ISSN: 1839-9940Journal of Genomics
J Genomics 2019; 7:1-6. doi:10.7150/jgen.28153 This volume Cite
Research Paper
Genomic Insights Into A Novel, Alkalitolerant Nitrogen Fixing Bacteria, Azonexus sp. Strain ZS02
1. Department of Biological and Geographical Sciences, University of Huddersfield, Queensgate Campus, Huddersfield, United Kingdom, HD1 3DH.
2. Department of Chemical Sciences, University of Huddersfield, Queensgate Campus, Huddersfield, United Kingdom, HD1 3DH.
Abstract
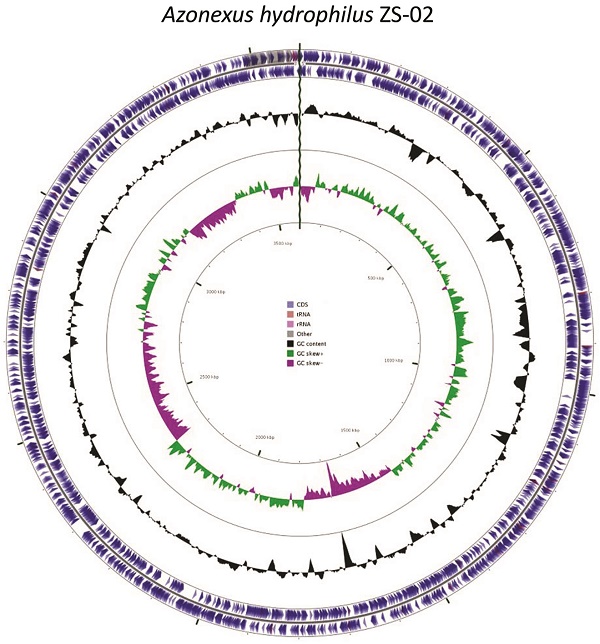
Alkaline environments represent a significant challenge to the growth of micro-organisms. Despite this, there are a number of alkaline environments which contain active microbial communities. Here we describe the genome of a diazotrophic, alkalitolerant strain of Azonexus, which was isolated from a microcosm seeded with hyperalkaline soils resulting from lime depositions. The isolate has a genome size 3.60 Mb with 3431 protein coding genes. The proteome indicated the presence of genes associated with the cycling of nitrogen, in particular the fixation of atmospheric nitrogen. Although closely related to Azonexus hydrophilus strain d8-1 by both 16S (97.9%) and in silico gDNA (84.1%) relatedness, the isolate demonstrates a pH tolerance above that reported for this strain. The proteome contained genes for the complete Na+/H+ antiporter (subunits A to G) for cytoplasmic pH regulation; this may account for the phenotypic characteristics of this strain which exhibited optimal growth conditions of pH 9 and 30°C.
Keywords: Azonexus, facultative aerobic, diazotroph, alkalitolerant, whole genome sequence, Nitrogen fixation, Nitrogen cycle, nitrogen fixing bacteria.
