool A molecules generate a constant 1-to-1 proportion between the pools. Subpools B-E have differing molar concentrations that produce a trend in relative abundance across the pools of 1, 1.5, 2.5, 4-fold. For example, if pool 12 and pool 13 were used for two different samples, the log2-transformed ratios between different subpools will be 0, -0.585, -0.687, -0.737, and 2 [6]. The second set of pools, 78A and 78B, provide a pair of samples with reciprocal changes in relative abundance, i.e. 1.5-fold up and down, producing log2 transformed ratios of 0, 0.585, and -0.585.
While production of the spike-in control RNAs was tightly controlled, it was a test set, and there are multiple cases where measurements of spike-in molecules do not match the original description and/or expectations. In this short note, we summarize data outlining problematic ERCC spike-ins. This information should be used in re-evaluating datasets using the test version, as well as any future work that may use remaining aliquots in circulation (Table 1, and Supplementary Material for more details).
Summarized information on NIST distributed ERCC spike-in control test version.
| ERCC Control | GenBanka | DNAb | Length (nt)c | % GCc | MW | Subpool in pool 12 to 15 | Subpool in pool 78 | |
|---|---|---|---|---|---|---|---|---|
| ERCC-00002a | DQ459430 | Syn | 1061 | 51 | 341,162 | B | B | |
| ERCC-00003a | DQ516784 | Mjan | 1023 | 33 | 327,530 | A | A | |
| ERCC-00004a | DQ516752 | Mjan | 523 | 34 | 167,216 | C | C | |
| ERCC-00007l | EF011068 | Bsub | 1135 | 46 | 362,636 | D | A | |
| ERCC-00009d | DQ668364 | Bsub | 984 | 47 | 316,584 | E | C | |
| ERCC-00012 | DQ883670 | Syn | 994 | 51 | 320,263 | A | A | |
| ERCC-00013a | EF011062 | Bsub | 808 | 43 | 261,415 | B | B | |
| ERCC-00014a,d | DQ875385 | Mjan | 1957 | 44 | 631,409 | C | B | |
| ERCC-00016 | DQ883664 | Syn | 844 | 48 | 271,684 | D | A | |
| ERCC-00017a | DQ459420 | Syn | 1136 | 51 | 367,042 | E | C | |
| ERCC-00018a,l | EF011065 | Bsub | 1026 | 43 | 330,493 | C | C | |
| ERCC-00019 | DQ883651 | Syn | 644 | 49 | 207,543 | B | B | |
| ERCC-00022 | DQ855004 | Syn | 751 | 47 | 241,178 | C | C | |
| ERCC-00023l | DQ516744 | Mjan | 273 | 31 | 88,186 | D | A | |
| ERCC-00024 | DQ854993 | Syn | 536 | 46 | 173,128 | E | C | |
| ERCC-00025a | DQ883689 | Syn | 1994 | 50 | 640,941 | A | A | |
| ERCC-00028a | DQ459419 | Syn | 1130 | 51 | 364,285 | B | B | |
| ERCC-00031a | DQ459431 | Syn | 1138 | 48 | 365,732 | E | C | |
| ERCC-00033 | DQ516796 | Mjan | 2022 | 33 | 651,534 | D | B | |
| ERCC-00034a | DQ855001 | Syn | 1019 | 49 | 328,139 | E | A | |
| ERCC-00035a | DQ459413 | Syn | 1130 | 51 | 364,378 | A | A | |
| ERCC-00039 | DQ883656 | Syn | 740 | 49 | 238,322 | B | B | |
| ERCC-00040a | DQ883661 | Syn | 744 | 53 | 239,738 | C | B | |
| ERCC-00041 | EF011069 | Bsub | 1123 | 45 | 363,007 | D | C | |
| ERCC-00042a | DQ516783 | Mjan | 1023 | 39 | 325,750 | E | B | |
| ERCC-00043a | DQ516787 | Mjan | 1023 | 33 | 330,122 | A | C | |
| ERCC-00044a | DQ459424 | Syn | 1156 | 50 | 372,347 | B | B | |
| ERCC-00046a | DQ516748 | Mjan | 522 | 35 | 168,087 | C | C | |
| ERCC-00048 | DQ883671 | Syn | 992 | 48 | 320,110 | D | B | |
| ERCC-00051 | DQ516740 | Mjan | 274 | 34 | 88,356 | C | A | |
| ERCC-00053a | DQ516785 | Mjan | 1023 | 31 | 327,971 | A | C | |
| ERCC-00054 | DQ516731 | Mjan | 274 | 37 | 88,966 | B | B | |
| ERCC-00057d | DQ668366 | Bsub | 1021 | 50 | 328,287 | C | A | |
| ERCC-00058a | DQ459418 | Syn | 1136 | 50 | 366,548 | D | C | |
| ERCC-00059d | DQ668356 | Bsub | 525 | 48 | 168,750 | E | A | |
| ERCC-00060a | DQ516763 | Mjan | 523 | 31 | 168,195 | A | C | |
| ERCC-00061a | DQ459426 | Syn | 1136 | 50 | 366,454 | B | B | |
| ERCC-00062a | DQ516786 | Mjan | 1023 | 31 | 328,505 | C | A | |
| ERCC-00067 | DQ883653 | Syn | 644 | 47 | 207,451 | D | A | |
| ERCC-00069a | DQ459421 | Syn | 1137 | 50 | 366,664 | E | A | |
| ERCC-00071 | DQ883654 | Syn | 642 | 48 | 206,115 | A | C | |
| ERCC-00073g | DQ668358 | Bsub | 603 | 47 | 193,958 | B | B | |
| ERCC-00074a | DQ516754 | Mjan | 522 | 35 | 167,539 | C | A | |
| ERCC-00075a | DQ516778 | Mjan | 1023 | 36 | 325,442 | D | B | |
| ERCC-00076a | DQ883650 | Syn | 642 | 50 | 206,436 | E | B | |
| ERCC-00077 | DQ516742 | Mjan | 273 | 33 | 87,694 | A | A | |
| ERCC-00078 | DQ883673 | Syn | 993 | 50 | 320,094 | B | B | |
| ERCC-00079 | DQ883652 | Syn | 644 | 49 | 207,757 | A | C | |
| ERCC-00081a | DQ854991 | Syn | 534 | 49 | 172,323 | D | A | |
| ERCC-00083a | DQ516780 | Mjan | 1023 | 35 | 325,668 | E | A | |
| ERCC-00084e | DQ883682 | Syn | 994 | 50 | 320,445 | A | C | |
| ERCC-00085e | DQ883669 | Syn | 844 | 49 | 271,323 | B | B | |
| ERCC-00086a | DQ516791 | Mjan | 1020 | 32 | 328,632 | C | B | |
| ERCC-00092a | DQ459425 | Syn | 1124 | 50 | 361,716 | D | B | |
| ERCC-00095a | DQ516759 | Mjan | 521 | 37 | 166,307 | E | B | |
| ERCC-00096a,i | DQ459429 | Syn | 1107 | 51 | 356,565 | A | C | |
| ERCC-00097a | DQ516758 | Mjan | 523 | 36 | 167,189 | B | B | |
| ERCC-00098a | DQ459415 | Syn | 1143 | 51 | 368,970 | C | C | |
| ERCC-00099a,d | DQ875387 | Bsub | 1350 | 41 | 434,408 | D | A | |
| ERCC-00104a,k | DQ516815 | Mjan | 2022 | 33 | 647,370 | E | C | |
| ERCC-00108d | DQ668365 | Bsub | 1022 | 49 | 328,424 | A | A | |
| ERCC-00109a | DQ854998 | Syn | 536 | 46 | 172,925 | B | B | |
| ERCC-00111 | DQ883685 | Syn | 994 | 47 | 319,359 | C | A | |
| ERCC-00112a | DQ459422 | Syn | 1136 | 47 | 364,932 | D | C | |
| ERCC-00113a,f | DQ883663 | Syn | 840 | 50 | 270,697 | D | A | |
| ERCC-00116d,j | DQ668367 | Bsub | 1991 | 50 | 639,986 | B | B | |
| ERCC-00117a | DQ459412 | Syn | 1136 | 51 | 365,757 | C | A | |
| ERCC-00120a | DQ854992 | Syn | 536 | 48 | 172,605 | D | A | |
| ERCC-00123a | DQ516782 | Mjan | 1022 | 36 | 324,911 | E | C | |
| ERCC-00126a | DQ459427 | Syn | 1119 | 51 | 359,790 | A | C | |
| ERCC-00128a,l | DQ459428 | Syn | 1133 | 48 | 364,405 | B | B | |
| ERCC-00130 | EF011072 | Bsub | 1059 | 46 | 342,268 | C | C | |
| ERCC-00131a | DQ855003 | Syn | 771 | 47 | 248,276 | D | A | |
| ERCC-00134a | DQ516739 | Mjan | 274 | 31 | 88,594 | E | C | |
| ERCC-00136a | EF011063 | Bsub | 1033 | 42 | 333,363 | A | C | |
| ERCC-00137a | DQ855000 | Syn | 537 | 50 | 173,218 | B | B | |
| ERCC-00138a | DQ516777 | Mjan | 1022 | 33 | 327,949 | C | C | |
| ERCC-00142a | DQ883646 | Syn | 493 | 50 | 159,090 | D | C | |
| ERCC-00143 | DQ668362 | Bsub | 784 | 49 | 251,705 | E | A | |
| ERCC-00144h | DQ854995 | Syn | 538 | 46 | 173,404 | A | C | |
| ERCC-00145 | DQ875386 | Bsub | 1042 | 44 | 336,179 | B | B | |
| ERCC-00147a | DQ516790 | Mjan | 1023 | 36 | 331,125 | C | A | |
| ERCC-00148 | DQ883642 | Syn | 494 | 49 | 159,911 | D | B | |
| ERCC-00150 | DQ883659 | Syn | 743 | 47 | 239,128 | E | A | |
| ERCC-00154a | DQ854997 | Syn | 537 | 50 | 173,317 | A | C | |
| ERCC-00156 | DQ883643 | Syn | 494 | 49 | 159,199 | B | B | |
| ERCC-00157a | DQ839618 | Syn | 1019 | 50 | 328,635 | C | C | |
| ERCC-00158a | DQ516795 | Mjan | 1021 | 34 | 328,797 | D | A | |
| ERCC-00160a | DQ883658 | Syn | 743 | 46 | 239,437 | E | C | |
| ERCC-00162a | DQ516750 | Mjan | 523 | 36 | 166,409 | A | A | |
| ERCC-00163a | DQ668359 | Bsub | 543 | 47 | 174,949 | B | B | |
| ERCC-00164a | DQ516779 | Mjan | 1022 | 37 | 324,758 | C | A | |
| ERCC-00165 | DQ668363 | Bsub | 872 | 50 | 279,788 | D | C | |
| ERCC-00168a | DQ516776 | Mjan | 1024 | 34 | 326,399 | E | A | |
| ERCC-00170a | DQ516773 | Mjan | 1024 | 34 | 330,808 | A | B | |
| ERCC-00171 | DQ854994 | Syn | 505 | 48 | 163,022 | B | B |
(a) Sequence mismatches between the GenBank entries and the resequenced RNAs (see [4]).
(b) Syn: De novo synthetic design, Mjan: Methanocaldococcus jannaschii, Bsub: Bacillus subtilis.
(c) Length and GC content include poly(A) sequence.
(d) Reversed (anti-sense) in Pools 12 -15.
(e) ERCC-00084 is not detected. E.g. ERCC-00084 and ERCC-00085, may have both been prepared from ERCC-00085 plasmid. ERCC-00085 behaves as C in some batches of Pool 78A and 78B.
(f) Corrected Pool membership to D and corrected Pool concentrations accordingly.
(g) Poor performing.
(h) Consistently under-reports abundance.
(i) Consistently over-reports abundance in Pools 78A and 78B.
(j) Particularly unsuitable for polyA+ isolation.
(k) ERCC-00104 has a length of either 2202 nt or 2203 nt.
(l) Not present in current commercial collections.
The plasmid DNAs were sequenced and deposited in GenBank, however, the in vitro transcribed RNAs were not sequenced except during testing in RNA-Seq experiments. These experiments made it clear that seven ERCC controls had the complementary sequence indicating that the transcripts were from the other strand (ERCC-00009, -00014, -00057, -00059, -00099, -00108, and -00116). As a result, these spike-in controls would not be measurable in hybridization-based assays [6]. Similarly, they would not be aligned in a strand-specific RNA-Seq analysis unless strand specificity was “turned off” in read quantification steps, or complementary sequences were provided for alignment. Additionally, plasmids are replicated in bacteria, where errors can be introduced. Differences in the sequences of the actual RNAs and the plasmids used for transcript templates are known [4], suggesting that such mutations occurred during plasmid propagation in the test set. The certified values of SRM 2374 are the sequences of the plasmids as distributed in the final set, and were determined by exhaustive sequencing [8].
There were instances of pooling errors in the test set. From multiple experiments that used 78A and 78B, we recognized that ERCC-00085 behaves like Subpool “C”, rather than the intended Subpool “B”. Therefore, when pools 78A and 78B were compared, ERCC-00085 displayed 33.3% increased fold changes than the original description. We have not detected ERCC-00084 in our experiments and it is possible that this RNA was prepared from ERCC-00085 plasmid DNA, effectively increasing the measurement of ERCC-00085. Similarly, we have corrected pool membership of ERCC-00113 from Subpool C to Subpool D from pools 12-15. ERCC-00073 and ERCC-00144 did not provide accurate measurements [4, 6]. One reason for poor measurement may be due to the molecular properties of individual spike-in RNA species (e.g. size and secondary structure). Additionally, a previous study pointed out discrepancy in ERCC-00116 measurements between poly-dT based mRNA enrichment and rRNA depletion protocols [3, 9]. The polyA tails on the ERCC spike-ins are not optimal for PolyA+ selection, and using them prior to library production is not recommended [4]. While there could well be additional instances of unexpected behavior of ERCC spike-in measurements, the information we provide here explains the unexpected ERCC behaviors that we have encountered to date.
Supplementary Material
Supplemental file 1.
Abbreviations
ERCC - External RNA Controls Consortium, NIST - National Institute of Standards and Technology, SRM - Standard Reference Material, ENCODE - Encyclopedia of DNA Elements.
Acknowledgements
The authors would like to acknowledge the careful experimental work by Sarah Helber to prepare the RNA and complex mixtures required for the test pools. This work supported in part by the Intramural Research program of the National Institutes of Health, NIDDK.
Disclaimer
Certain commercial equipment, instruments, or materials are identified in this paper in order to specify the experimental procedure adequately. Such identification is not intended to imply recommendation or endorsement by the National Institute of Standards and Technology (NIST), nor is it intended to imply that the materials or equipment identified are necessarily the best available for the purpose.
Competing Interests
The authors declare no competing interests.
References
1. Baker SC, Bauer SR, Beyer RP, Brenton JD, Bromley B, Burrill J, Causton H, Conley MP, Elespuru R, Fero M. et al. The External RNA Controls Consortium: a progress report. Nature methods. 2005;2(10):731-734
2. ERCC. Proposed methods for testing and selecting the ERCC external RNA controls. BMC genomics. 2005;6:150
3. Munro SA, Lund SP, Pine PS, Binder H, Clevert DA, Conesa A, Dopazo J, Fasold M, Hochreiter S, Hong H. et al. Assessing technical performance in differential gene expression experiments with external spike-in RNA control ratio mixtures. Nature communications. 2014;5:5125
4. Jiang L, Schlesinger F, Davis CA, Zhang Y, Li R, Salit M, Gingeras TR, Oliver B. Synthetic spike-in standards for RNA-seq experiments. Genome research. 2011;21(9):1543-1551
5. https://genome.ucsc.edu/ENCODE/protocols/dataStandards/
6. Pine PS, Munro SA, Parsons JR, McDaniel J, Lucas AB, Lozach J, Myers TG, Su Q, Jacobs-Helber SM, Salit M. Evaluation of the External RNA Controls Consortium (ERCC) reference material using a modified Latin square design. BMC Biotechnol. 2016;16(1):54
7. Malone JH, Oliver B. Microarrays, deep sequencing and the true measure of the transcriptome. BMC biology. 2011;9:34
8. Standard reference material 2374; DNA sequence library for external RNA controls. https://www-s.nist.gov/srmors/certificates/view_certGIF.cfm?certificate=2374
9. Qing T, Yu Y, Du T, Shi L. mRNA enrichment protocols determine the quantification characteristics of external RNA spike-in controls in RNA-Seq studies. Science China Life sciences. 2013;56(2):134-142
Author contact
Corresponding author: Hangnoh Lee, Ph.D. Email: hangnoh.leegov Phone: 301-594-1716/ Fax: 301-496-5239.

 Global reach, higher impact
Global reach, higher impact