ISSN: 1839-9940
J Genomics 2015; 3:57-58. doi:10.7150/jgen.11253 This volume Cite
Short Research Communication
Draft Genome Sequence of Multidrug Resistant Salmonella enterica serovar Weltevreden Isolated from Seafood
1. Faculty of Biomedical Sciences, Nitte University Center for Science Education and Research, UNESCO MIRCEN for Medical & Marine Biotechnology, University Enclave, Medical Sciences Complex, Deralakatte, Mangalore-575018, India
2. School of Information Science (SOIS), Manipal University, Manipal, India
3. FAO Consultant, Subba-Meena, Jayanagar, Mangalore, India
Published 2015-3-1
Abstract
Salmonella enterica subsp. enterica serovar Weltevereden is the most frequent serovar isolated from Asia. Here, we report a draft genome sequence of multidrug resistant Salmonella Weltevreden 9 isolated from seafood. Whole-genome of this isolate and annotation will help enhance the understanding of this pathogenic multidrug-resistant serovar.
Keywords: Salmonella Weltevreden, multidrug resistance, seafood, genome sequence
Introduction
Salmonellosis is a common foodborne disease caused by Salmonella spp. and is of global significance. It is a significant pathogen of food producing animals and remains a primary source of salmonellosis (1). Salmonella spp. has been frequently reported in environmental samples since they find their way to the environment through the excreta of humans and animals. In 2004, the European Union (EU) alone recorded 192,703 human cases of salmonellosis (2). The majority of seafood associated illness is due to the consumption of shellfish harvested from sewage polluted waters. It is well known that fish/shellfish normally do not harbour microorganism like Salmonella but acquire it from contaminated water from which they have been harvested (3). The work of Koonse et al. (2005) (4) showed S. Weltevreden to be the most frequent serovar at 21 % prevalence of the total serovars reported from aquaculture shrimp farms in three different countries. Similar observation were also made by Ponce et al. (2008) (5) who confirmed S. Weltevreden as the most important serovar among the 64 different serovars isolated from seafood. It has been the predominant serovar in seafood in the Asian region reported in several studies (6-8).
In this report the availability of draft genome sequence of S. Weltevreden (SW9) isolated from seafood in India has been announced. S. Weltevreden (SW9) was isolated from seafood (fish) obtained from the fish landing centre in Mangalore, located in the Southwest coast of India. Sample was immediately iced and transported to the laboratory for further analysis. Culture based technique for the isolation and phenotypic identification of the isolate using a battery of biochemical tests was performed as per FDA Bacteriological Analytical Manual. Genotypic identification was done by polymerase chain reaction using genus specific primers.
The isolate was resistant to 6 antimicrobials including tetracycline, chloramphenicol, nalidixic acid, ampicillin, co-trimoxazole and erythromycin. Genomic DNA was extracted from S. Weltevreden (SW9) using a QIAamp DNA minikit (Qiagen, Germany). A concentration of 50ng/µl was used for the genome sequencing. The raw sequence data was generated after library preparation on the Ion Torrent PGM platform and assembled using CLC Genomics Workbench version 6. Structural gene prediction and functional annotation was performed using the Rapid Annotations Subsystems Technology (RAST) server (http://rast.nmpdr.org/) as it was shown to work exceedingly well in our previous studies (9).
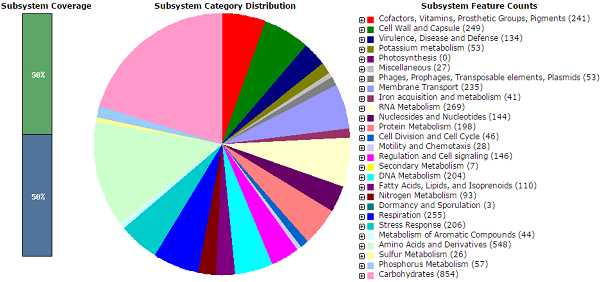
A total of 2, 64,365 reads with a mean read length of 150.2 bp for 200 bp fragmentation chemistry obtained from the Ion PGM was assembled into 716 contigs. The median contig length N50 = 6276. The draft genome had a length of 3,825,753 bp, with 6239 coding sequences and a GC content of 51.8%. The 716 contigs from SW9 were also assembled to NC_003198.1 using Geneious 8.0.3. The analysis obtained from the RAST server revealed 456 subsystems. The annotated genome had 109 genes responsible for resistance to antibiotics and toxic compounds, including 17 genes for multidrug resistance efflux pumps and 18 genes for mdtABCD multidrug resistance cluster and 8 fluoroquinolone resistant genes. There were 235 genes coding for membrane transport proteins. The multidrug resistant isolate (SW9) harbored the resistance genes to antibiotics like tetracycline, chloramphenicol, co-trimoxazole, florfenicol and presented point mutation in the quinolone resistance determining region (QRDR) responsible for resistance to nalidixic acid. The isolate was found to carry resistant genes such as tetB, catA1 for tetracycline and chloramphenicol respectively (10).
Nucleotide sequence accession number
This whole-genome shotgun project has been deposited at DDBJ/EMBL/GenBank under the accession number JPIO00000000.1. The version described in this paper is the first version.
Acknowledgements
The financial support provided by the Indian Council of Medical Research through the project on Antimicrobial resistance (AMR/37/2011-ECD -I) is gratefully acknowledged.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Forshell PL, Wierup M. Salmonella contamination: a significant challenge to the global marketing of animal food products. Rev sci tech Off int Epiz. 2006;25:541-554
2. European food safety authority (EFSA). Trends and sources of zoonoses, zoonotic agents and antimicrobial resistance in the European Union in 2004. EFSA J. 2006;310:23-95
3. Shewan JM. Salmonella. In: (ed.) Borgstrom G. Fish as food, 2nd ed. New York: Academic Press. 1962:443
4. Koonse B, Burkhardt III W, Chirtel S, Hoskin GP. Salmonella and sanitary quality of aquacultured shrimp. J Food Prot. 2005;68:2527-2532
5. Ponce E, Khan AA, Cheng CM, Summage-West C, Cerniglia CE. Prevalence and characterization of Salmonella enterica serovar Weltevreden from imported seafood. Food Microbiol. 2008;25:29-35
6. Boonmar S, Bangtrakulnonth A, Pornrunangwong S, Terajima J, Watanabe H, Kaneko KI, Ogawa M. Epidemiological analysis of Salmonella Enteritidis isolates from humans and broiler chickens in Thailand by phage typing and pulsed-field gel electrophoresis. J Clin Microbiol. 1998;36:971-974
7. Phan TT, Khai LTL, Ogasawara N, Tam NT, Okatani AT, Akiba M, Hayashidani H. Contamination of Salmonella in retail meats and shrimps in the Mekong Delta, Vietnam. J Food Prot. 2005;68:1077-1080
8. Shabarinath S, Sanath KH, Khushiramani R, Karunasagar I, Karunasagar I. Detection and characterization of Salmonella associated with tropical seafood. Int J Food Microbiol. 2007;114:227-233
9. Kumar BK, Deekshit VK, Rai P, Gurtler V, Karunasagar I, Karunasagar I. Draft Genome Sequence of trh+ Vibrio parahaemolyticus VP-49, Isolated from Seafood Harvested along the Mangalore Coast, India. Genome Announc. 2014;2:e00607-14
10. Deekshit VK, Kumar BK, Rai P, Srikumar S, Karunasagar I, Karunasagar I. Detection of class 1 integrons in Salmonella Weltevreden and silent antibiotic resistance genes in some seafood associated nontyphoidal isolates of Salmonella in south-west coast of India. J Appl Microbiol. 2012;112:1113-1122
Author contact
Corresponding author: Prof. (Dr.) Indrani Karunasagar, Director (R & D), UNESCO MIRCEN for Medical & Marine Biotechnology, Faculty of Biomedical Sciences, Nitte University Center for Science Education and Research, University Enclave, Medical Sciences Complex, Deralakatte, Mangalore-575018, India. E.mail: karuna8sagarcom

 Global reach, higher impact
Global reach, higher impact