ISSN: 1839-9940Journal of Genomics
J Genomics 2017; 5:75-76. doi:10.7150/jgen.19673 This volume Cite
Short Research Paper
Complete Genome Sequence of Actinomadura Parvosata Subsp. Kistnae, A Rich Source of Novel Natural Product (Bio-)Chemistry
Biosystems Chemistry, Department of Chemistry and Center for Integrated Protein Science Munich (CIPSM), Technical University of Munich, Lichtenbergstraße 4, 85748 Garching, Germany.
Abstract
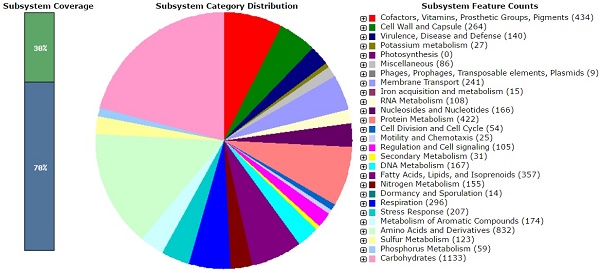
The soil dwelling actinomycete strain Actinomadura parvosata subsp. kistnae is the producer of the antiviral antibiotics kistamicin A and B. Genome sequencing and bioinformatic analysis revealed the presence of the kistamycin biosynthetic gene cluster responsible for the formation of these non-ribosomal peptides as well as an impressive number of yet uncharacterized biosynthetic pathways. This includes polyketide, ribosomal and non-ribosomal peptide and a large number of terpenoid biosynthetic loci encoding yet unknown natural products. The genomic data of this strain is thus a treasure trove for genome mining for novel functional metabolites and new biocatalysts.
Keywords: Actinomadura parvosata subsp. kistnae, whole-genome sequencing, Kistamicin.
Introduction
Actinomycetes are well known to produce a wealth of secondary metabolites with diverse biological activities and complex structures [1]. Included in this broad range of compounds are antibiotics, antivirals and anticancer agents from important natural product classes, such as peptides, polyketides or terpenes [2]. Unfortunately, many more promising secondary metabolites encoded in actinomycete genomes remain undiscovered because their biosynthetic gene clusters are not expressed under standard fermentation conditions [3, 4]. In recent years, these putatively silent gene clusters have become accessible by genome mining [5, 6]. In our search for yet undiscovered natural product chemistry and biochemistry we selected the actinomycete strain Actinomadura parvosata subsp. kistnae (strain designation S382-8) as a promising target organism. It was isolated from a soil sample collected near the Kistna River in India and shown to produce the potent antiviral antibiotics kistamicin A and B [7]. These secondary metabolites exhibit potent antiviral activity against influenza virus type A as well as antimicrobial activity against Gram-positive bacteria and putatively belong to the class of non-ribosomal peptides [7]. To get further insights into the kistamicin biosynthesis and to explore the strains whole metabolic potential we sequenced the genome of Actinomadura parvosata subsp. kistnae (Table 1).
In the draft genome sequence we indeed identified the 48 kb kistamicin biosynthetic gene cluster, showing the typical NRPS assembly line organization [8, 9]. Furthermore, in silico analysis with the program antiSMASH [8] revealed 33 other gene clusters for the biosynthesis of secondary metabolites. This places Actinomadura parvosata subsp. kistnae among those actinomycete strains that harbour the most biosynthetic gene clusters found in this group of bacteria [10]. antiSMASH [8] revealed six ribosomal and non-ribosomal peptide Synthetase (NRPS) systems ranging from 30 kb to 69 kb in size as well as six polyketide synthase (PKS) clusters ranging from 34 kb to 51 kb (3 type I PKS, 1 type II PKS, 1 type III PKS, 1 unspecified) and two mixed NRPS-PKS (both >100 kb). Additionally, seven terpene biosynthetic gene clusters (21 kb to 22 kb), 4 lassopeptide, 3 bacteriocin, 2 lantipeptide, 1 mixed lassopeptide-lantipeptide, 2 siderophore and 1 unspecified cluster have been detected by antiSMASH. Only two of these clusters show 100 % similarity to already characterized pathways, namely those encoding geosmin and 2-methylisoborneol. All other biosynthetic loci have low similarity to previously characterized gene clusters. Together with the sheer number of biosynthetic gene clusters these findings underline the high potential of this organism for the discovery of novel chemistry and biochemical transformations useful in biomedicine and biotechnological applications.
Genome features of Actinomadura parvosata subsp. kistnae.
| Features | Chromosome |
|---|---|
| Length (bp) | 13,559,781 |
| G+C content (%) | 71.7 % |
| CDS | 12784 |
| rRNA genes | 17 |
| tRNA genes | 63 |
| Biosynthetic gene clusters | 34 |
The genomic sequence of Actinomadura parvosata subsp. kistnae was obtained by PacBio single molecule real-time (SMRT®) sequencing. The raw data was generated from three SMRT cells, resulting in a total of approx. 173.600 reads with an average size of 5.9 kb and a N50 read length of 10 kb. Assembly of this data by GATC resulted in 36 polished contigs with an average coverage of 61.3. In total this leads to a size of the chromosome of this strain of approx. 13.56 MBp and thus places it among the largest actinobacterial genome sequences known to date [10]. The G+C content is 71.7%. Importantly, approx. 10% of the very large genome of the strain is devoted to the biosynthesis of specialized metabolites, as shown by antiSMASH analysis [8] described above. The genome was annotated using the RAST server [11] and the software Geneious version 8.1.8 [12], resulting in the annotation of 12784 coding sequences (CDSs) as well as 17 rRNA genes and 34 tRNA loci.
The complete genome sequence was deposited at EMBL under Accession Np. [PRJEB19374].
Acknowledgements
We thank the German Research Foundation DFG (GU-1233/1-1 and CIPSM) and the Hans-Fischer-Gesellschaft e. V. for generous funding. This work was supported by the German Research Foundation (DFG) and the Technical University of Munich (TUM) in the framework of the Open Access Publishing Program.
Competing Interests
The authors have declared that no competing interests exist.
References
1. Nett M, Ikeda H, Moore B.S. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat. Prod. Rep. 2009;26:1362-1384
2. Nawani N, Aigle B, Mandal A, Bodas M, Ghorbel S, Prakash D. Actinomycetes: Role in biotechnology and medicine. Biomed. Res. Int. 2013;2013:687190
3. Hertweck C. Hidden biosynthetic treasures brought to light. Nat. Chem. Biol. 2009;5:450-452
4. Devine R, Hutchings M.I, Holmes N.A. Future directions for the discovery of antibiotics from actinomycete bacteria. Emerging Topics in Life Sciences. 2017 [Epub ahead of print]
5. Rutledge P.J, Challis G.L. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat. Rev. Micro. 2015;13:509-523
6. Ziemert N, Alanjary M, Weber T. The evolution of genome mining in microbes - a review. Nat. Prod. Rep. 2016;33:988-1005
7. Naruse N, Tenmyo O, Kobaru S, Hatori M, Tomita K, Hamagishi Y, Oki T. New antiviral antibiotics, kistamicins A and B I. Taxonomy, production, isolation, physico-chemical properties and biological activities. J. Antibiot. 1993;46:1804-1811
8. Weber T, Blin K, Duddela S. et al. antiSMASH 3.0 a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 2015;43:W237-243
9. Bachmann B.O, Ravel J. Methods for in silico prediction of microbial polyketide and nonribosomal peptide biosynthetic pathways from DNA sequencing data. Methods Enzymol. 2009;458:181-217
10. Doroghazi J.R, Metcalf W.W. Comparative genomics of actinomycetes with a focus on natural product biosynthetic genes. BMC Genomics. 2013;14:611-623
11. Aziz R.K, Bartels D, Best A.A. et al. The RAST Server: Rapid Annotations using Subsystems Technology. BMC Genomics. 2008;9:75-89
12. Kearse M, Moir R, Wilson A. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28:1647-1649
Author contact
![]() Corresponding author: E.Mail: tobias.guldertum.de
Corresponding author: E.Mail: tobias.guldertum.de
Received 2017-2-15
Accepted 2017-5-14
Published 2017-7-2
