ISSN: 1839-9940
J Genomics 2016; 4:1-3. doi:10.7150/jgen.13585 This volume Cite
Short Research Communication
Draft Genome Sequence of Colletotrichum falcatum - A Prelude on Screening of Red Rot Pathogen in Sugarcane
Plant Pathology Section, ICAR-Sugarcane Breeding Institute, Coimbatore 641007, India.
Published 2016-1-30
Abstract

Colletotrichum falcatum, a concealed fungal ascomycete causes red rot, which is a serious disease in sugarcane. It infects economically important stalk tissues, considered as store house of sugar in sugarcane. The study is to find genetic complexities of C. falcatum in establishing this as a stalk infecting pathogen and to decipher the unique lifestyle of this pathogen using NGS technology. We report the draft genome of C. falcatum of about 48.16 Mb in size with 12,270 genes. The genome sequences were compared with other fungal species which revealed that C. falcatum is closely related to C. graminicola and C.sublineola the causal organisms of anthracnose in maize and sorghum. These results brought a new revelation to explore the lifestyle of this unique pathogen which is specialized to infect sugarcane stalk tissues in detail.
Keywords: Genetic complexities, Phylogenomic analysis, NGS Technology.
Introduction
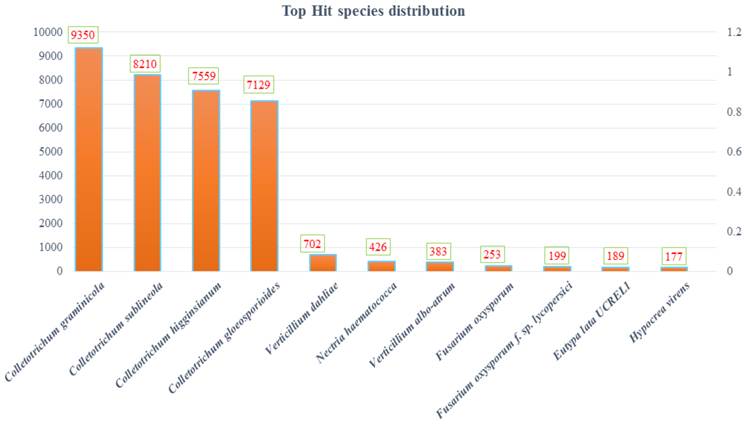
Colletotrichum is one of the most important fungal groups which is devastating and causes huge loss in many economically important crops. C. falcatum, a concealed fungal ascomycete which causes red rot in sugarcane. This pathogen infects sugarcane stalks which is economically important and considered as sugar reservoir in this tall perennial grass. Ever since its report from India during 1902, the disease continues to be a major challenge for sugarcane cultivation in India and many other countries (1). C. falcatum enters the host through nodal tissues of sugarcane, does not form haustoria, and no apparent mechanical pressure is observed during its penetration into stalk. This is an unique pathogen with a different lifestyle in occupying the host compared to other Colletotrichum spp. Earlier in Colletotrichum species several attempts were made to study the lifestyle and pathogenicity using NGS technology (2, 3, 4, 5). Recently, the sugarcane infecting smut genome and transcriptome sequences have been reported (6, 7). This study is focused mainly on finding the genomic signature of C. falcatum and to decipher their uniqueness among Colletotrichum spp. Total nucleic acids were isolated from C. falcatum pathotype Cf671 (MTCC accession number-12142) and cDNA was sequenced on the Illumina Hi Seq 2000 (Genotypic solutions, Bengaluru, India). The read qualities were normalized and primary assembling has been done using Velvet assembler (8) and contig extension/ scaffolding was done by SSPACE (9). The generated high quality, alignable sequence data have been achieved by 2x100 bp library with >150x coverage. The genome of C. falcatum was 48.16 Mb in size with 12,270 predicted genes and 9891 genes were annotated. The gene annotations and structure predictions have been done with AUGUSTUS (10), FGENESH (11) and GeneMarkS (12). Most of the annotated genes were found to be similar with C.graminicola and C.sublineola shared 90% and 84% genes respectively (Fig. 1). The genes encoding biotrophy-necrotrophy transition and membrane transporters were identified and comparative analysis revealed that the number of transporters encoded by C. falcatum is significantly more as compared to that encoded by several other Colletotrichum spp infecting other crops (2, 3, 4, 5). The genome of C. falcatum revealed presence of plant cell wall degrading enzymes (CWDE), Candidate Secretory Effectors (CSEPs), transposable elements, primary and secondary metabolites, membrane transporters, signaling molecules, CAZy, mating proteins and sclerotic development proteins. This study presents a draft genome sequence of C. falcatum, the destructive pathogen of sugarcane and an unique member in Colletotrichum family. The genomic information represents a high resource of the pathogen biology and we identified several putative genes/functions required for its pathogenesis. Further, this work will lead to understand biology, lifestyle and uniqueness of C. falcatum in establishing itself as an intriguing stalk infecting pathogen of sugarcane.
Genomic relation of Colletotrichum falcatum with other fungi based on whole genome consensus from Uniprot database.
Nucleotide Sequence Accession Numbers
This Whole Genome Shotgun project has been deposited at DDBJ/EMBL/GenBank under the accession LPVI00000000. The version described in this paper is version LPVI01000000.
Acknowledgements
The authors are grateful to the Director of ICAR-Sugarcane Breeding Institute, Coimbatore, India for providing facilities and encouragement.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Viswanathan R. Plant Disease: Red rot of sugarcane. New Delhi, India: Anmol Publishers. 2010
2. O'Connell RJ, Thon MR, Hacquard S, Amyotte SG, Kleemann J, Torres MF, Vaillancourt LJ. Lifestyle transitions in plant pathogenic Colletotrichum fungi deciphered by genome and transcriptome analyses. Nat Gene. 2012;44:1060-1065
3. Gan P, Ikeda K, Irieda H, Narusaka M, O'Connell RJ, Narusaka Y, Shirasu K. Comparative genomic and transcriptomic analyses reveal the hemibiotrophic stage shift of Colletotrichum fungi. New Phytol. 2012;197:1236-1249
4. Baroncelli R, Sanz-martín JM, Rech GE, Sukno SA, Thon MR. Draft genome sequence of Colletotrichum sublineola, a destructive pathogen of cultivated sorghum. Genome Announc. 2014;2(3):10-11 doi:10.1128/genomeA.00540-14
5. Baroncelli R, Sreenivasaprasad S, Sukno SA, Thon MR, Holub E. Draft genome sequence of Colletotrichum acutatum Sensu Lato (Colletotrichum fioriniae). Genome Announc. 2014;2(2):e00112-14 doi:10.1128/genomeA.00112-14
6. Wu Q, Xu L, Guo J, Su Y, Que Y. Transcriptome profile analysis of sugarcane responses to Sporisorium scitamineum infection using solexa sequencing technology. BioMed Res Intern. 2013:298920. doi:10.1155/2013/298920
7. Que Y, Xu L, Wu Q, Liu Y, Ling H, Liu Y, Zhang Y, Guo J, Su Y, Chen J, Wang S, Zhang C. Genome sequencing of Sporisorium scitamineum provides insights into the pathogenic mechanisms of sugarcane smut. BMC Genomics. 2014;15(1):996. doi: 10.1186/1471-2164-15-996
8. Zerbino DR, Birney E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008;18:821-829
9. Boetzer M, Henkel CV, Jansen HJ, Butler D, Pirovano W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics. 2011;27(4):578-579 doi:10.1093/bioinformatics/btq683
10. Stanke M, Steinkamp R, Waack S, Morgenstern B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucl Acids Res. 2004;32:W309-W312 doi:10.1093/nar/gkh379
11. Besemer J, Borodovsky M. GeneMark: web software for gene finding in prokaryotes, eukaryotes and viruses. Nucl Acids Res. 2005;33(suppl 2):W451-W454 doi:10.1093/nar/gki487
12. Salamov AA, Solovyey VV. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 2000;10:516-522
Author contact
Corresponding author: E-mail: rasaviswanathanco.in.

 Global reach, higher impact
Global reach, higher impact