ISSN: 1839-9940
J Genomics 2017; 5:124-127. doi:10.7150/jgen.21724 This volume Cite
Short Research Paper
Draft genome sequence of a biosurfactant producing, Bacillus aquimaris strain SAMM MCC 3014 isolated from Indian Arabian coastline sea water
1. Department of Microbiology, Elphinstone College, Mumbai 400032;
2. Department of Zoology, DBF Dayanand College of Arts and Science, Solapur 413002;
3. Biochemical Science Division, CSIR-National Chemical Laboratory, Homi Bhabha Road, Pune 411008;
4. Department of Botany, Savitribai Phule Pune University, Ganeshkhind, Pune 411007;
5. Institute of Medical Microbiology, Biomedizinisches Forschungszentrum Seltersberg Schubertstr. 81 Giessen, Germany 35392;
6. Microbial Culture Collection, National Centre for Cell Science, Pune 411021;
7. Yenepoya Research Centre, Yenepoya University, Deralakatte, Mangalore 575018.
Received 2017-7-2; Accepted 2017-8-31; Published 2017-10-8
Abstract
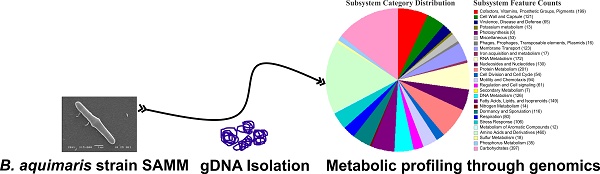
Bacillus aquimaris strain SAMM, a biosurfactant producing moderately halophilic marine bacterium was isolated from Indian Arabian coastline sea water. The strain was found to tolerate up to 2.7 M NaCl indicating osmotic stress sustainable physiological systems. We report here the draft genome sequence of B. aquimaris SAMM, as a candidate bacterium for bioactive surfactant producer. The whole genome sequence with 161 scaffolds, 4,414,932 bp and 44.8% of G+C content for SAMM was obtained using Illumina MiSeq sequencing technology. Annotation was added by the PGAP and RAST prokaryotic genome annotation service and shown 4,247 coding sequences, 123 RNAs genes, classified in 453 subsystems. Several genes encoding enzymatic activities against the high molecular weight polysaccharides, osmotic stress response and siderophore synthesis of potential biotechnological importance were identified in the genome.
Keywords: moderately halophilic, biosurfactant producer, Bacillus aquimaris strain SAMM.
Introduction
Members of the genus Bacillus has been documented to be versatile in metabolism, thus garnering a great amount of interest in studying and exploring its biotechnological perspectives. Nonetheless, marine Bacillus aquimaris outshoot for expedient candidature for the bioactive compound production but is not limited to: starch-degrading enzymes [1], proteases [2, 3], and organic solvent stable alkaline cellulases [4]. We screened marine environment and isolated the beta haemolytic, moderately halophilic SAMM strain from Indian Arabian coastline sea water (17.24 N, 73.37 E) and exploited for antibacterial bio-surfactants. Here we presented the draft genome sequence features of the moderately halophilic SAMM strain. Since the type strain Bacillus aquimaris TF12 DSM 16205 was isolated from the marine sea, this moderate halophilic genome exhibited a range of proteolytic protein genes and new sub-family of alpha-amylases (GH13) reservoir [5]. Moderately halophilic SAMM strain was grown on the nutrient agar medium with 1.5 M NaCl and was found that it can tolerate salts up to 2.7 M. Bacterial genomic DNA which were extracted, purified and sequenced following the protocol as described by Suryavanshi et al., (2017) [6]. The 2×300 bp paired-end sequencing library was constructed using Nextera DNA kits and sequenced on the Illumina® MiSeq platform. A total of 1,366,666 paired reads were obtained. Sequence quality of the genome was analysed for quality control using FastQC software [7]. The reads were trimmed, and only the bases with quality above 20 (Q20) were used for assembly. After analysis, raw sequences were trimmed and assembled using de novo assemblers SPAdes 3.6.1 [8].
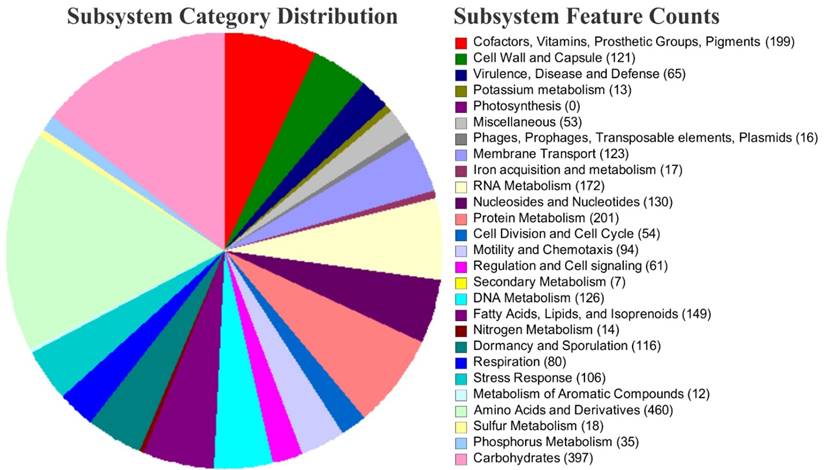
The obtained contigs shorter than 200 bp were eliminated. Annotation and identification of metabolic pathways encoded on the draft genome was carried out using the NCBI Prokaryotic Genome Annotation Pipeline (ver3.3, released 2013) (Available at https://www.ncbi.nlm.nih.gov/genome/annotation_prok/) was used for annotation refinement and registration of the genome at the International Nucleotide Sequence Database Collaboration (GenBank, USA) [9]. The de novo genome assembly of strain SAMM resulted in 161 Scaffolds with a genome size of 4,414,932 bp and G+C content of 44.8%. A total of 4,247 protein-coding sequences (CDSs), 123 copies of RNA: 32 rRNA, 86 tRNAs and 5 ncRNA were identified. Protein coding genes were analyzed by COG database [10] using COGsoft [11] and Pfam domains were predicted using NCBI Batch CD-Search Tool [12]. We obtained 3199 total number of COGs assignments under different classes. In addition, using the RAST server version 2.0 [13], annotation based strain SAMM metabolic profile distributed in 453 subsystems.
Bioinformatic analysis suggested SAMM containing 106 stress response genes, out of 12 sub-categorized for osmotic stress which codes for Choline and Betaine uptake and Betaine Biosynthesis, 28 genes related to oxidative stress, 6 were observed for cold shock and 18 involved in heat shock which is extended dnaK gene cluster. It also possesses the genes clusters for secondary metabolism such as thiazole/oxazole-modified microcin (TOMM) synthesis and auxin biosynthesis. The 17 genes are associated with the acquisition of Iron and their metabolism, out of which 14 codes for siderophore biosynthesis. The SAMM encoding many genes for membrane transport system (123) including 32 for ABC transporters and 12 are related to Uni-, Sym- and Antiporters.
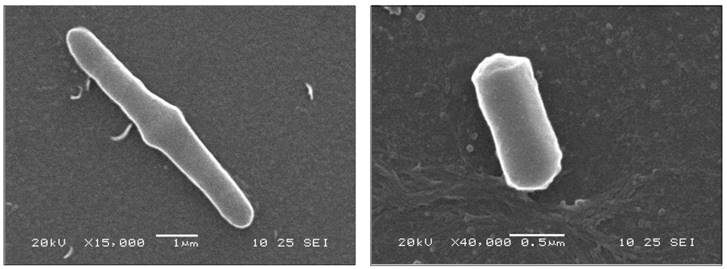
Being marine moderately halophilic, the molecular functionality of the bacterium in the osmotic environment carries osmoadaptive glycine betaine synthesis genes [14]. These were found to be annotated in MINN01000074, MINN01000096 and MINN01000106 accessions along with the stress responsive osmoadaptive ectoine/hydroxyectoine ABC transporter ATP-binding protein cassette under MINN01000011 accessions. Interestingly, SAMM harbours the complete synthesis cluster for polyamine spermidine. Polyamine-spermidine is recognized as a natural cardio-protectant and providing an economically viable natural source in contrast to the chemical synthesis [15]. This bacterium has elongated rod shaped morphology (Figure 1) and having spores generation and Mn (II) oxidization genes were found as similar to its close relative Mn (II) oxidizing marine bacterium Bacillus sp. SG-1 (RAST server Genome ID 161544.4) [16].
Scanning electron micrograph of Bacillus aquimaris SAMM MCC 3014 describing the rod shaped morphology.
Various reports describe the presence of large scale and widespread biological compounds showing the abiogenic and surfactants phenotypes in marine bacteria [17]. Since the strain SAMM showing the bio-surfactant activity phenotype, molecular basis of the origin of this activity is still elusive. The SAMM genome displaying the enterobactin siderophore production transporter (Iron-enterobactin transporter, ATP-binding protein) may be associated with the surfactin molecule synthesis pathway [18].
Genome project information summary.
| Property/Attributes | MCC 3014 | Property/Attributes | MCC 3014 |
|---|---|---|---|
| Finishing quality | High-quality draft | GOLD ID | Gs0127396 |
| Libraries used | 300 bp | RAST genome ID | 189382.10 |
| Sequencing platform | Illumina MiSeq | Genome size (bp) | 4,414,932 |
| Fold coverage | 100.0 X | Total genes | 4,523 |
| Assemblers | SPAdes v. 3.6.1 | Protein coding genes | 4,247 |
| GenBank ID | MINN00000000.1 | RNA genes | 123 |
| NCBI Taxonomy ID | 189382 | Pseudo genes | 153 |
| BioProject ID | PRJNA341790 | Genes assigned to COGs | 3199 |
The overview of subsystem category coverage Bacillus aquimaris SAMM MCC 3014 genome based on RAST server.
Accession numbers
Bacillus aquimaris strain SAMM has been deposited at Microbial Culture Collection, National Centre for Cell Science Pune, under the accession number MCC 3014. This whole-genome shotgun project has been deposited at DDBJ/EMBL/GenBank under the BioProject ID PRJNA341790 with accession number MINN00000000. The version described in this paper is in the first version, MINN00000000.1.
Acknowledgements
MS acknowledges Council of Scientific and Industrial Research, New Delhi, for fellowship. Authors are thankful to Dr. Yogesh Shouche and Microbial Culture Collection project staff (BT/PR10054/NDB/52/94/2007). Authors are thankful to Dr. Shashi Ghodake, DDS, Middletown, CT 06457, USA for proofreading of the manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Puspasari F, Radjasa OK, Noer AS. et al. Raw starch-degrading α-amylase from Bacillus aquimaris MKSC 6.2: isolation and expression of the gene, bioinformatics and biochemical characterization of the recombinant enzyme. J Appl Microbiol. 2013;114(1):108-120
2. Thaz CJ, Jayaraman G. Stability and Detergent Compatibility of a Predominantly β-Sheet Serine Protease from Halotolerant B. aquimaris VITP4 Strain. Appl Biochem Biotechnol. 2014;172(2):687-700
3. Phelan RW, O'Halloran JA, Kennedy J. et al. Diversity and bioactive potential of endospore-forming bacteria cultured from the marine sponge Haliclona simulans. J Appl Microbiol. 2012;112(1):65-78
4. Trivedi N, Gupta V, Kumar M, Kumari P, Reddy CRK, Jha B. Solvent tolerant marine bacterium Bacillus aquimaris secreting organic solvent stable alkaline cellulase. Chemosphere. 2011;83(5):706-712
5. Hernández-González IL, Olmedo-Álvarez G. Draft Whole-Genome Sequence of the Type Strain Bacillus aquimaris. Genome Announc. 2016;4(4):4-5
6. Suryavanshi M, Paul D, Doijad S. et al. Draft genome sequence of Lactobacillus plantarum strains E2C2 and E2C5 isolated from human stool culture. Stand Genomic Sci. 2017;12:15
7. Andrews S. FastQC: a quality control tool for high throughput sequence data. 2010. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
8. Bankevich A, Nurk S, Antipov D. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19(5):455-77
9. Tatusova T, DiCuccio M, Badretdin A. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44(14):6614-24
10. Tatusov RL, Galperin MY, Natale DA, Koonin E V. The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000;28(1):33-6
11. Wu S, Zhu Z, Fu L, Niu B, Li W. WebMGA: a customizable web server for fast metagenomic sequence analysis. BMC Genomics. 2011;12:444
12. Marchler-Bauer A, Derbyshire MK, Gonzales NR. et al. CDD: NCBI's conserved domain database. Nucleic Acids Res. 2015;43(Database issue):D222-6
13. Overbeek R, Olson R, Pusch GD. et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014;42(Database issue):D206-14
14. Nau-Wagner G, Opper D, Rolbetzki A. et al. Genetic Control of Osmoadaptive Glycine Betaine Synthesis in Bacillus subtilis through the Choline-Sensing and Glycine Betaine-Responsive GbsR Repressor. J Bacteriol. 2012;194(10):2703-2714
15. Eisenberg T, Abdellatif M, Schroeder S. et al. Cardioprotection and lifespan extension by the natural polyamine spermidine. Nat Med. 2016;22(12):1428-1438
16. Toyoda K, Tebo BM. Kinetics of Mn(II) oxidation by spores of the marine Bacillus sp. SG-1. Geochim Cosmochim Acta. 2016;189:58-69
17. Oliveira JS, Araújo W, Lopes Sales AI. et al. BioSurfDB: knowledge and algorithms to support biosurfactants and biodegradation studies. Database (Oxford). 2015
18. Ryan RP, Monchy S, Cardinale M. et al. The versatility and adaptation of bacteria from the genus Stenotrophomonas. Nat Rev Microbiol. 2009;7(7):514-525
Author contact
Corresponding author: Suryavanshi Mangesh Vasant, Microbial Culture Collection, National Centre for Cell Science, Pune 411021 and Yenepoya Research Centre, Yenepoya University, Deralakatte, Mangalore 575018 Phone: +912025329000 Email: mangeshnccscom

 Global reach, higher impact
Global reach, higher impact